RidgeCV#
- class sklearn.linear_model.RidgeCV(alphas=(0.1, 1.0, 10.0), *, fit_intercept=True, scoring=None, cv=None, gcv_mode=None, store_cv_results=False, alpha_per_target=False)[source]#
Ridge regression with built-in cross-validation.
See glossary entry for cross-validation estimator.
By default, it performs efficient Leave-One-Out Cross-Validation.
Read more in the User Guide.
- Parameters:
- alphasarray-like of shape (n_alphas,), default=(0.1, 1.0, 10.0)
Array of alpha values to try. Regularization strength; must be a positive float. Regularization improves the conditioning of the problem and reduces the variance of the estimates. Larger values specify stronger regularization. Alpha corresponds to
1 / (2C)in other linear models such asLogisticRegressionorLinearSVC. If using Leave-One-Out cross-validation, alphas must be strictly positive.For an example on how regularization strength affects the model coefficients, see Ridge coefficients as a function of the L2 Regularization.
- fit_interceptbool, default=True
Whether to calculate the intercept for this model. If set to false, no intercept will be used in calculations (i.e. data is expected to be centered).
- scoringstr, callable, default=None
The scoring method to use for cross-validation. Options:
str: see String name scorers for options.
callable: a scorer callable object (e.g., function) with signature
scorer(estimator, X, y). See Callable scorers for details.None: negative mean squared error if cv is None (i.e. when using leave-one-out cross-validation), or coefficient of determination (\(R^2\)) otherwise.
- cvint, cross-validation generator or an iterable, default=None
Determines the cross-validation splitting strategy. Possible inputs for cv are:
None, to use the efficient Leave-One-Out cross-validation
integer, to specify the number of folds,
an iterable yielding (train, test) splits as arrays of indices.
For integer/None inputs, if
yis binary or multiclass,StratifiedKFoldis used, else,KFoldis used.Refer User Guide for the various cross-validation strategies that can be used here.
- gcv_mode{‘auto’, ‘svd’, ‘eigen’}, default=’auto’
Flag indicating which strategy to use when performing Leave-One-Out Cross-Validation. Options are:
'auto' : same as 'eigen' 'svd' : use singular value decomposition of X when X is dense, fallback to 'eigen' when X is sparse 'eigen' : use eigendecomposition of X X' when n_samples <= n_features or X' X when n_features < n_samples
The ‘auto’ mode is the default and is intended to pick the cheaper option depending on the shape and sparsity of the training data.
- store_cv_resultsbool, default=False
Flag indicating if the cross-validation values corresponding to each alpha should be stored in the
cv_results_attribute (see below). This flag is only compatible withcv=None(i.e. using Leave-One-Out Cross-Validation).Changed in version 1.5: Parameter name changed from
store_cv_valuestostore_cv_results.- alpha_per_targetbool, default=False
Flag indicating whether to optimize the alpha value (picked from the
alphasparameter list) for each target separately (for multi-output settings: multiple prediction targets). When set toTrue, after fitting, thealpha_attribute will contain a value for each target. When set toFalse, a single alpha is used for all targets. This flag is only compatible withcv=None(i.e. using Leave-One-Out Cross-Validation).Added in version 0.24.
- Attributes:
- cv_results_ndarray of shape (n_samples, n_alphas) or shape (n_samples, n_targets, n_alphas), optional
Cross-validation values for each alpha (only available if
store_cv_results=Trueandcv=None). Afterfit()has been called, this attribute will contain the mean squared errors ifscoring is Noneotherwise it will contain standardized per point prediction values.Changed in version 1.5:
cv_values_changed tocv_results_.- coef_ndarray of shape (n_features) or (n_targets, n_features)
Weight vector(s).
- intercept_float or ndarray of shape (n_targets,)
Independent term in decision function. Set to 0.0 if
fit_intercept = False.- alpha_float or ndarray of shape (n_targets,)
Estimated regularization parameter, or, if
alpha_per_target=True, the estimated regularization parameter for each target.- best_score_float or ndarray of shape (n_targets,)
Score of base estimator with best alpha, or, if
alpha_per_target=True, a score for each target.Added in version 0.23.
- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
See also
RidgeRidge regression.
RidgeClassifierClassifier based on ridge regression on {-1, 1} labels.
RidgeClassifierCVRidge classifier with built-in cross validation.
Examples
>>> from sklearn.datasets import load_diabetes >>> from sklearn.linear_model import RidgeCV >>> X, y = load_diabetes(return_X_y=True) >>> clf = RidgeCV(alphas=[1e-3, 1e-2, 1e-1, 1]).fit(X, y) >>> clf.score(X, y) 0.5166...
- fit(X, y, sample_weight=None, **params)[source]#
Fit Ridge regression model with cv.
- Parameters:
- Xndarray of shape (n_samples, n_features)
Training data. If using GCV, will be cast to float64 if necessary.
- yndarray of shape (n_samples,) or (n_samples, n_targets)
Target values. Will be cast to X’s dtype if necessary.
- sample_weightfloat or ndarray of shape (n_samples,), default=None
Individual weights for each sample. If given a float, every sample will have the same weight.
- **paramsdict, default=None
Parameters to be passed to the underlying scorer.
Added in version 1.5: Only available if
enable_metadata_routing=True, which can be set by usingsklearn.set_config(enable_metadata_routing=True). See Metadata Routing User Guide for more details.
- Returns:
- selfobject
Fitted estimator.
Notes
When sample_weight is provided, the selected hyperparameter may depend on whether we use leave-one-out cross-validation (cv=None) or another form of cross-validation, because only leave-one-out cross-validation takes the sample weights into account when computing the validation score.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
Added in version 1.5.
- Returns:
- routingMetadataRouter
A
MetadataRouterencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- predict(X)[source]#
Predict using the linear model.
- Parameters:
- Xarray-like or sparse matrix of shape (n_samples, n_features)
Samples.
- Returns:
- Cndarray of shape (n_samples,) or (n_samples, n_outputs)
Predicted values.
- score(X, y, sample_weight=None)[source]#
Return coefficient of determination on test data.
The coefficient of determination, \(R^2\), is defined as \((1 - \frac{u}{v})\), where \(u\) is the residual sum of squares
((y_true - y_pred)** 2).sum()and \(v\) is the total sum of squares((y_true - y_true.mean()) ** 2).sum(). The best possible score is 1.0 and it can be negative (because the model can be arbitrarily worse). A constant model that always predicts the expected value ofy, disregarding the input features, would get a \(R^2\) score of 0.0.- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples. For some estimators this may be a precomputed kernel matrix or a list of generic objects instead with shape
(n_samples, n_samples_fitted), wheren_samples_fittedis the number of samples used in the fitting for the estimator.- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True values for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
\(R^2\) of
self.predict(X)w.r.t.y.
Notes
The \(R^2\) score used when calling
scoreon a regressor usesmultioutput='uniform_average'from version 0.23 to keep consistent with default value ofr2_score. This influences thescoremethod of all the multioutput regressors (except forMultiOutputRegressor).
- set_fit_request(*, sample_weight: bool | None | str = '$UNCHANGED$') RidgeCV[source]#
Configure whether metadata should be requested to be passed to the
fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter infit.
- Returns:
- selfobject
The updated object.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') RidgeCV[source]#
Configure whether metadata should be requested to be passed to the
scoremethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.
Gallery examples#
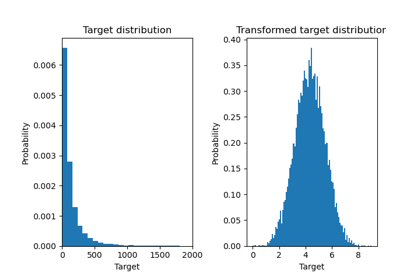
Effect of transforming the targets in regression model
Common pitfalls in the interpretation of coefficients of linear models