LocallyLinearEmbedding#
- class sklearn.manifold.LocallyLinearEmbedding(*, n_neighbors=5, n_components=2, reg=0.001, eigen_solver='auto', tol=1e-06, max_iter=100, method='standard', hessian_tol=0.0001, modified_tol=1e-12, neighbors_algorithm='auto', random_state=None, n_jobs=None)[source]#
Locally Linear Embedding.
Read more in the User Guide.
- Parameters:
- n_neighborsint, default=5
Number of neighbors to consider for each point.
- n_componentsint, default=2
Number of coordinates for the manifold.
- regfloat, default=1e-3
Regularization constant, multiplies the trace of the local covariance matrix of the distances.
- eigen_solver{‘auto’, ‘arpack’, ‘dense’}, default=’auto’
The solver used to compute the eigenvectors. The available options are:
'auto': algorithm will attempt to choose the best method for input data.'arpack': use arnoldi iteration in shift-invert mode. For this method, M may be a dense matrix, sparse matrix, or general linear operator.'dense': use standard dense matrix operations for the eigenvalue decomposition. For this method, M must be an array or matrix type. This method should be avoided for large problems.
Warning
ARPACK can be unstable for some problems. It is best to try several random seeds in order to check results.
- tolfloat, default=1e-6
Tolerance for ‘arpack’ method Not used if eigen_solver==’dense’.
- max_iterint, default=100
Maximum number of iterations for the arpack solver. Not used if eigen_solver==’dense’.
- method{‘standard’, ‘hessian’, ‘modified’, ‘ltsa’}, default=’standard’
standard: use the standard locally linear embedding algorithm. see reference [1]hessian: use the Hessian eigenmap method. This method requiresn_neighbors > n_components * (1 + (n_components + 1) / 2. see reference [2]modified: use the modified locally linear embedding algorithm. see reference [3]ltsa: use local tangent space alignment algorithm. see reference [4]
- hessian_tolfloat, default=1e-4
Tolerance for Hessian eigenmapping method. Only used if
method == 'hessian'.- modified_tolfloat, default=1e-12
Tolerance for modified LLE method. Only used if
method == 'modified'.- neighbors_algorithm{‘auto’, ‘brute’, ‘kd_tree’, ‘ball_tree’}, default=’auto’
Algorithm to use for nearest neighbors search, passed to
NearestNeighborsinstance.- random_stateint, RandomState instance, default=None
Determines the random number generator when
eigen_solver== ‘arpack’. Pass an int for reproducible results across multiple function calls. See Glossary.- n_jobsint or None, default=None
The number of parallel jobs to run.
Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.
- Attributes:
- embedding_array-like, shape [n_samples, n_components]
Stores the embedding vectors
- reconstruction_error_float
Reconstruction error associated with
embedding_- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
- nbrs_NearestNeighbors object
Stores nearest neighbors instance, including BallTree or KDtree if applicable.
See also
SpectralEmbeddingSpectral embedding for non-linear dimensionality reduction.
TSNEDistributed Stochastic Neighbor Embedding.
References
[1]Roweis, S. & Saul, L. Nonlinear dimensionality reduction by locally linear embedding. Science 290:2323 (2000).
[2]Donoho, D. & Grimes, C. Hessian eigenmaps: Locally linear embedding techniques for high-dimensional data. Proc Natl Acad Sci U S A. 100:5591 (2003).
[4]Zhang, Z. & Zha, H. Principal manifolds and nonlinear dimensionality reduction via tangent space alignment. Journal of Shanghai Univ. 8:406 (2004)
Examples
>>> from sklearn.datasets import load_digits >>> from sklearn.manifold import LocallyLinearEmbedding >>> X, _ = load_digits(return_X_y=True) >>> X.shape (1797, 64) >>> embedding = LocallyLinearEmbedding(n_components=2) >>> X_transformed = embedding.fit_transform(X[:100]) >>> X_transformed.shape (100, 2)
- fit(X, y=None)[source]#
Compute the embedding vectors for data X.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Training set.
- yIgnored
Not used, present here for API consistency by convention.
- Returns:
- selfobject
Fitted
LocallyLinearEmbeddingclass instance.
- fit_transform(X, y=None)[source]#
Compute the embedding vectors for data X and transform X.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Training set.
- yIgnored
Not used, present here for API consistency by convention.
- Returns:
- X_newarray-like, shape (n_samples, n_components)
Returns the instance itself.
- get_feature_names_out(input_features=None)[source]#
Get output feature names for transformation.
The feature names out will prefixed by the lowercased class name. For example, if the transformer outputs 3 features, then the feature names out are:
["class_name0", "class_name1", "class_name2"].- Parameters:
- input_featuresarray-like of str or None, default=None
Only used to validate feature names with the names seen in
fit.
- Returns:
- feature_names_outndarray of str objects
Transformed feature names.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- set_output(*, transform=None)[source]#
Set output container.
Refer to the user guide for more details and Introducing the set_output API for an example on how to use the API.
- Parameters:
- transform{“default”, “pandas”, “polars”}, default=None
Configure output of
transformandfit_transform."default": Default output format of a transformer"pandas": DataFrame output"polars": Polars outputNone: Transform configuration is unchanged
Added in version 1.4:
"polars"option was added.
- Returns:
- selfestimator instance
Estimator instance.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- transform(X)[source]#
Transform new points into embedding space.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Training set.
- Returns:
- X_newndarray of shape (n_samples, n_components)
Returns the instance itself.
Notes
Because of scaling performed by this method, it is discouraged to use it together with methods that are not scale-invariant (like SVMs).
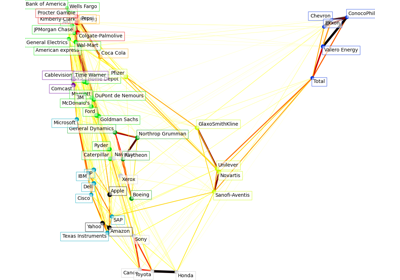
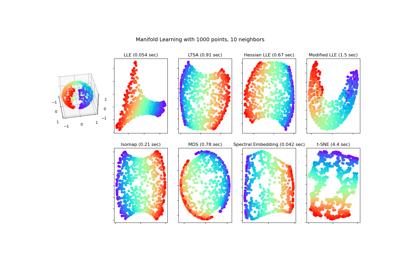
Gallery examples#

Manifold learning on handwritten digits: Locally Linear Embedding, Isomap…