silhouette_samples#
- sklearn.metrics.silhouette_samples(X, labels, *, metric='euclidean', **kwds)[source]#
Compute the Silhouette Coefficient for each sample.
The Silhouette Coefficient is a measure of how well samples are clustered with samples that are similar to themselves. Clustering models with a high Silhouette Coefficient are said to be dense, where samples in the same cluster are similar to each other, and well separated, where samples in different clusters are not very similar to each other.
The Silhouette Coefficient is calculated using the mean intra-cluster distance (
a) and the mean nearest-cluster distance (b) for each sample. The Silhouette Coefficient for a sample is(b - a) / max(a, b). Note that Silhouette Coefficient is only defined if number of labels is 2<= n_labels <= n_samples - 1.This function returns the Silhouette Coefficient for each sample.
The best value is 1 and the worst value is -1. Values near 0 indicate overlapping clusters.
Read more in the User Guide.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples_a, n_samples_a) if metric == “precomputed” or (n_samples_a, n_features) otherwise
An array of pairwise distances between samples, or a feature array. If a sparse matrix is provided, CSR format should be favoured avoiding an additional copy.
- labelsarray-like of shape (n_samples,)
Label values for each sample.
- metricstr or callable, default=’euclidean’
The metric to use when calculating distance between instances in a feature array. If metric is a string, it must be one of the options allowed by
pairwise_distances. IfXis the distance array itself, use “precomputed” as the metric. Precomputed distance matrices must have 0 along the diagonal.- **kwdsoptional keyword parameters
Any further parameters are passed directly to the distance function. If using a
scipy.spatial.distancemetric, the parameters are still metric dependent. See the scipy docs for usage examples.
- Returns:
- silhouettearray-like of shape (n_samples,)
Silhouette Coefficients for each sample.
References
Examples
>>> from sklearn.metrics import silhouette_samples >>> from sklearn.datasets import make_blobs >>> from sklearn.cluster import KMeans >>> X, y = make_blobs(n_samples=50, random_state=42) >>> kmeans = KMeans(n_clusters=3, random_state=42) >>> labels = kmeans.fit_predict(X) >>> silhouette_samples(X, labels) array([...])
Gallery examples#
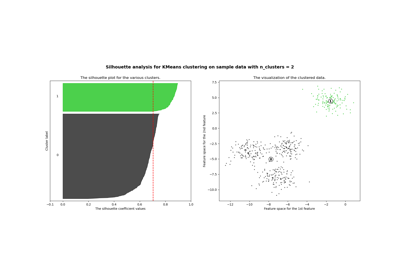
Selecting the number of clusters with silhouette analysis on KMeans clustering
