IsolationForest#
- class sklearn.ensemble.IsolationForest(*, n_estimators=100, max_samples='auto', contamination='auto', max_features=1.0, bootstrap=False, n_jobs=None, random_state=None, verbose=0, warm_start=False)[source]#
Isolation Forest Algorithm.
Return the anomaly score of each sample using the IsolationForest algorithm
The IsolationForest ‘isolates’ observations by randomly selecting a feature and then randomly selecting a split value between the maximum and minimum values of the selected feature.
Since recursive partitioning can be represented by a tree structure, the number of splittings required to isolate a sample is equivalent to the path length from the root node to the terminating node.
This path length, averaged over a forest of such random trees, is a measure of normality and our decision function.
Random partitioning produces noticeably shorter paths for anomalies. Hence, when a forest of random trees collectively produce shorter path lengths for particular samples, they are highly likely to be anomalies.
Read more in the User Guide.
Added in version 0.18.
- Parameters:
- n_estimatorsint, default=100
The number of base estimators in the ensemble.
- max_samples“auto”, int or float, default=”auto”
The number of samples to draw from X to train each base estimator.
If int, then draw
max_samplessamples.If float, then draw
max_samples * X.shape[0]samples.If “auto”, then
max_samples=min(256, n_samples).
If max_samples is larger than the number of samples provided, all samples will be used for all trees (no sampling).
- contamination‘auto’ or float, default=’auto’
The amount of contamination of the data set, i.e. the proportion of outliers in the data set. Used when fitting to define the threshold on the scores of the samples.
If ‘auto’, the threshold is determined as in the original paper.
If float, the contamination should be in the range (0, 0.5].
Changed in version 0.22: The default value of
contaminationchanged from 0.1 to'auto'.- max_featuresint or float, default=1.0
The number of features to draw from X to train each base estimator.
If int, then draw
max_featuresfeatures.If float, then draw
max(1, int(max_features * n_features_in_))features.
Note: using a float number less than 1.0 or integer less than number of features will enable feature subsampling and leads to a longer runtime.
- bootstrapbool, default=False
If True, individual trees are fit on random subsets of the training data sampled with replacement. If False, sampling without replacement is performed.
- n_jobsint, default=None
The number of jobs to run in parallel for
fit.Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.- random_stateint, RandomState instance or None, default=None
Controls the pseudo-randomness of the selection of the feature and split values for each branching step and each tree in the forest.
Pass an int for reproducible results across multiple function calls. See Glossary.
- verboseint, default=0
Controls the verbosity of the tree building process.
- warm_startbool, default=False
When set to
True, reuse the solution of the previous call to fit and add more estimators to the ensemble, otherwise, just fit a whole new forest. See the Glossary.Added in version 0.21.
- Attributes:
- estimator_
ExtraTreeRegressorinstance The child estimator template used to create the collection of fitted sub-estimators.
Added in version 1.2:
base_estimator_was renamed toestimator_.- estimators_list of ExtraTreeRegressor instances
The collection of fitted sub-estimators.
- estimators_features_list of ndarray
The subset of drawn features for each base estimator.
estimators_samples_list of ndarrayThe subset of drawn samples for each base estimator.
- max_samples_int
The actual number of samples.
- offset_float
Offset used to define the decision function from the raw scores. We have the relation:
decision_function = score_samples - offset_.offset_is defined as follows. When the contamination parameter is set to “auto”, the offset is equal to -0.5 as the scores of inliers are close to 0 and the scores of outliers are close to -1. When a contamination parameter different than “auto” is provided, the offset is defined in such a way we obtain the expected number of outliers (samples with decision function < 0) in training.Added in version 0.20.
- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
- estimator_
See also
sklearn.covariance.EllipticEnvelopeAn object for detecting outliers in a Gaussian distributed dataset.
sklearn.svm.OneClassSVMUnsupervised Outlier Detection. Estimate the support of a high-dimensional distribution. The implementation is based on libsvm.
sklearn.neighbors.LocalOutlierFactorUnsupervised Outlier Detection using Local Outlier Factor (LOF).
Notes
The implementation is based on an ensemble of ExtraTreeRegressor. The maximum depth of each tree is set to
ceil(log_2(n))where \(n\) is the number of samples used to build the tree (see [1] for more details).References
[1]F. T. Liu, K. M. Ting and Z. -H. Zhou. “Isolation forest.” 2008 Eighth IEEE International Conference on Data Mining (ICDM), 2008, pp. 413-422.
[2]F. T. Liu, K. M. Ting and Z. -H. Zhou. “Isolation-based anomaly detection.” ACM Transactions on Knowledge Discovery from Data (TKDD) 6.1 (2012): 1-39.
Examples
>>> from sklearn.ensemble import IsolationForest >>> X = [[-1.1], [0.3], [0.5], [100]] >>> clf = IsolationForest(random_state=0).fit(X) >>> clf.predict([[0.1], [0], [90]]) array([ 1, 1, -1])
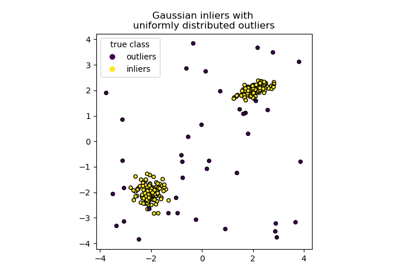
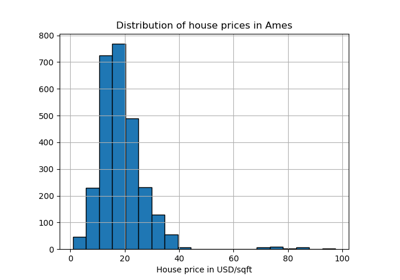
For an example of using isolation forest for anomaly detection see IsolationForest example.
- decision_function(X)[source]#
Average anomaly score of X of the base classifiers.
The anomaly score of an input sample is computed as the mean anomaly score of the trees in the forest.
The measure of normality of an observation given a tree is the depth of the leaf containing this observation, which is equivalent to the number of splittings required to isolate this point. In case of several observations n_left in the leaf, the average path length of an n_left samples isolation tree is added.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The input samples. Internally, it will be converted to
dtype=np.float32and if a sparse matrix is provided to a sparsecsr_matrix.
- Returns:
- scoresndarray of shape (n_samples,)
The anomaly score of the input samples. The lower, the more abnormal. Negative scores represent outliers, positive scores represent inliers.
Notes
The decision_function method can be parallelized by setting a joblib context. This inherently does NOT use the
n_jobsparameter initialized in the class, which is used duringfit. This is because, calculating the score may actually be faster without parallelization for a small number of samples, such as for 1000 samples or less. The user can set the number of jobs in the joblib context to control the number of parallel jobs.from joblib import parallel_backend # Note, we use threading here as the decision_function method is # not CPU bound. with parallel_backend("threading", n_jobs=4): model.decision_function(X)
- fit(X, y=None, sample_weight=None)[source]#
Fit estimator.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The input samples. Use
dtype=np.float32for maximum efficiency. Sparse matrices are also supported, use sparsecsc_matrixfor maximum efficiency.- yIgnored
Not used, present for API consistency by convention.
- sample_weightarray-like of shape (n_samples,), default=None
Sample weights. If None, then samples are equally weighted.
- Returns:
- selfobject
Fitted estimator.
- fit_predict(X, y=None, **kwargs)[source]#
Perform fit on X and returns labels for X.
Returns -1 for outliers and 1 for inliers.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The input samples.
- yIgnored
Not used, present for API consistency by convention.
- **kwargsdict
Arguments to be passed to
fit.Added in version 1.4.
- Returns:
- yndarray of shape (n_samples,)
1 for inliers, -1 for outliers.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
Added in version 1.5.
- Returns:
- routingMetadataRouter
A
MetadataRouterencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- predict(X)[source]#
Predict if a particular sample is an outlier or not.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The input samples. Internally, it will be converted to
dtype=np.float32and if a sparse matrix is provided to a sparsecsr_matrix.
- Returns:
- is_inlierndarray of shape (n_samples,)
For each observation, tells whether or not (+1 or -1) it should be considered as an inlier according to the fitted model.
Notes
The predict method can be parallelized by setting a joblib context. This inherently does NOT use the
n_jobsparameter initialized in the class, which is used duringfit. This is because, predict may actually be faster without parallelization for a small number of samples, such as for 1000 samples or less. The user can set the number of jobs in the joblib context to control the number of parallel jobs.from joblib import parallel_backend # Note, we use threading here as the predict method is not CPU bound. with parallel_backend("threading", n_jobs=4): model.predict(X)
- score_samples(X)[source]#
Opposite of the anomaly score defined in the original paper.
The anomaly score of an input sample is computed as the mean anomaly score of the trees in the forest.
The measure of normality of an observation given a tree is the depth of the leaf containing this observation, which is equivalent to the number of splittings required to isolate this point. In case of several observations n_left in the leaf, the average path length of an n_left samples isolation tree is added.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The input samples.
- Returns:
- scoresndarray of shape (n_samples,)
The anomaly score of the input samples. The lower, the more abnormal.
Notes
The score function method can be parallelized by setting a joblib context. This inherently does NOT use the
n_jobsparameter initialized in the class, which is used duringfit. This is because, calculating the score may actually be faster without parallelization for a small number of samples, such as for 1000 samples or less. The user can set the number of jobs in the joblib context to control the number of parallel jobs.from joblib import parallel_backend # Note, we use threading here as the score_samples method is not CPU bound. with parallel_backend("threading", n_jobs=4): model.score(X)
- set_fit_request(*, sample_weight: bool | None | str = '$UNCHANGED$') IsolationForest[source]#
Configure whether metadata should be requested to be passed to the
fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter infit.
- Returns:
- selfobject
The updated object.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
Gallery examples#
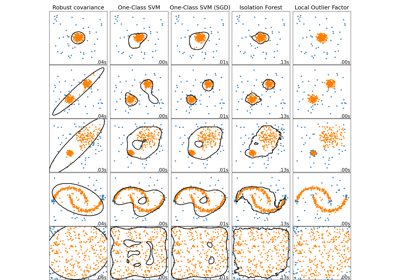
Comparing anomaly detection algorithms for outlier detection on toy datasets