completeness_score#
- sklearn.metrics.completeness_score(labels_true, labels_pred)[source]#
Compute completeness metric of a cluster labeling given a ground truth.
A clustering result satisfies completeness if all the data points that are members of a given class are elements of the same cluster.
This metric is independent of the absolute values of the labels: a permutation of the class or cluster label values won’t change the score value in any way.
This metric is not symmetric: switching
label_truewithlabel_predwill return thehomogeneity_scorewhich will be different in general.Read more in the User Guide.
- Parameters:
- labels_truearray-like of shape (n_samples,)
Ground truth class labels to be used as a reference.
- labels_predarray-like of shape (n_samples,)
Cluster labels to evaluate.
- Returns:
- completenessfloat
Score between 0.0 and 1.0. 1.0 stands for perfectly complete labeling.
See also
homogeneity_scoreHomogeneity metric of cluster labeling.
v_measure_scoreV-Measure (NMI with arithmetic mean option).
References
Examples
Perfect labelings are complete:
>>> from sklearn.metrics.cluster import completeness_score >>> completeness_score([0, 0, 1, 1], [1, 1, 0, 0]) 1.0
Non-perfect labelings that assign all classes members to the same clusters are still complete:
>>> print(completeness_score([0, 0, 1, 1], [0, 0, 0, 0])) 1.0 >>> print(completeness_score([0, 1, 2, 3], [0, 0, 1, 1])) 0.999
If classes members are split across different clusters, the assignment cannot be complete:
>>> print(completeness_score([0, 0, 1, 1], [0, 1, 0, 1])) 0.0 >>> print(completeness_score([0, 0, 0, 0], [0, 1, 2, 3])) 0.0
Gallery examples#
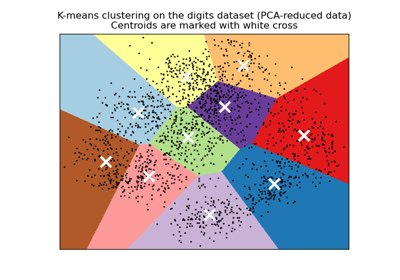
A demo of K-Means clustering on the handwritten digits data