pairwise_distances_argmin#
- sklearn.metrics.pairwise_distances_argmin(X, Y, *, axis=1, metric='euclidean', metric_kwargs=None)[source]#
Compute minimum distances between one point and a set of points.
This function computes for each row in X, the index of the row of Y which is closest (according to the specified distance).
This is mostly equivalent to calling:
pairwise_distances(X, Y=Y, metric=metric).argmin(axis=axis)
but uses much less memory, and is faster for large arrays.
This function works with dense 2D arrays only.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples_X, n_features)
Array containing points.
- Y{array-like, sparse matrix} of shape (n_samples_Y, n_features)
Arrays containing points.
- axisint, default=1
Axis along which the argmin and distances are to be computed.
- metricstr or callable, default=”euclidean”
Metric to use for distance computation. Any metric from scikit-learn or
scipy.spatial.distancecan be used.If metric is a callable function, it is called on each pair of instances (rows) and the resulting value recorded. The callable should take two arrays as input and return one value indicating the distance between them. This works for Scipy’s metrics, but is less efficient than passing the metric name as a string.
Distance matrices are not supported.
Valid values for metric are:
from scikit-learn: [‘cityblock’, ‘cosine’, ‘euclidean’, ‘l1’, ‘l2’, ‘manhattan’, ‘nan_euclidean’]
from
scipy.spatial.distance: [‘braycurtis’, ‘canberra’, ‘chebyshev’, ‘correlation’, ‘dice’, ‘hamming’, ‘jaccard’, ‘kulsinski’, ‘mahalanobis’, ‘minkowski’, ‘rogerstanimoto’, ‘russellrao’, ‘seuclidean’, ‘sokalmichener’, ‘sokalsneath’, ‘sqeuclidean’, ‘yule’]
See the documentation for
scipy.spatial.distancefor details on these metrics.Note
'kulsinski'is deprecated from SciPy 1.9 and will be removed in SciPy 1.11.Note
'matching'has been removed in SciPy 1.9 (use'hamming'instead).- metric_kwargsdict, default=None
Keyword arguments to pass to specified metric function.
- Returns:
- argminnumpy.ndarray
Y[argmin[i], :] is the row in Y that is closest to X[i, :].
See also
pairwise_distancesDistances between every pair of samples of X and Y.
pairwise_distances_argmin_minSame as
pairwise_distances_argminbut also returns the distances.
Examples
>>> from sklearn.metrics.pairwise import pairwise_distances_argmin >>> X = [[0, 0, 0], [1, 1, 1]] >>> Y = [[1, 0, 0], [1, 1, 0]] >>> pairwise_distances_argmin(X, Y) array([0, 1])
Gallery examples#
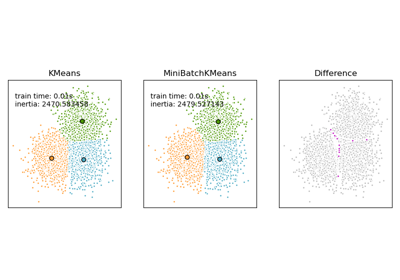
Comparison of the K-Means and MiniBatchKMeans clustering algorithms
