OneVsRestClassifier#
- class sklearn.multiclass.OneVsRestClassifier(estimator, *, n_jobs=None, verbose=0)[source]#
One-vs-the-rest (OvR) multiclass strategy.
Also known as one-vs-all, this strategy consists in fitting one classifier per class. For each classifier, the class is fitted against all the other classes. In addition to its computational efficiency (only
n_classesclassifiers are needed), one advantage of this approach is its interpretability. Since each class is represented by one and one classifier only, it is possible to gain knowledge about the class by inspecting its corresponding classifier. This is the most commonly used strategy for multiclass classification and is a fair default choice.OneVsRestClassifier can also be used for multilabel classification. To use this feature, provide an indicator matrix for the target
ywhen calling.fit. In other words, the target labels should be formatted as a 2D binary (0/1) matrix, where [i, j] == 1 indicates the presence of label j in sample i. This estimator uses the binary relevance method to perform multilabel classification, which involves training one binary classifier independently for each label.Read more in the User Guide.
- Parameters:
- estimatorestimator object
A regressor or a classifier that implements fit. When a classifier is passed, decision_function will be used in priority and it will fallback to predict_proba if it is not available. When a regressor is passed, predict is used.
- n_jobsint, default=None
The number of jobs to use for the computation: the
n_classesone-vs-rest problems are computed in parallel.Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.Changed in version 0.20:
n_jobsdefault changed from 1 to None- verboseint, default=0
The verbosity level, if non zero, progress messages are printed. Below 50, the output is sent to stderr. Otherwise, the output is sent to stdout. The frequency of the messages increases with the verbosity level, reporting all iterations at 10. See
joblib.Parallelfor more details.Added in version 1.1.
- Attributes:
- estimators_list of
n_classesestimators Estimators used for predictions.
- classes_array, shape = [
n_classes] Class labels.
n_classes_intNumber of classes.
- label_binarizer_LabelBinarizer object
Object used to transform multiclass labels to binary labels and vice-versa.
multilabel_booleanWhether this is a multilabel classifier.
- n_features_in_int
Number of features seen during fit. Only defined if the underlying estimator exposes such an attribute when fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Only defined if the underlying estimator exposes such an attribute when fit.
Added in version 1.0.
- estimators_list of
See also
OneVsOneClassifierOne-vs-one multiclass strategy.
OutputCodeClassifier(Error-Correcting) Output-Code multiclass strategy.
sklearn.multioutput.MultiOutputClassifierAlternate way of extending an estimator for multilabel classification.
sklearn.preprocessing.MultiLabelBinarizerTransform iterable of iterables to binary indicator matrix.
Examples
>>> import numpy as np >>> from sklearn.multiclass import OneVsRestClassifier >>> from sklearn.svm import SVC >>> X = np.array([ ... [10, 10], ... [8, 10], ... [-5, 5.5], ... [-5.4, 5.5], ... [-20, -20], ... [-15, -20] ... ]) >>> y = np.array([0, 0, 1, 1, 2, 2]) >>> clf = OneVsRestClassifier(SVC()).fit(X, y) >>> clf.predict([[-19, -20], [9, 9], [-5, 5]]) array([2, 0, 1])
- decision_function(X)[source]#
Decision function for the OneVsRestClassifier.
Return the distance of each sample from the decision boundary for each class. This can only be used with estimators which implement the
decision_functionmethod.- Parameters:
- Xarray-like of shape (n_samples, n_features)
Input data.
- Returns:
- Tarray-like of shape (n_samples, n_classes) or (n_samples,) for binary classification.
Result of calling
decision_functionon the final estimator.Changed in version 0.19: output shape changed to
(n_samples,)to conform to scikit-learn conventions for binary classification.
- fit(X, y, **fit_params)[source]#
Fit underlying estimators.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Data.
- y{array-like, sparse matrix} of shape (n_samples,) or (n_samples, n_classes)
Multi-class targets. An indicator matrix turns on multilabel classification.
- **fit_paramsdict
Parameters passed to the
estimator.fitmethod of each sub-estimator.Added in version 1.4: Only available if
enable_metadata_routing=True. See Metadata Routing User Guide for more details.
- Returns:
- selfobject
Instance of fitted estimator.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
Added in version 1.4.
- Returns:
- routingMetadataRouter
A
MetadataRouterencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- partial_fit(X, y, classes=None, **partial_fit_params)[source]#
Partially fit underlying estimators.
Should be used when memory is inefficient to train all data. Chunks of data can be passed in several iterations.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Data.
- y{array-like, sparse matrix} of shape (n_samples,) or (n_samples, n_classes)
Multi-class targets. An indicator matrix turns on multilabel classification.
- classesarray, shape (n_classes, )
Classes across all calls to partial_fit. Can be obtained via
np.unique(y_all), where y_all is the target vector of the entire dataset. This argument is only required in the first call of partial_fit and can be omitted in the subsequent calls.- **partial_fit_paramsdict
Parameters passed to the
estimator.partial_fitmethod of each sub-estimator.Added in version 1.4: Only available if
enable_metadata_routing=True. See Metadata Routing User Guide for more details.
- Returns:
- selfobject
Instance of partially fitted estimator.
- predict(X)[source]#
Predict multi-class targets using underlying estimators.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Data.
- Returns:
- y{array-like, sparse matrix} of shape (n_samples,) or (n_samples, n_classes)
Predicted multi-class targets.
- predict_proba(X)[source]#
Probability estimates.
The returned estimates for all classes are ordered by label of classes.
Note that in the multilabel case, each sample can have any number of labels. This returns the marginal probability that the given sample has the label in question. For example, it is entirely consistent that two labels both have a 90% probability of applying to a given sample.
In the single label multiclass case, the rows of the returned matrix sum to 1.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Input data.
- Returns:
- Tarray-like of shape (n_samples, n_classes)
Returns the probability of the sample for each class in the model, where classes are ordered as they are in
self.classes_.
- score(X, y, sample_weight=None)[source]#
Return accuracy on provided data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True labels for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
Mean accuracy of
self.predict(X)w.r.t.y.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_partial_fit_request(*, classes: bool | None | str = '$UNCHANGED$') OneVsRestClassifier[source]#
Configure whether metadata should be requested to be passed to the
partial_fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed topartial_fitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it topartial_fit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- classesstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
classesparameter inpartial_fit.
- Returns:
- selfobject
The updated object.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') OneVsRestClassifier[source]#
Configure whether metadata should be requested to be passed to the
scoremethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.
Gallery examples#
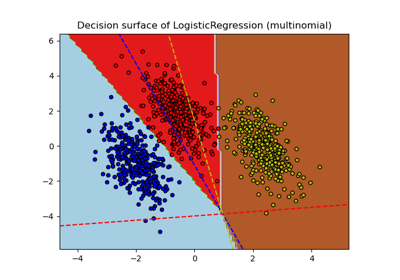
Decision Boundaries of Multinomial and One-vs-Rest Logistic Regression
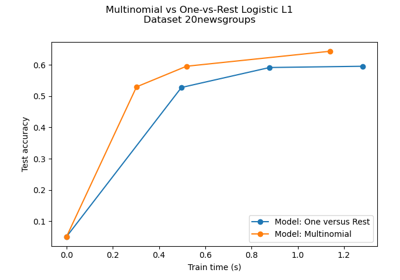
Multiclass sparse logistic regression on 20newgroups
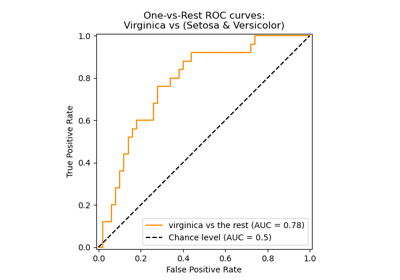
Multiclass Receiver Operating Characteristic (ROC)
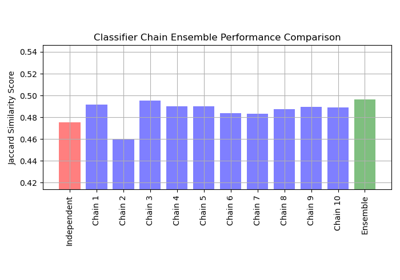
Multilabel classification using a classifier chain