DBSCAN#
- class sklearn.cluster.DBSCAN(eps=0.5, *, min_samples=5, metric='euclidean', metric_params=None, algorithm='auto', leaf_size=30, p=None, n_jobs=None)[source]#
Perform DBSCAN clustering from vector array or distance matrix.
DBSCAN - Density-Based Spatial Clustering of Applications with Noise. Finds core samples of high density and expands clusters from them. This algorithm is particularly good for data which contains clusters of similar density and can find clusters of arbitrary shape.
Unlike K-means, DBSCAN does not require specifying the number of clusters in advance and can identify outliers as noise points.
This implementation has a worst case memory complexity of \(O({n}^2)\), which can occur when the
epsparam is large andmin_samplesis low, while the original DBSCAN only uses linear memory. For further details, see the Notes below.Read more in the User Guide.
- Parameters:
- epsfloat, default=0.5
The maximum distance between two samples for one to be considered as in the neighborhood of the other. This is not a maximum bound on the distances of points within a cluster. This is the most important DBSCAN parameter to choose appropriately for your data set and distance function. Smaller values generally lead to more clusters.
- min_samplesint, default=5
The number of samples (or total weight) in a neighborhood for a point to be considered as a core point. This includes the point itself. If
min_samplesis set to a higher value, DBSCAN will find denser clusters, whereas if it is set to a lower value, the found clusters will be more sparse.- metricstr, or callable, default=’euclidean’
The metric to use when calculating distance between instances in a feature array. If metric is a string or callable, it must be one of the options allowed by
sklearn.metrics.pairwise_distancesfor its metric parameter. If metric is “precomputed”, X is assumed to be a distance matrix and must be square. X may be a sparse graph, in which case only “nonzero” elements may be considered neighbors for DBSCAN.Added in version 0.17: metric precomputed to accept precomputed sparse matrix.
- metric_paramsdict, default=None
Additional keyword arguments for the metric function.
Added in version 0.19.
- algorithm{‘auto’, ‘ball_tree’, ‘kd_tree’, ‘brute’}, default=’auto’
The algorithm to be used by the NearestNeighbors module to compute pointwise distances and find nearest neighbors. ‘auto’ will attempt to decide the most appropriate algorithm based on the values passed to
fitmethod. SeeNearestNeighborsdocumentation for details.- leaf_sizeint, default=30
Leaf size passed to BallTree or cKDTree. This can affect the speed of the construction and query, as well as the memory required to store the tree. The optimal value depends on the nature of the problem.
- pfloat, default=None
The power of the Minkowski metric to be used to calculate distance between points. If None, then
p=2(equivalent to the Euclidean distance). When p=1, this is equivalent to Manhattan distance.- n_jobsint, default=None
The number of parallel jobs to run.
Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.
- Attributes:
- core_sample_indices_ndarray of shape (n_core_samples,)
Indices of core samples.
- components_ndarray of shape (n_core_samples, n_features)
Copy of each core sample found by training.
- labels_ndarray of shape (n_samples,)
Cluster labels for each point in the dataset given to fit(). Noisy samples are given the label -1. Non-negative integers indicate cluster membership.
- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
See also
OPTICSA similar clustering at multiple values of eps. Our implementation is optimized for memory usage.
Notes
This implementation bulk-computes all neighborhood queries, which increases the memory complexity to O(n.d) where d is the average number of neighbors, while original DBSCAN had memory complexity O(n). It may attract a higher memory complexity when querying these nearest neighborhoods, depending on the
algorithm.One way to avoid the query complexity is to pre-compute sparse neighborhoods in chunks using
NearestNeighbors.radius_neighbors_graphwithmode='distance', then usingmetric='precomputed'here.Another way to reduce memory and computation time is to remove (near-)duplicate points and use
sample_weightinstead.OPTICSprovides a similar clustering with lower memory usage.References
Ester, M., H. P. Kriegel, J. Sander, and X. Xu, “A Density-Based Algorithm for Discovering Clusters in Large Spatial Databases with Noise”. In: Proceedings of the 2nd International Conference on Knowledge Discovery and Data Mining, Portland, OR, AAAI Press, pp. 226-231. 1996
Schubert, E., Sander, J., Ester, M., Kriegel, H. P., & Xu, X. (2017). “DBSCAN revisited, revisited: why and how you should (still) use DBSCAN.” ACM Transactions on Database Systems (TODS), 42(3), 19.
Examples
>>> from sklearn.cluster import DBSCAN >>> import numpy as np >>> X = np.array([[1, 2], [2, 2], [2, 3], ... [8, 7], [8, 8], [25, 80]]) >>> clustering = DBSCAN(eps=3, min_samples=2).fit(X) >>> clustering.labels_ array([ 0, 0, 0, 1, 1, -1]) >>> clustering DBSCAN(eps=3, min_samples=2)
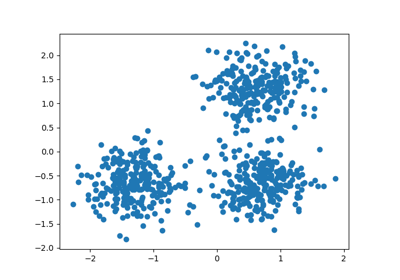
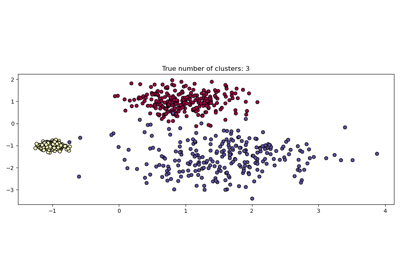
For an example, see Demo of DBSCAN clustering algorithm.
For a comparison of DBSCAN with other clustering algorithms, see Comparing different clustering algorithms on toy datasets
- fit(X, y=None, sample_weight=None)[source]#
Perform DBSCAN clustering from features, or distance matrix.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features), or (n_samples, n_samples)
Training instances to cluster, or distances between instances if
metric='precomputed'. If a sparse matrix is provided, it will be converted into a sparsecsr_matrix.- yIgnored
Not used, present here for API consistency by convention.
- sample_weightarray-like of shape (n_samples,), default=None
Weight of each sample, such that a sample with a weight of at least
min_samplesis by itself a core sample; a sample with a negative weight may inhibit its eps-neighbor from being core. Note that weights are absolute, and default to 1.
- Returns:
- selfobject
Returns a fitted instance of self.
- fit_predict(X, y=None, sample_weight=None)[source]#
Compute clusters from a data or distance matrix and predict labels.
This method fits the model and returns the cluster labels in a single step. It is equivalent to calling fit(X).labels_.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features), or (n_samples, n_samples)
Training instances to cluster, or distances between instances if
metric='precomputed'. If a sparse matrix is provided, it will be converted into a sparsecsr_matrix.- yIgnored
Not used, present here for API consistency by convention.
- sample_weightarray-like of shape (n_samples,), default=None
Weight of each sample, such that a sample with a weight of at least
min_samplesis by itself a core sample; a sample with a negative weight may inhibit its eps-neighbor from being core. Note that weights are absolute, and default to 1.
- Returns:
- labelsndarray of shape (n_samples,)
Cluster labels. Noisy samples are given the label -1. Non-negative integers indicate cluster membership.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- set_fit_request(*, sample_weight: bool | None | str = '$UNCHANGED$') DBSCAN[source]#
Configure whether metadata should be requested to be passed to the
fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter infit.
- Returns:
- selfobject
The updated object.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
Gallery examples#
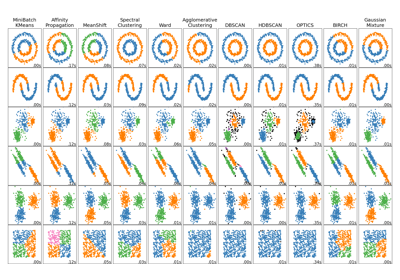
Comparing different clustering algorithms on toy datasets