TimeSeriesSplit#
- class sklearn.model_selection.TimeSeriesSplit(n_splits=5, *, max_train_size=None, test_size=None, gap=0)[source]#
Time Series cross-validator.
Provides train/test indices to split time-ordered data, where other cross-validation methods are inappropriate, as they would lead to training on future data and evaluating on past data. To ensure comparable metrics across folds, samples must be equally spaced. Once this condition is met, each test set covers the same time duration, while the train set size accumulates data from previous splits.
This cross-validation object is a variation of
KFold. In the k-th split, it returns the first k folds as the train set and the (k+1)-th fold as the test set.Note that, unlike standard cross-validation methods, successive training sets are supersets of those that come before them.
Read more in the User Guide.
For visualisation of cross-validation behaviour and comparison between common scikit-learn split methods refer to Visualizing cross-validation behavior in scikit-learn
Added in version 0.18.
- Parameters:
- n_splitsint, default=5
Number of splits. Must be at least 2.
Changed in version 0.22:
n_splitsdefault value changed from 3 to 5.- max_train_sizeint, default=None
Maximum size for a single training set.
- test_sizeint, default=None
Used to limit the size of the test set. Defaults to
n_samples // (n_splits + 1), which is the maximum allowed value withgap=0.Added in version 0.24.
- gapint, default=0
Number of samples to exclude from the end of each train set before the test set.
Added in version 0.24.
Notes
The training set has size
i * n_samples // (n_splits + 1) + n_samples % (n_splits + 1)in theith split, with a test set of sizen_samples//(n_splits + 1)by default, wheren_samplesis the number of samples. Note that this formula is only valid whentest_sizeandmax_train_sizeare left to their default values.Examples
>>> import numpy as np >>> from sklearn.model_selection import TimeSeriesSplit >>> X = np.array([[1, 2], [3, 4], [1, 2], [3, 4], [1, 2], [3, 4]]) >>> y = np.array([1, 2, 3, 4, 5, 6]) >>> tscv = TimeSeriesSplit() >>> print(tscv) TimeSeriesSplit(gap=0, max_train_size=None, n_splits=5, test_size=None) >>> for i, (train_index, test_index) in enumerate(tscv.split(X)): ... print(f"Fold {i}:") ... print(f" Train: index={train_index}") ... print(f" Test: index={test_index}") Fold 0: Train: index=[0] Test: index=[1] Fold 1: Train: index=[0 1] Test: index=[2] Fold 2: Train: index=[0 1 2] Test: index=[3] Fold 3: Train: index=[0 1 2 3] Test: index=[4] Fold 4: Train: index=[0 1 2 3 4] Test: index=[5] >>> # Fix test_size to 2 with 12 samples >>> X = np.random.randn(12, 2) >>> y = np.random.randint(0, 2, 12) >>> tscv = TimeSeriesSplit(n_splits=3, test_size=2) >>> for i, (train_index, test_index) in enumerate(tscv.split(X)): ... print(f"Fold {i}:") ... print(f" Train: index={train_index}") ... print(f" Test: index={test_index}") Fold 0: Train: index=[0 1 2 3 4 5] Test: index=[6 7] Fold 1: Train: index=[0 1 2 3 4 5 6 7] Test: index=[8 9] Fold 2: Train: index=[0 1 2 3 4 5 6 7 8 9] Test: index=[10 11] >>> # Add in a 2 period gap >>> tscv = TimeSeriesSplit(n_splits=3, test_size=2, gap=2) >>> for i, (train_index, test_index) in enumerate(tscv.split(X)): ... print(f"Fold {i}:") ... print(f" Train: index={train_index}") ... print(f" Test: index={test_index}") Fold 0: Train: index=[0 1 2 3] Test: index=[6 7] Fold 1: Train: index=[0 1 2 3 4 5] Test: index=[8 9] Fold 2: Train: index=[0 1 2 3 4 5 6 7] Test: index=[10 11]
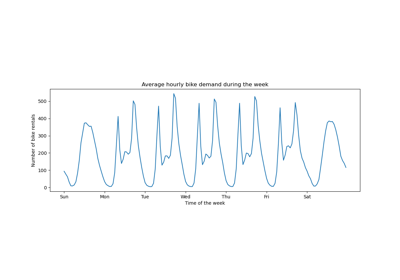
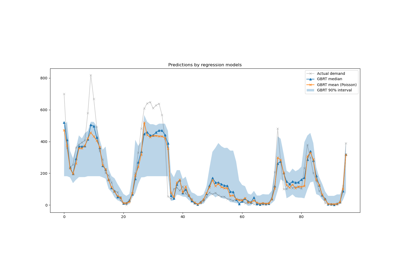
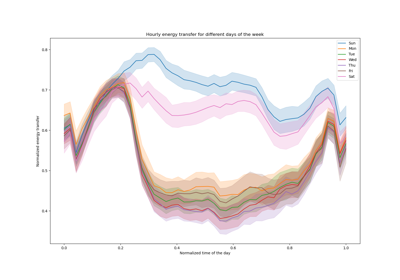
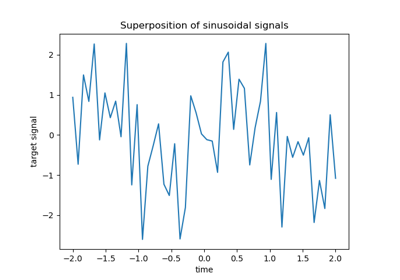
For a more extended example see Time-related feature engineering.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_n_splits(X=None, y=None, groups=None)[source]#
Returns the number of splitting iterations as set with the
n_splitsparam when instantiating the cross-validator.- Parameters:
- Xarray-like of shape (n_samples, n_features), default=None
Always ignored, exists for API compatibility.
- yarray-like of shape (n_samples,), default=None
Always ignored, exists for API compatibility.
- groupsarray-like of shape (n_samples,), default=None
Always ignored, exists for API compatibility.
- Returns:
- n_splitsint
Returns the number of splitting iterations in the cross-validator.
- split(X, y=None, groups=None)[source]#
Generate indices to split data into training and test set.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Training data, where
n_samplesis the number of samples andn_featuresis the number of features.- yarray-like of shape (n_samples,), default=None
Always ignored, exists for API compatibility.
- groupsarray-like of shape (n_samples,), default=None
Always ignored, exists for API compatibility.
- Yields:
- trainndarray
The training set indices for that split.
- testndarray
The testing set indices for that split.
Gallery examples#
Visualizing cross-validation behavior in scikit-learn