GaussianProcessRegressor#
- class sklearn.gaussian_process.GaussianProcessRegressor(kernel=None, *, alpha=1e-10, optimizer='fmin_l_bfgs_b', n_restarts_optimizer=0, normalize_y=False, copy_X_train=True, n_targets=None, random_state=None)[source]#
Gaussian process regression (GPR).
The implementation is based on Algorithm 2.1 of [RW2006].
In addition to standard scikit-learn estimator API,
GaussianProcessRegressor:allows prediction without prior fitting (based on the GP prior)
provides an additional method
sample_y(X), which evaluates samples drawn from the GPR (prior or posterior) at given inputsexposes a method
log_marginal_likelihood(theta), which can be used externally for other ways of selecting hyperparameters, e.g., via Markov chain Monte Carlo.
To learn the difference between a point-estimate approach vs. a more Bayesian modelling approach, refer to the example entitled Comparison of kernel ridge and Gaussian process regression.
Read more in the User Guide.
Added in version 0.18.
- Parameters:
- kernelkernel instance, default=None
The kernel specifying the covariance function of the GP. If
Noneis passed, the kernelConstantKernel() * RBF()is used as default. Note that the kernel hyperparameters are optimized during fitting unless the bounds are marked as"fixed"or the argumentoptimizeris set toNone.- alphafloat or ndarray of shape (n_samples,), default=1e-10
Value added to the diagonal of the kernel matrix during fitting. This can prevent a potential numerical issue during fitting, by ensuring that the calculated values form a positive definite matrix. It can also be interpreted as the variance of additional Gaussian measurement noise on the training observations. Note that this is different from using a
WhiteKernel. If an array is passed, it must have the same number of entries as the data used for fitting and is used as datapoint-dependent noise level. Allowing to specify the noise level directly as a parameter is mainly for convenience and for consistency withRidge. For an example illustrating how the alpha parameter controls the noise variance in Gaussian Process Regression, see Gaussian Processes regression: basic introductory example.- optimizer“fmin_l_bfgs_b”, callable or None, default=”fmin_l_bfgs_b”
Can either be one of the internally supported optimizers for optimizing the kernel’s parameters, specified by a string, or an externally defined optimizer passed as a callable. If a callable is passed, it must have the signature:
def optimizer(obj_func, initial_theta, bounds): # * 'obj_func': the objective function to be minimized, which # takes the hyperparameters theta as a parameter and an # optional flag eval_gradient, which determines if the # gradient is returned additionally to the function value # * 'initial_theta': the initial value for theta, which can be # used by local optimizers # * 'bounds': the bounds on the values of theta .... # Returned are the best found hyperparameters theta and # the corresponding value of the target function. return theta_opt, func_min
Per default, the L-BFGS-B algorithm from
scipy.optimize.minimizeis used. If None is passed, the kernel’s parameters are kept fixed. Available internal optimizers are:{'fmin_l_bfgs_b'}.- n_restarts_optimizerint, default=0
The number of restarts of the optimizer for finding the kernel’s parameters which maximize the log-marginal likelihood. The first run of the optimizer is performed from the kernel’s initial parameters, the remaining ones (if any) from thetas sampled log-uniform randomly from the space of allowed theta-values. If greater than 0, all bounds must be finite. Note that
n_restarts_optimizer == 0implies that one run is performed.- normalize_ybool, default=False
Whether or not to normalize the target values
yby removing the mean and scaling to unit-variance. This is recommended for cases where zero-mean, unit-variance priors are used. Note that, in this implementation, the normalisation is reversed before the GP predictions are reported.Changed in version 0.23.
- copy_X_trainbool, default=True
If True, a persistent copy of the training data is stored in the object. Otherwise, just a reference to the training data is stored, which might cause predictions to change if the data is modified externally.
- n_targetsint, default=None
The number of dimensions of the target values. Used to decide the number of outputs when sampling from the prior distributions (i.e. calling
sample_ybeforefit). This parameter is ignored oncefithas been called.Added in version 1.3.
- random_stateint, RandomState instance or None, default=None
Determines random number generation used to initialize the centers. Pass an int for reproducible results across multiple function calls. See Glossary.
- Attributes:
- X_train_array-like of shape (n_samples, n_features) or list of object
Feature vectors or other representations of training data (also required for prediction).
- y_train_array-like of shape (n_samples,) or (n_samples, n_targets)
Target values in training data (also required for prediction).
- kernel_kernel instance
The kernel used for prediction. The structure of the kernel is the same as the one passed as parameter but with optimized hyperparameters.
- L_array-like of shape (n_samples, n_samples)
Lower-triangular Cholesky decomposition of the kernel in
X_train_.- alpha_array-like of shape (n_samples,)
Dual coefficients of training data points in kernel space.
- log_marginal_likelihood_value_float
The log-marginal-likelihood of
self.kernel_.theta.- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
See also
GaussianProcessClassifierGaussian process classification (GPC) based on Laplace approximation.
References
Examples
>>> from sklearn.datasets import make_friedman2 >>> from sklearn.gaussian_process import GaussianProcessRegressor >>> from sklearn.gaussian_process.kernels import DotProduct, WhiteKernel >>> X, y = make_friedman2(n_samples=500, noise=0, random_state=0) >>> kernel = DotProduct() + WhiteKernel() >>> gpr = GaussianProcessRegressor(kernel=kernel, ... random_state=0).fit(X, y) >>> gpr.score(X, y) 0.3680... >>> gpr.predict(X[:2,:], return_std=True) (array([653.0, 592.1]), array([316.6, 316.6]))
- fit(X, y)[source]#
Fit Gaussian process regression model.
- Parameters:
- Xarray-like of shape (n_samples, n_features) or list of object
Feature vectors or other representations of training data.
- yarray-like of shape (n_samples,) or (n_samples, n_targets)
Target values.
- Returns:
- selfobject
GaussianProcessRegressor class instance.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- log_marginal_likelihood(theta=None, eval_gradient=False, clone_kernel=True)[source]#
Return log-marginal likelihood of theta for training data.
- Parameters:
- thetaarray-like of shape (n_kernel_params,) default=None
Kernel hyperparameters for which the log-marginal likelihood is evaluated. If None, the precomputed log_marginal_likelihood of
self.kernel_.thetais returned.- eval_gradientbool, default=False
If True, the gradient of the log-marginal likelihood with respect to the kernel hyperparameters at position theta is returned additionally. If True, theta must not be None.
- clone_kernelbool, default=True
If True, the kernel attribute is copied. If False, the kernel attribute is modified, but may result in a performance improvement.
- Returns:
- log_likelihoodfloat
Log-marginal likelihood of theta for training data.
- log_likelihood_gradientndarray of shape (n_kernel_params,), optional
Gradient of the log-marginal likelihood with respect to the kernel hyperparameters at position theta. Only returned when eval_gradient is True.
- predict(X, return_std=False, return_cov=False)[source]#
Predict using the Gaussian process regression model.
We can also predict based on an unfitted model by using the GP prior. In addition to the mean of the predictive distribution, optionally also returns its standard deviation (
return_std=True) or covariance (return_cov=True). Note that at most one of the two can be requested.- Parameters:
- Xarray-like of shape (n_samples, n_features) or list of object
Query points where the GP is evaluated.
- return_stdbool, default=False
If True, the standard-deviation of the predictive distribution at the query points is returned along with the mean.
- return_covbool, default=False
If True, the covariance of the joint predictive distribution at the query points is returned along with the mean.
- Returns:
- y_meanndarray of shape (n_samples,) or (n_samples, n_targets)
Mean of predictive distribution at query points.
- y_stdndarray of shape (n_samples,) or (n_samples, n_targets), optional
Standard deviation of predictive distribution at query points. Only returned when
return_stdis True.- y_covndarray of shape (n_samples, n_samples) or (n_samples, n_samples, n_targets), optional
Covariance of joint predictive distribution at query points. Only returned when
return_covis True.
- sample_y(X, n_samples=1, random_state=0)[source]#
Draw samples from Gaussian process and evaluate at X.
- Parameters:
- Xarray-like of shape (n_samples_X, n_features) or list of object
Query points where the GP is evaluated.
- n_samplesint, default=1
Number of samples drawn from the Gaussian process per query point.
- random_stateint, RandomState instance or None, default=0
Determines random number generation to randomly draw samples. Pass an int for reproducible results across multiple function calls. See Glossary.
- Returns:
- y_samplesndarray of shape (n_samples_X, n_samples), or (n_samples_X, n_targets, n_samples)
Values of n_samples samples drawn from Gaussian process and evaluated at query points.
- score(X, y, sample_weight=None)[source]#
Return coefficient of determination on test data.
The coefficient of determination, \(R^2\), is defined as \((1 - \frac{u}{v})\), where \(u\) is the residual sum of squares
((y_true - y_pred)** 2).sum()and \(v\) is the total sum of squares((y_true - y_true.mean()) ** 2).sum(). The best possible score is 1.0 and it can be negative (because the model can be arbitrarily worse). A constant model that always predicts the expected value ofy, disregarding the input features, would get a \(R^2\) score of 0.0.- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples. For some estimators this may be a precomputed kernel matrix or a list of generic objects instead with shape
(n_samples, n_samples_fitted), wheren_samples_fittedis the number of samples used in the fitting for the estimator.- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True values for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
\(R^2\) of
self.predict(X)w.r.t.y.
Notes
The \(R^2\) score used when calling
scoreon a regressor usesmultioutput='uniform_average'from version 0.23 to keep consistent with default value ofr2_score. This influences thescoremethod of all the multioutput regressors (except forMultiOutputRegressor).
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_predict_request(*, return_cov: bool | None | str = '$UNCHANGED$', return_std: bool | None | str = '$UNCHANGED$') GaussianProcessRegressor[source]#
Configure whether metadata should be requested to be passed to the
predictmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed topredictif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it topredict.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- return_covstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
return_covparameter inpredict.- return_stdstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
return_stdparameter inpredict.
- Returns:
- selfobject
The updated object.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') GaussianProcessRegressor[source]#
Configure whether metadata should be requested to be passed to the
scoremethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.
Gallery examples#
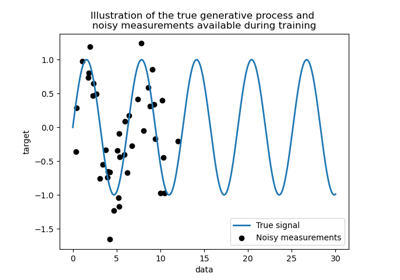
Comparison of kernel ridge and Gaussian process regression
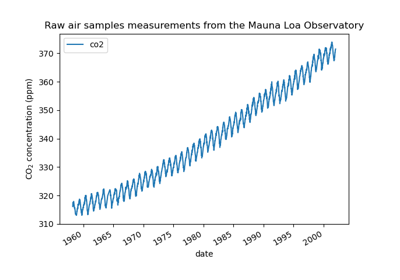
Forecasting of CO2 level on Mona Loa dataset using Gaussian process regression (GPR)
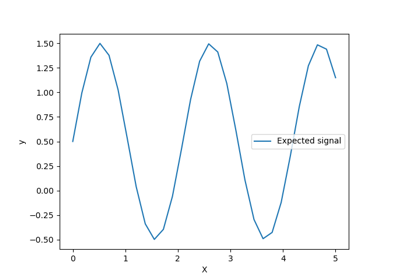
Ability of Gaussian process regression (GPR) to estimate data noise-level
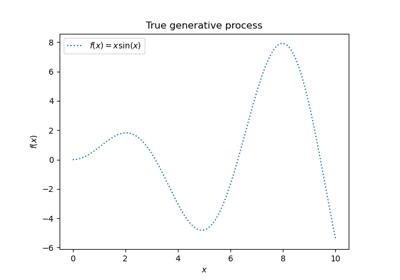
Gaussian Processes regression: basic introductory example
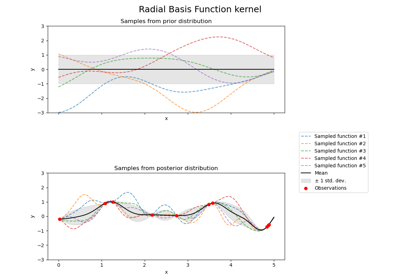
Illustration of prior and posterior Gaussian process for different kernels