SGDRegressor#
- class sklearn.linear_model.SGDRegressor(loss='squared_error', *, penalty='l2', alpha=0.0001, l1_ratio=0.15, fit_intercept=True, max_iter=1000, tol=0.001, shuffle=True, verbose=0, epsilon=0.1, random_state=None, learning_rate='invscaling', eta0=0.01, power_t=0.25, early_stopping=False, validation_fraction=0.1, n_iter_no_change=5, warm_start=False, average=False)[source]#
Linear model fitted by minimizing a regularized empirical loss with SGD.
SGD stands for Stochastic Gradient Descent: the gradient of the loss is estimated each sample at a time and the model is updated along the way with a decreasing strength schedule (aka learning rate).
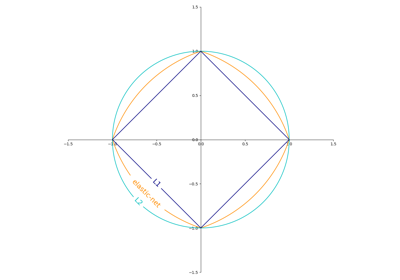
The regularizer is a penalty added to the loss function that shrinks model parameters towards the zero vector using either the squared euclidean norm L2 or the absolute norm L1 or a combination of both (Elastic Net). If the parameter update crosses the 0.0 value because of the regularizer, the update is truncated to 0.0 to allow for learning sparse models and achieve online feature selection.
This implementation works with data represented as dense numpy arrays of floating point values for the features.
Read more in the User Guide.
- Parameters:
- lossstr, default=’squared_error’
The loss function to be used. The possible values are ‘squared_error’, ‘huber’, ‘epsilon_insensitive’, or ‘squared_epsilon_insensitive’
The ‘squared_error’ refers to the ordinary least squares fit. ‘huber’ modifies ‘squared_error’ to focus less on getting outliers correct by switching from squared to linear loss past a distance of epsilon. ‘epsilon_insensitive’ ignores errors less than epsilon and is linear past that; this is the loss function used in SVR. ‘squared_epsilon_insensitive’ is the same but becomes squared loss past a tolerance of epsilon.
More details about the losses formulas can be found in the User Guide.
- penalty{‘l2’, ‘l1’, ‘elasticnet’, None}, default=’l2’
The penalty (aka regularization term) to be used. Defaults to ‘l2’ which is the standard regularizer for linear SVM models. ‘l1’ and ‘elasticnet’ might bring sparsity to the model (feature selection) not achievable with ‘l2’. No penalty is added when set to
None.You can see a visualisation of the penalties in SGD: Penalties.
- alphafloat, default=0.0001
Constant that multiplies the regularization term. The higher the value, the stronger the regularization. Also used to compute the learning rate when
learning_rateis set to ‘optimal’. Values must be in the range[0.0, inf).- l1_ratiofloat, default=0.15
The Elastic Net mixing parameter, with 0 <= l1_ratio <= 1. l1_ratio=0 corresponds to L2 penalty, l1_ratio=1 to L1. Only used if
penaltyis ‘elasticnet’. Values must be in the range[0.0, 1.0]or can beNoneifpenaltyis notelasticnet.Changed in version 1.7:
l1_ratiocan beNonewhenpenaltyis not “elasticnet”.- fit_interceptbool, default=True
Whether the intercept should be estimated or not. If False, the data is assumed to be already centered.
- max_iterint, default=1000
The maximum number of passes over the training data (aka epochs). It only impacts the behavior in the
fitmethod, and not thepartial_fitmethod. Values must be in the range[1, inf).Added in version 0.19.
- tolfloat or None, default=1e-3
The stopping criterion. If it is not None, training will stop when (loss > best_loss - tol) for
n_iter_no_changeconsecutive epochs. Convergence is checked against the training loss or the validation loss depending on theearly_stoppingparameter. Values must be in the range[0.0, inf).Added in version 0.19.
- shufflebool, default=True
Whether or not the training data should be shuffled after each epoch.
- verboseint, default=0
The verbosity level. Values must be in the range
[0, inf).- epsilonfloat, default=0.1
Epsilon in the epsilon-insensitive loss functions; only if
lossis ‘huber’, ‘epsilon_insensitive’, or ‘squared_epsilon_insensitive’. For ‘huber’, determines the threshold at which it becomes less important to get the prediction exactly right. For epsilon-insensitive, any differences between the current prediction and the correct label are ignored if they are less than this threshold. Values must be in the range[0.0, inf).- random_stateint, RandomState instance, default=None
Used for shuffling the data, when
shuffleis set toTrue. Pass an int for reproducible output across multiple function calls. See Glossary.- learning_ratestr, default=’invscaling’
The learning rate schedule:
‘constant’:
eta = eta0‘optimal’:
eta = 1.0 / (alpha * (t + t0))where t0 is chosen by a heuristic proposed by Leon Bottou.‘invscaling’:
eta = eta0 / pow(t, power_t)‘adaptive’: eta = eta0, as long as the training keeps decreasing. Each time n_iter_no_change consecutive epochs fail to decrease the training loss by tol or fail to increase validation score by tol if early_stopping is True, the current learning rate is divided by 5.
‘pa1’: passive-aggressive algorithm 1, see [1]. Only with
loss='epsilon_insensitive'. Update isw += eta y xwitheta = min(eta0, loss/||x||**2).‘pa2’: passive-aggressive algorithm 2, see [1]. Only with
loss='epsilon_insensitive'. Update isw += eta y xwitheta = hinge_loss / (||x||**2 + 1/(2 eta0)).
Added in version 0.20: Added ‘adaptive’ option.
Added in version 1.8: Added options ‘pa1’ and ‘pa2’
- eta0float, default=0.01
The initial learning rate for the ‘constant’, ‘invscaling’ or ‘adaptive’ schedules. The default value is 0.01. Values must be in the range
(0.0, inf).For PA-1 (
learning_rate=pa1) and PA-II (pa2), it specifies the aggressiveness parameter for the passive-agressive algorithm, see [1] where it is called C:For PA-I it is the maximum step size.
For PA-II it regularizes the step size (the smaller
eta0the more it regularizes).
As a general rule-of-thumb for PA,
eta0should be small when the data is noisy.- power_tfloat, default=0.25
The exponent for inverse scaling learning rate. Values must be in the range
[0.0, inf).Deprecated since version 1.8: Negative values for
power_tare deprecated in version 1.8 and will raise an error in 1.10. Use values in the range [0.0, inf) instead.- early_stoppingbool, default=False
Whether to use early stopping to terminate training when validation score is not improving. If set to True, it will automatically set aside a fraction of training data as validation and terminate training when validation score returned by the
scoremethod is not improving by at leasttolforn_iter_no_changeconsecutive epochs.See Early stopping of Stochastic Gradient Descent for an example of the effects of early stopping.
Added in version 0.20: Added ‘early_stopping’ option
- validation_fractionfloat, default=0.1
The proportion of training data to set aside as validation set for early stopping. Must be between 0 and 1. Only used if
early_stoppingis True. Values must be in the range(0.0, 1.0).Added in version 0.20: Added ‘validation_fraction’ option
- n_iter_no_changeint, default=5
Number of iterations with no improvement to wait before stopping fitting. Convergence is checked against the training loss or the validation loss depending on the
early_stoppingparameter. Integer values must be in the range[1, max_iter).Added in version 0.20: Added ‘n_iter_no_change’ option
- warm_startbool, default=False
When set to True, reuse the solution of the previous call to fit as initialization, otherwise, just erase the previous solution. See the Glossary.
Repeatedly calling fit or partial_fit when warm_start is True can result in a different solution than when calling fit a single time because of the way the data is shuffled. If a dynamic learning rate is used, the learning rate is adapted depending on the number of samples already seen. Calling
fitresets this counter, whilepartial_fitwill result in increasing the existing counter.- averagebool or int, default=False
When set to True, computes the averaged SGD weights across all updates and stores the result in the
coef_attribute. If set to an int greater than 1, averaging will begin once the total number of samples seen reachesaverage. Soaverage=10will begin averaging after seeing 10 samples.
- Attributes:
- coef_ndarray of shape (n_features,)
Weights assigned to the features.
- intercept_ndarray of shape (1,)
The intercept term.
- n_iter_int
The actual number of iterations before reaching the stopping criterion.
- t_int
Number of weight updates performed during training. Same as
(n_iter_ * n_samples + 1).- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
See also
HuberRegressorLinear regression model that is robust to outliers.
LarsLeast Angle Regression model.
LassoLinear Model trained with L1 prior as regularizer.
RANSACRegressorRANSAC (RANdom SAmple Consensus) algorithm.
RidgeLinear least squares with l2 regularization.
sklearn.svm.SVREpsilon-Support Vector Regression.
TheilSenRegressorTheil-Sen Estimator robust multivariate regression model.
References
[1] (1,2)Online Passive-Aggressive Algorithms <http://jmlr.csail.mit.edu/papers/volume7/crammer06a/crammer06a.pdf> K. Crammer, O. Dekel, J. Keshat, S. Shalev-Shwartz, Y. Singer - JMLR (2006)
Examples
>>> import numpy as np >>> from sklearn.linear_model import SGDRegressor >>> from sklearn.pipeline import make_pipeline >>> from sklearn.preprocessing import StandardScaler >>> n_samples, n_features = 10, 5 >>> rng = np.random.RandomState(0) >>> y = rng.randn(n_samples) >>> X = rng.randn(n_samples, n_features) >>> # Always scale the input. The most convenient way is to use a pipeline. >>> reg = make_pipeline(StandardScaler(), ... SGDRegressor(max_iter=1000, tol=1e-3)) >>> reg.fit(X, y) Pipeline(steps=[('standardscaler', StandardScaler()), ('sgdregressor', SGDRegressor())])
- densify()[source]#
Convert coefficient matrix to dense array format.
Converts the
coef_member (back) to a numpy.ndarray. This is the default format ofcoef_and is required for fitting, so calling this method is only required on models that have previously been sparsified; otherwise, it is a no-op.- Returns:
- self
Fitted estimator.
- fit(X, y, coef_init=None, intercept_init=None, sample_weight=None)[source]#
Fit linear model with Stochastic Gradient Descent.
- Parameters:
- X{array-like, sparse matrix}, shape (n_samples, n_features)
Training data.
- yndarray of shape (n_samples,)
Target values.
- coef_initndarray of shape (n_features,), default=None
The initial coefficients to warm-start the optimization.
- intercept_initndarray of shape (1,), default=None
The initial intercept to warm-start the optimization.
- sample_weightarray-like, shape (n_samples,), default=None
Weights applied to individual samples (1. for unweighted).
- Returns:
- selfobject
Fitted
SGDRegressorestimator.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- partial_fit(X, y, sample_weight=None)[source]#
Perform one epoch of stochastic gradient descent on given samples.
Internally, this method uses
max_iter = 1. Therefore, it is not guaranteed that a minimum of the cost function is reached after calling it once. Matters such as objective convergence and early stopping should be handled by the user.- Parameters:
- X{array-like, sparse matrix}, shape (n_samples, n_features)
Subset of training data.
- ynumpy array of shape (n_samples,)
Subset of target values.
- sample_weightarray-like, shape (n_samples,), default=None
Weights applied to individual samples. If not provided, uniform weights are assumed.
- Returns:
- selfobject
Returns an instance of self.
- predict(X)[source]#
Predict using the linear model.
- Parameters:
- X{array-like, sparse matrix}, shape (n_samples, n_features)
Input data.
- Returns:
- ndarray of shape (n_samples,)
Predicted target values per element in X.
- score(X, y, sample_weight=None)[source]#
Return coefficient of determination on test data.
The coefficient of determination, \(R^2\), is defined as \((1 - \frac{u}{v})\), where \(u\) is the residual sum of squares
((y_true - y_pred)** 2).sum()and \(v\) is the total sum of squares((y_true - y_true.mean()) ** 2).sum(). The best possible score is 1.0 and it can be negative (because the model can be arbitrarily worse). A constant model that always predicts the expected value ofy, disregarding the input features, would get a \(R^2\) score of 0.0.- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples. For some estimators this may be a precomputed kernel matrix or a list of generic objects instead with shape
(n_samples, n_samples_fitted), wheren_samples_fittedis the number of samples used in the fitting for the estimator.- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True values for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
\(R^2\) of
self.predict(X)w.r.t.y.
Notes
The \(R^2\) score used when calling
scoreon a regressor usesmultioutput='uniform_average'from version 0.23 to keep consistent with default value ofr2_score. This influences thescoremethod of all the multioutput regressors (except forMultiOutputRegressor).
- set_fit_request(*, coef_init: bool | None | str = '$UNCHANGED$', intercept_init: bool | None | str = '$UNCHANGED$', sample_weight: bool | None | str = '$UNCHANGED$') SGDRegressor[source]#
Configure whether metadata should be requested to be passed to the
fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- coef_initstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
coef_initparameter infit.- intercept_initstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
intercept_initparameter infit.- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter infit.
- Returns:
- selfobject
The updated object.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_partial_fit_request(*, sample_weight: bool | None | str = '$UNCHANGED$') SGDRegressor[source]#
Configure whether metadata should be requested to be passed to the
partial_fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed topartial_fitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it topartial_fit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inpartial_fit.
- Returns:
- selfobject
The updated object.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') SGDRegressor[source]#
Configure whether metadata should be requested to be passed to the
scoremethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.
- sparsify()[source]#
Convert coefficient matrix to sparse format.
Converts the
coef_member to a scipy.sparse matrix, which for L1-regularized models can be much more memory- and storage-efficient than the usual numpy.ndarray representation.The
intercept_member is not converted.- Returns:
- self
Fitted estimator.
Notes
For non-sparse models, i.e. when there are not many zeros in
coef_, this may actually increase memory usage, so use this method with care. A rule of thumb is that the number of zero elements, which can be computed with(coef_ == 0).sum(), must be more than 50% for this to provide significant benefits.After calling this method, further fitting with the partial_fit method (if any) will not work until you call densify.