AdaBoostClassifier#
- class sklearn.ensemble.AdaBoostClassifier(estimator=None, *, n_estimators=50, learning_rate=1.0, random_state=None)[source]#
An AdaBoost classifier.
An AdaBoost [1] classifier is a meta-estimator that begins by fitting a classifier on the original dataset and then fits additional copies of the classifier on the same dataset but where the weights of incorrectly classified instances are adjusted such that subsequent classifiers focus more on difficult cases.
This class implements the algorithm based on [2].
Read more in the User Guide.
Added in version 0.14.
- Parameters:
- estimatorobject, default=None
The base estimator from which the boosted ensemble is built. Support for sample weighting is required, as well as proper
classes_andn_classes_attributes. IfNone, then the base estimator isDecisionTreeClassifierinitialized withmax_depth=1.Added in version 1.2:
base_estimatorwas renamed toestimator.- n_estimatorsint, default=50
The maximum number of estimators at which boosting is terminated. In case of perfect fit, the learning procedure is stopped early. Values must be in the range
[1, inf).- learning_ratefloat, default=1.0
Weight applied to each classifier at each boosting iteration. A higher learning rate increases the contribution of each classifier. There is a trade-off between the
learning_rateandn_estimatorsparameters. Values must be in the range(0.0, inf).- random_stateint, RandomState instance or None, default=None
Controls the random seed given at each
estimatorat each boosting iteration. Thus, it is only used whenestimatorexposes arandom_state. Pass an int for reproducible output across multiple function calls. See Glossary.
- Attributes:
- estimator_estimator
The base estimator from which the ensemble is grown.
Added in version 1.2:
base_estimator_was renamed toestimator_.- estimators_list of classifiers
The collection of fitted sub-estimators.
- classes_ndarray of shape (n_classes,)
The classes labels.
- n_classes_int
The number of classes.
- estimator_weights_ndarray of floats
Weights for each estimator in the boosted ensemble.
- estimator_errors_ndarray of floats
Classification error for each estimator in the boosted ensemble.
feature_importances_ndarray of shape (n_features,)The impurity-based feature importances.
- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
See also
AdaBoostRegressorAn AdaBoost regressor that begins by fitting a regressor on the original dataset and then fits additional copies of the regressor on the same dataset but where the weights of instances are adjusted according to the error of the current prediction.
GradientBoostingClassifierGB builds an additive model in a forward stage-wise fashion. Regression trees are fit on the negative gradient of the binomial or multinomial deviance loss function. Binary classification is a special case where only a single regression tree is induced.
sklearn.tree.DecisionTreeClassifierA non-parametric supervised learning method used for classification. Creates a model that predicts the value of a target variable by learning simple decision rules inferred from the data features.
References
[1]Y. Freund, R. Schapire, “A Decision-Theoretic Generalization of on-Line Learning and an Application to Boosting”, 1995.
Examples
>>> from sklearn.ensemble import AdaBoostClassifier >>> from sklearn.datasets import make_classification >>> X, y = make_classification(n_samples=1000, n_features=4, ... n_informative=2, n_redundant=0, ... random_state=0, shuffle=False) >>> clf = AdaBoostClassifier(n_estimators=100, random_state=0) >>> clf.fit(X, y) AdaBoostClassifier(n_estimators=100, random_state=0) >>> clf.predict([[0, 0, 0, 0]]) array([1]) >>> clf.score(X, y) 0.96
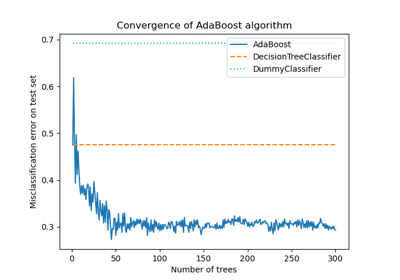
For a detailed example of using AdaBoost to fit a sequence of DecisionTrees as weaklearners, please refer to Multi-class AdaBoosted Decision Trees.
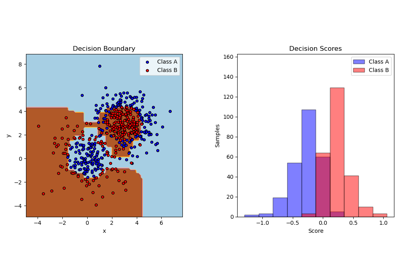
For a detailed example of using AdaBoost to fit a non-linearly separable classification dataset composed of two Gaussian quantiles clusters, please refer to Two-class AdaBoost.
- decision_function(X)[source]#
Compute the decision function of
X.- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrix can be CSC, CSR, COO, DOK, or LIL. COO, DOK, and LIL are converted to CSR.
- Returns:
- scorendarray of shape of (n_samples, k)
The decision function of the input samples. The order of outputs is the same as that of the classes_ attribute. Binary classification is a special cases with
k == 1, otherwisek==n_classes. For binary classification, values closer to -1 or 1 mean more like the first or second class inclasses_, respectively.
- fit(X, y, sample_weight=None)[source]#
Build a boosted classifier/regressor from the training set (X, y).
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrix can be CSC, CSR, COO, DOK, or LIL. COO, DOK, and LIL are converted to CSR.
- yarray-like of shape (n_samples,)
The target values.
- sample_weightarray-like of shape (n_samples,), default=None
Sample weights. If None, the sample weights are initialized to 1 / n_samples.
- Returns:
- selfobject
Fitted estimator.
- get_metadata_routing()[source]#
Raise
NotImplementedError.This estimator does not support metadata routing yet.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- predict(X)[source]#
Predict classes for X.
The predicted class of an input sample is computed as the weighted mean prediction of the classifiers in the ensemble.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrix can be CSC, CSR, COO, DOK, or LIL. COO, DOK, and LIL are converted to CSR.
- Returns:
- yndarray of shape (n_samples,)
The predicted classes.
- predict_log_proba(X)[source]#
Predict class log-probabilities for X.
The predicted class log-probabilities of an input sample is computed as the weighted mean predicted class log-probabilities of the classifiers in the ensemble.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrix can be CSC, CSR, COO, DOK, or LIL. COO, DOK, and LIL are converted to CSR.
- Returns:
- pndarray of shape (n_samples, n_classes)
The class probabilities of the input samples. The order of outputs is the same of that of the classes_ attribute.
- predict_proba(X)[source]#
Predict class probabilities for X.
The predicted class probabilities of an input sample is computed as the weighted mean predicted class probabilities of the classifiers in the ensemble.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrix can be CSC, CSR, COO, DOK, or LIL. COO, DOK, and LIL are converted to CSR.
- Returns:
- pndarray of shape (n_samples, n_classes)
The class probabilities of the input samples. The order of outputs is the same of that of the classes_ attribute.
- score(X, y, sample_weight=None)[source]#
Return accuracy on provided data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True labels for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
Mean accuracy of
self.predict(X)w.r.t.y.
- set_fit_request(*, sample_weight: bool | None | str = '$UNCHANGED$') AdaBoostClassifier[source]#
Configure whether metadata should be requested to be passed to the
fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter infit.
- Returns:
- selfobject
The updated object.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') AdaBoostClassifier[source]#
Configure whether metadata should be requested to be passed to the
scoremethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.
- staged_decision_function(X)[source]#
Compute decision function of
Xfor each boosting iteration.This method allows monitoring (i.e. determine error on testing set) after each boosting iteration.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrix can be CSC, CSR, COO, DOK, or LIL. COO, DOK, and LIL are converted to CSR.
- Yields:
- scoregenerator of ndarray of shape (n_samples, k)
The decision function of the input samples. The order of outputs is the same of that of the classes_ attribute. Binary classification is a special cases with
k == 1, otherwisek==n_classes. For binary classification, values closer to -1 or 1 mean more like the first or second class inclasses_, respectively.
- staged_predict(X)[source]#
Return staged predictions for X.
The predicted class of an input sample is computed as the weighted mean prediction of the classifiers in the ensemble.
This generator method yields the ensemble prediction after each iteration of boosting and therefore allows monitoring, such as to determine the prediction on a test set after each boost.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
The input samples. Sparse matrix can be CSC, CSR, COO, DOK, or LIL. COO, DOK, and LIL are converted to CSR.
- Yields:
- ygenerator of ndarray of shape (n_samples,)
The predicted classes.
- staged_predict_proba(X)[source]#
Predict class probabilities for X.
The predicted class probabilities of an input sample is computed as the weighted mean predicted class probabilities of the classifiers in the ensemble.
This generator method yields the ensemble predicted class probabilities after each iteration of boosting and therefore allows monitoring, such as to determine the predicted class probabilities on a test set after each boost.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrix can be CSC, CSR, COO, DOK, or LIL. COO, DOK, and LIL are converted to CSR.
- Yields:
- pgenerator of ndarray of shape (n_samples,)
The class probabilities of the input samples. The order of outputs is the same of that of the classes_ attribute.
- staged_score(X, y, sample_weight=None)[source]#
Return staged scores for X, y.
This generator method yields the ensemble score after each iteration of boosting and therefore allows monitoring, such as to determine the score on a test set after each boost.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrix can be CSC, CSR, COO, DOK, or LIL. COO, DOK, and LIL are converted to CSR.
- yarray-like of shape (n_samples,)
Labels for X.
- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Yields:
- zfloat
Gallery examples#
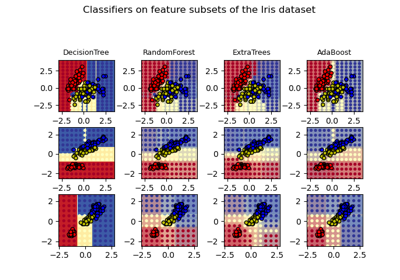
Plot the decision surfaces of ensembles of trees on the iris dataset