Isomap#
- class sklearn.manifold.Isomap(*, n_neighbors=5, radius=None, n_components=2, eigen_solver='auto', tol=0, max_iter=None, path_method='auto', neighbors_algorithm='auto', n_jobs=None, metric='minkowski', p=2, metric_params=None)[source]#
Isomap Embedding.
Non-linear dimensionality reduction through Isometric Mapping
Read more in the User Guide.
- Parameters:
- n_neighborsint or None, default=5
Number of neighbors to consider for each point. If
n_neighborsis an int, thenradiusmust beNone.- radiusfloat or None, default=None
Limiting distance of neighbors to return. If
radiusis a float, thenn_neighborsmust be set toNone.Added in version 1.1.
- n_componentsint, default=2
Number of coordinates for the manifold.
- eigen_solver{‘auto’, ‘arpack’, ‘dense’}, default=’auto’
‘auto’ : Attempt to choose the most efficient solver for the given problem.
‘arpack’ : Use Arnoldi decomposition to find the eigenvalues and eigenvectors.
‘dense’ : Use a direct solver (i.e. LAPACK) for the eigenvalue decomposition.
- tolfloat, default=0
Convergence tolerance passed to arpack or lobpcg. not used if eigen_solver == ‘dense’.
- max_iterint, default=None
Maximum number of iterations for the arpack solver. not used if eigen_solver == ‘dense’.
- path_method{‘auto’, ‘FW’, ‘D’}, default=’auto’
Method to use in finding shortest path.
‘auto’ : attempt to choose the best algorithm automatically.
‘FW’ : Floyd-Warshall algorithm.
‘D’ : Dijkstra’s algorithm.
- neighbors_algorithm{‘auto’, ‘brute’, ‘kd_tree’, ‘ball_tree’}, default=’auto’
Algorithm to use for nearest neighbors search, passed to neighbors.NearestNeighbors instance.
- n_jobsint or None, default=None
The number of parallel jobs to run.
Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.- metricstr, or callable, default=”minkowski”
The metric to use when calculating distance between instances in a feature array. If metric is a string or callable, it must be one of the options allowed by
sklearn.metrics.pairwise_distancesfor its metric parameter. If metric is “precomputed”, X is assumed to be a distance matrix and must be square. X may be a Glossary.Added in version 0.22.
- pfloat, default=2
Parameter for the Minkowski metric from sklearn.metrics.pairwise.pairwise_distances. When p = 1, this is equivalent to using manhattan_distance (l1), and euclidean_distance (l2) for p = 2. For arbitrary p, minkowski_distance (l_p) is used.
Added in version 0.22.
- metric_paramsdict, default=None
Additional keyword arguments for the metric function.
Added in version 0.22.
- Attributes:
- embedding_array-like, shape (n_samples, n_components)
Stores the embedding vectors.
- kernel_pca_object
KernelPCAobject used to implement the embedding.- nbrs_sklearn.neighbors.NearestNeighbors instance
Stores nearest neighbors instance, including BallTree or KDtree if applicable.
- dist_matrix_array-like, shape (n_samples, n_samples)
Stores the geodesic distance matrix of training data.
- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
See also
sklearn.decomposition.PCAPrincipal component analysis that is a linear dimensionality reduction method.
sklearn.decomposition.KernelPCANon-linear dimensionality reduction using kernels and PCA.
MDSManifold learning using multidimensional scaling.
TSNET-distributed Stochastic Neighbor Embedding.
LocallyLinearEmbeddingManifold learning using Locally Linear Embedding.
SpectralEmbeddingSpectral embedding for non-linear dimensionality.
References
[1]Tenenbaum, J.B.; De Silva, V.; & Langford, J.C. A global geometric framework for nonlinear dimensionality reduction. Science 290 (5500)
Examples
>>> from sklearn.datasets import load_digits >>> from sklearn.manifold import Isomap >>> X, _ = load_digits(return_X_y=True) >>> X.shape (1797, 64) >>> embedding = Isomap(n_components=2) >>> X_transformed = embedding.fit_transform(X[:100]) >>> X_transformed.shape (100, 2)
- fit(X, y=None)[source]#
Compute the embedding vectors for data X.
- Parameters:
- X{array-like, sparse matrix, BallTree, KDTree, NearestNeighbors}
Sample data, shape = (n_samples, n_features), in the form of a numpy array, sparse matrix, precomputed tree, or NearestNeighbors object.
- yIgnored
Not used, present for API consistency by convention.
- Returns:
- selfobject
Returns a fitted instance of self.
- fit_transform(X, y=None)[source]#
Fit the model from data in X and transform X.
- Parameters:
- X{array-like, sparse matrix, BallTree, KDTree}
Training vector, where
n_samplesis the number of samples andn_featuresis the number of features.- yIgnored
Not used, present for API consistency by convention.
- Returns:
- X_newarray-like, shape (n_samples, n_components)
X transformed in the new space.
- get_feature_names_out(input_features=None)[source]#
Get output feature names for transformation.
The feature names out will prefixed by the lowercased class name. For example, if the transformer outputs 3 features, then the feature names out are:
["class_name0", "class_name1", "class_name2"].- Parameters:
- input_featuresarray-like of str or None, default=None
Only used to validate feature names with the names seen in
fit.
- Returns:
- feature_names_outndarray of str objects
Transformed feature names.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- reconstruction_error()[source]#
Compute the reconstruction error for the embedding.
- Returns:
- reconstruction_errorfloat
Reconstruction error.
Notes
The cost function of an isomap embedding is
E = frobenius_norm[K(D) - K(D_fit)] / n_samplesWhere D is the matrix of distances for the input data X, D_fit is the matrix of distances for the output embedding X_fit, and K is the isomap kernel:
K(D) = -0.5 * (I - 1/n_samples) * D^2 * (I - 1/n_samples)
- set_output(*, transform=None)[source]#
Set output container.
Refer to the user guide for more details and Introducing the set_output API for an example on how to use the API.
- Parameters:
- transform{“default”, “pandas”, “polars”}, default=None
Configure output of
transformandfit_transform."default": Default output format of a transformer"pandas": DataFrame output"polars": Polars outputNone: Transform configuration is unchanged
Added in version 1.4:
"polars"option was added.
- Returns:
- selfestimator instance
Estimator instance.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- transform(X)[source]#
Transform X.
This is implemented by linking the points X into the graph of geodesic distances of the training data. First the
n_neighborsnearest neighbors of X are found in the training data, and from these the shortest geodesic distances from each point in X to each point in the training data are computed in order to construct the kernel. The embedding of X is the projection of this kernel onto the embedding vectors of the training set.- Parameters:
- X{array-like, sparse matrix}, shape (n_queries, n_features)
If neighbors_algorithm=’precomputed’, X is assumed to be a distance matrix or a sparse graph of shape (n_queries, n_samples_fit).
- Returns:
- X_newarray-like, shape (n_queries, n_components)
X transformed in the new space.
Gallery examples#

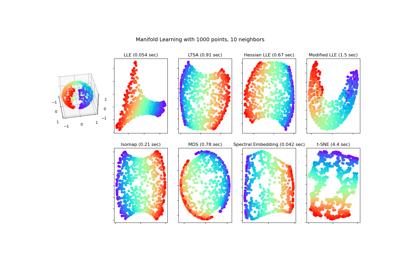
Manifold learning on handwritten digits: Locally Linear Embedding, Isomap…