Note
Go to the end to download the full example code or to run this example in your browser via JupyterLite or Binder.
Time-related feature engineering#
This notebook introduces different strategies to leverage time-related features for a bike sharing demand regression task that is highly dependent on business cycles (days, weeks, months) and yearly season cycles.
In the process, we introduce how to perform periodic feature engineering using
the sklearn.preprocessing.SplineTransformer class and its
extrapolation="periodic" option.
# Authors: The scikit-learn developers
# SPDX-License-Identifier: BSD-3-Clause
Data exploration on the Bike Sharing Demand dataset#
We start by loading the data from the OpenML repository.
from sklearn.datasets import fetch_openml
bike_sharing = fetch_openml("Bike_Sharing_Demand", version=2, as_frame=True)
df = bike_sharing.frame
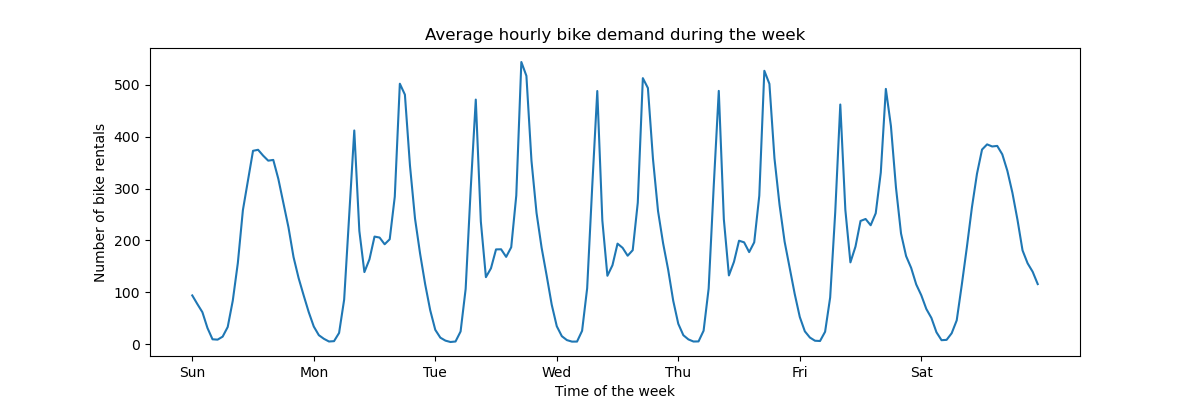
To get a quick understanding of the periodic patterns of the data, let us have a look at the average demand per hour during a week.
Note that the week starts on a Sunday, during the weekend. We can clearly distinguish the commute patterns in the morning and evenings of the work days and the leisure use of the bikes on the weekends with a more spread peak demand around the middle of the days:
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(12, 4))
average_week_demand = df.groupby(["weekday", "hour"])["count"].mean()
average_week_demand.plot(ax=ax)
_ = ax.set(
title="Average hourly bike demand during the week",
xticks=[i * 24 for i in range(7)],
xticklabels=["Sun", "Mon", "Tue", "Wed", "Thu", "Fri", "Sat"],
xlabel="Time of the week",
ylabel="Number of bike rentals",
)
The target of the prediction problem is the absolute count of bike rentals on an hourly basis:
df["count"].max()
np.int64(977)
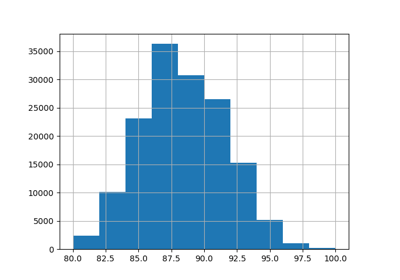
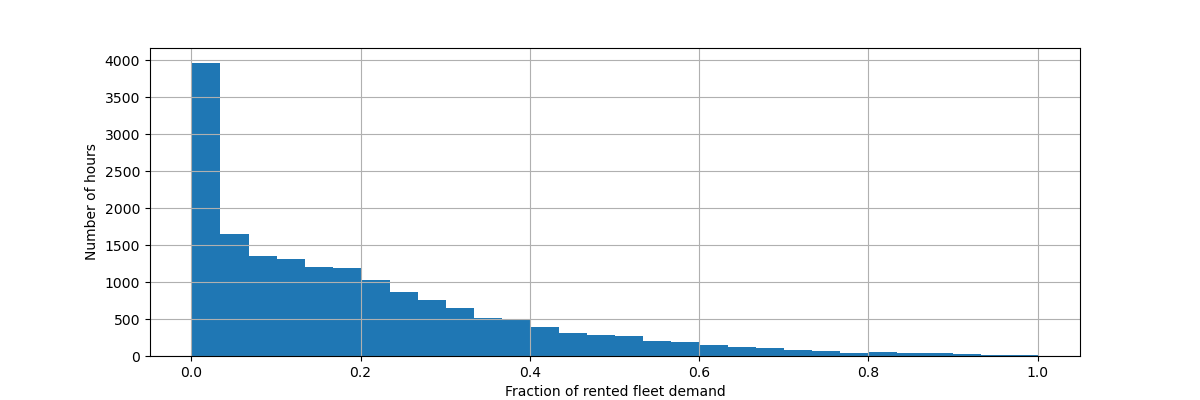
Let us rescale the target variable (number of hourly bike rentals) to predict a relative demand so that the mean absolute error is more easily interpreted as a fraction of the maximum demand.
Note
The fit method of the models used in this notebook all minimizes the mean squared error to estimate the conditional mean. The absolute error, however, would estimate the conditional median.
Nevertheless, when reporting performance measures on the test set in the discussion, we choose to focus on the mean absolute error instead of the (root) mean squared error because it is more intuitive to interpret. Note, however, that in this study the best models for one metric are also the best ones in terms of the other metric.
y = df["count"] / df["count"].max()
fig, ax = plt.subplots(figsize=(12, 4))
y.hist(bins=30, ax=ax)
_ = ax.set(
xlabel="Fraction of rented fleet demand",
ylabel="Number of hours",
)
The input feature data frame is a time annotated hourly log of variables describing the weather conditions. It includes both numerical and categorical variables. Note that the time information has already been expanded into several complementary columns.
X = df.drop("count", axis="columns")
X
| season | year | month | hour | holiday | weekday | workingday | weather | temp | feel_temp | humidity | windspeed | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | spring | 0 | 1 | 0 | False | 6 | False | clear | 9.84 | 14.395 | 0.81 | 0.0000 |
| 1 | spring | 0 | 1 | 1 | False | 6 | False | clear | 9.02 | 13.635 | 0.80 | 0.0000 |
| 2 | spring | 0 | 1 | 2 | False | 6 | False | clear | 9.02 | 13.635 | 0.80 | 0.0000 |
| 3 | spring | 0 | 1 | 3 | False | 6 | False | clear | 9.84 | 14.395 | 0.75 | 0.0000 |
| 4 | spring | 0 | 1 | 4 | False | 6 | False | clear | 9.84 | 14.395 | 0.75 | 0.0000 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 17374 | spring | 1 | 12 | 19 | False | 1 | True | misty | 10.66 | 12.880 | 0.60 | 11.0014 |
| 17375 | spring | 1 | 12 | 20 | False | 1 | True | misty | 10.66 | 12.880 | 0.60 | 11.0014 |
| 17376 | spring | 1 | 12 | 21 | False | 1 | True | clear | 10.66 | 12.880 | 0.60 | 11.0014 |
| 17377 | spring | 1 | 12 | 22 | False | 1 | True | clear | 10.66 | 13.635 | 0.56 | 8.9981 |
| 17378 | spring | 1 | 12 | 23 | False | 1 | True | clear | 10.66 | 13.635 | 0.65 | 8.9981 |
17379 rows × 12 columns
Note
If the time information was only present as a date or datetime column, we could have expanded it into hour-in-the-day, day-in-the-week, day-in-the-month, month-in-the-year using pandas: https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#time-date-components
We now introspect the distribution of the categorical variables, starting
with "weather":
X["weather"].value_counts()
weather
clear 11413
misty 4544
rain 1419
heavy_rain 3
Name: count, dtype: int64
Since there are only 3 "heavy_rain" events, we cannot use this category to
train machine learning models with cross validation. Instead, we simplify the
representation by collapsing those into the "rain" category.
X["weather"] = (
X["weather"]
.astype(object)
.replace(to_replace="heavy_rain", value="rain")
.astype("category")
)
X["weather"].value_counts()
weather
clear 11413
misty 4544
rain 1422
Name: count, dtype: int64
As expected, the "season" variable is well balanced:
X["season"].value_counts()
season
fall 4496
summer 4409
spring 4242
winter 4232
Name: count, dtype: int64
Time-based cross-validation#
Since the dataset is a time-ordered event log (hourly demand), we will use a time-sensitive cross-validation splitter to evaluate our demand forecasting model as realistically as possible. We use a gap of 2 days between the train and test side of the splits. We also limit the training set size to make the performance of the CV folds more stable.
1000 test datapoints should be enough to quantify the performance of the model. This represents a bit less than a month and a half of contiguous test data:
from sklearn.model_selection import TimeSeriesSplit
ts_cv = TimeSeriesSplit(
n_splits=5,
gap=48,
max_train_size=10000,
test_size=1000,
)
Let us manually inspect the various splits to check that the
TimeSeriesSplit works as we expect, starting with the first split:
all_splits = list(ts_cv.split(X, y))
train_0, test_0 = all_splits[0]
X.iloc[test_0]
| season | year | month | hour | holiday | weekday | workingday | weather | temp | feel_temp | humidity | windspeed | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 12379 | summer | 1 | 6 | 0 | False | 2 | True | clear | 22.14 | 25.760 | 0.68 | 27.9993 |
| 12380 | summer | 1 | 6 | 1 | False | 2 | True | misty | 21.32 | 25.000 | 0.77 | 22.0028 |
| 12381 | summer | 1 | 6 | 2 | False | 2 | True | rain | 21.32 | 25.000 | 0.72 | 19.9995 |
| 12382 | summer | 1 | 6 | 3 | False | 2 | True | rain | 20.50 | 24.240 | 0.82 | 12.9980 |
| 12383 | summer | 1 | 6 | 4 | False | 2 | True | rain | 20.50 | 24.240 | 0.82 | 12.9980 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 13374 | fall | 1 | 7 | 11 | False | 1 | True | clear | 34.44 | 40.150 | 0.53 | 15.0013 |
| 13375 | fall | 1 | 7 | 12 | False | 1 | True | clear | 34.44 | 39.395 | 0.49 | 8.9981 |
| 13376 | fall | 1 | 7 | 13 | False | 1 | True | clear | 34.44 | 39.395 | 0.49 | 19.0012 |
| 13377 | fall | 1 | 7 | 14 | False | 1 | True | clear | 36.08 | 40.910 | 0.42 | 7.0015 |
| 13378 | fall | 1 | 7 | 15 | False | 1 | True | clear | 35.26 | 40.150 | 0.47 | 16.9979 |
1000 rows × 12 columns
X.iloc[train_0]
| season | year | month | hour | holiday | weekday | workingday | weather | temp | feel_temp | humidity | windspeed | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2331 | summer | 0 | 4 | 1 | False | 2 | True | misty | 25.42 | 31.060 | 0.50 | 6.0032 |
| 2332 | summer | 0 | 4 | 2 | False | 2 | True | misty | 24.60 | 31.060 | 0.53 | 8.9981 |
| 2333 | summer | 0 | 4 | 3 | False | 2 | True | misty | 23.78 | 27.275 | 0.56 | 8.9981 |
| 2334 | summer | 0 | 4 | 4 | False | 2 | True | misty | 22.96 | 26.515 | 0.64 | 8.9981 |
| 2335 | summer | 0 | 4 | 5 | False | 2 | True | misty | 22.14 | 25.760 | 0.68 | 8.9981 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 12326 | summer | 1 | 6 | 19 | False | 6 | False | clear | 26.24 | 31.060 | 0.36 | 11.0014 |
| 12327 | summer | 1 | 6 | 20 | False | 6 | False | clear | 25.42 | 31.060 | 0.35 | 19.0012 |
| 12328 | summer | 1 | 6 | 21 | False | 6 | False | clear | 24.60 | 31.060 | 0.40 | 7.0015 |
| 12329 | summer | 1 | 6 | 22 | False | 6 | False | clear | 23.78 | 27.275 | 0.46 | 8.9981 |
| 12330 | summer | 1 | 6 | 23 | False | 6 | False | clear | 22.96 | 26.515 | 0.52 | 7.0015 |
10000 rows × 12 columns
We now inspect the last split:
train_4, test_4 = all_splits[4]
X.iloc[test_4]
| season | year | month | hour | holiday | weekday | workingday | weather | temp | feel_temp | humidity | windspeed | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 16379 | winter | 1 | 11 | 5 | False | 2 | True | misty | 13.94 | 16.665 | 0.66 | 8.9981 |
| 16380 | winter | 1 | 11 | 6 | False | 2 | True | misty | 13.94 | 16.665 | 0.71 | 11.0014 |
| 16381 | winter | 1 | 11 | 7 | False | 2 | True | clear | 13.12 | 16.665 | 0.76 | 6.0032 |
| 16382 | winter | 1 | 11 | 8 | False | 2 | True | clear | 13.94 | 16.665 | 0.71 | 8.9981 |
| 16383 | winter | 1 | 11 | 9 | False | 2 | True | misty | 14.76 | 18.940 | 0.71 | 0.0000 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 17374 | spring | 1 | 12 | 19 | False | 1 | True | misty | 10.66 | 12.880 | 0.60 | 11.0014 |
| 17375 | spring | 1 | 12 | 20 | False | 1 | True | misty | 10.66 | 12.880 | 0.60 | 11.0014 |
| 17376 | spring | 1 | 12 | 21 | False | 1 | True | clear | 10.66 | 12.880 | 0.60 | 11.0014 |
| 17377 | spring | 1 | 12 | 22 | False | 1 | True | clear | 10.66 | 13.635 | 0.56 | 8.9981 |
| 17378 | spring | 1 | 12 | 23 | False | 1 | True | clear | 10.66 | 13.635 | 0.65 | 8.9981 |
1000 rows × 12 columns
X.iloc[train_4]
| season | year | month | hour | holiday | weekday | workingday | weather | temp | feel_temp | humidity | windspeed | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 6331 | winter | 0 | 9 | 9 | False | 1 | True | misty | 26.24 | 28.790 | 0.89 | 12.9980 |
| 6332 | winter | 0 | 9 | 10 | False | 1 | True | misty | 26.24 | 28.790 | 0.89 | 12.9980 |
| 6333 | winter | 0 | 9 | 11 | False | 1 | True | clear | 27.88 | 31.820 | 0.79 | 15.0013 |
| 6334 | winter | 0 | 9 | 12 | False | 1 | True | misty | 27.88 | 31.820 | 0.79 | 11.0014 |
| 6335 | winter | 0 | 9 | 13 | False | 1 | True | misty | 28.70 | 33.335 | 0.74 | 11.0014 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 16326 | winter | 1 | 11 | 0 | False | 0 | False | misty | 12.30 | 15.150 | 0.70 | 11.0014 |
| 16327 | winter | 1 | 11 | 1 | False | 0 | False | clear | 12.30 | 14.395 | 0.70 | 12.9980 |
| 16328 | winter | 1 | 11 | 2 | False | 0 | False | clear | 11.48 | 14.395 | 0.81 | 7.0015 |
| 16329 | winter | 1 | 11 | 3 | False | 0 | False | misty | 12.30 | 15.150 | 0.81 | 11.0014 |
| 16330 | winter | 1 | 11 | 4 | False | 0 | False | misty | 12.30 | 14.395 | 0.81 | 12.9980 |
10000 rows × 12 columns
All is well. We are now ready to do some predictive modeling!
Gradient Boosting#
Gradient Boosting Regression with decision trees is often flexible enough to efficiently handle heterogeneous tabular data with a mix of categorical and numerical features as long as the number of samples is large enough.
Here, we use the modern
HistGradientBoostingRegressor with native support
for categorical features. Therefore, we only need to set
categorical_features="from_dtype" such that features with categorical dtype
are considered categorical features. For reference, we extract the categorical
features from the dataframe based on the dtype. The internal trees use a dedicated
tree splitting rule for these features.
The numerical variables need no preprocessing and, for the sake of simplicity, we only try the default hyper-parameters for this model:
from sklearn.compose import ColumnTransformer
from sklearn.ensemble import HistGradientBoostingRegressor
from sklearn.model_selection import cross_validate
from sklearn.pipeline import make_pipeline
gbrt = HistGradientBoostingRegressor(categorical_features="from_dtype", random_state=42)
categorical_columns = X.columns[X.dtypes == "category"]
print("Categorical features:", categorical_columns.tolist())
Categorical features: ['season', 'holiday', 'workingday', 'weather']
Let’s evaluate our gradient boosting model with the mean absolute error of the relative demand averaged across our 5 time-based cross-validation splits:
import numpy as np
def evaluate(model, X, y, cv, model_prop=None, model_step=None):
cv_results = cross_validate(
model,
X,
y,
cv=cv,
scoring=["neg_mean_absolute_error", "neg_root_mean_squared_error"],
return_estimator=True,
)
if model_prop is not None:
if model_step is not None:
values = [
getattr(m[model_step], model_prop) for m in cv_results["estimator"]
]
else:
values = [getattr(m, model_prop) for m in cv_results["estimator"]]
print(f"Mean model.{model_prop} = {np.mean(values)}")
mae = -cv_results["test_neg_mean_absolute_error"]
rmse = -cv_results["test_neg_root_mean_squared_error"]
print(
f"Mean Absolute Error: {mae.mean():.3f} +/- {mae.std():.3f}\n"
f"Root Mean Squared Error: {rmse.mean():.3f} +/- {rmse.std():.3f}"
)
# To display the fitted estimator diagrams in the notebook.
return cv_results["estimator"][0]
evaluate(gbrt, X, y, cv=ts_cv, model_prop="n_iter_")
Mean model.n_iter_ = 100.0
Mean Absolute Error: 0.044 +/- 0.003
Root Mean Squared Error: 0.068 +/- 0.005
HistGradientBoostingRegressor(random_state=42)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
Fitted attributes
We see that we set max_iter large enough such that early stopping took place.
This model has an average error around 4 to 5% of the maximum demand. This is quite good for a first trial without any hyper-parameter tuning! We just had to make the categorical variables explicit. Note that the time related features are passed as is, i.e. without processing them. But this is not much of a problem for tree-based models as they can learn a non-monotonic relationship between ordinal input features and the target.
This is not the case for linear regression models as we will see in the following.
Naive linear regression#
As usual for linear models, categorical variables need to be one-hot encoded.
For consistency, we scale the numerical features to the same 0-1 range using
MinMaxScaler, although in this case it does not
impact the results much because they are already on comparable scales:
from sklearn.linear_model import RidgeCV
from sklearn.preprocessing import MinMaxScaler, OneHotEncoder
one_hot_encoder = OneHotEncoder(handle_unknown="ignore", sparse_output=False)
alphas = np.logspace(-6, 6, 25)
naive_linear_pipeline = make_pipeline(
ColumnTransformer(
transformers=[
("categorical", one_hot_encoder, categorical_columns),
],
remainder=MinMaxScaler(),
verbose_feature_names_out=False,
),
RidgeCV(alphas=alphas),
)
evaluate(
naive_linear_pipeline, X, y, cv=ts_cv, model_prop="alpha_", model_step="ridgecv"
)
Mean model.alpha_ = 2.7298221281347037
Mean Absolute Error: 0.142 +/- 0.014
Root Mean Squared Error: 0.184 +/- 0.020
Pipeline(steps=[('columntransformer',
ColumnTransformer(remainder=MinMaxScaler(),
transformers=[('categorical',
OneHotEncoder(handle_unknown='ignore',
sparse_output=False),
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str'))],
verbose_feature_names_out=False)),
('ridgecv',
RidgeCV(alphas=array([1.00000000e-06, 3.16227766e-06, 1.00000000e-05, 3.16227766e-05,
1.00000000e-04, 3.16227766e-04, 1.00000000e-03, 3.16227766e-03,
1.00000000e-02, 3.16227766e-02, 1.00000000e-01, 3.16227766e-01,
1.00000000e+00, 3.16227766e+00, 1.00000000e+01, 3.16227766e+01,
1.00000000e+02, 3.16227766e+02, 1.00000000e+03, 3.16227766e+03,
1.00000000e+04, 3.16227766e+04, 1.00000000e+05, 3.16227766e+05,
1.00000000e+06])))])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Only defined if the underlying estimator exposes such an attribute when fit. .. versionadded:: 1.0 |
ndarray[object](12,) | ['season','year','month',...,'feel_temp','humidity','windspeed'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. Only defined if the underlying first estimator in `steps` exposes such an attribute when fit. .. versionadded:: 0.24 |
int | 12 |
Parameters
Fitted attributes
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')
Parameters
Fitted attributes
11 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
['year', 'month', 'hour', 'weekday', 'temp', 'feel_temp', 'humidity', 'windspeed']
Parameters
Fitted attributes
8 features
| year |
| month |
| hour |
| weekday |
| temp |
| feel_temp |
| humidity |
| windspeed |
19 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
| year |
| month |
| hour |
| weekday |
| temp |
| feel_temp |
| humidity |
| windspeed |
Parameters
Fitted attributes
It is affirmative to see that the selected alpha_ is in our specified
range.
The performance is not good: the average error is around 14% of the maximum demand. This is more than three times higher than the average error of the gradient boosting model. We can suspect that the naive original encoding (merely min-max scaled) of the periodic time-related features might prevent the linear regression model to properly leverage the time information: linear regression does not automatically model non-monotonic relationships between the input features and the target. Non-linear terms have to be engineered in the input.
For example, the raw numerical encoding of the "hour" feature prevents the
linear model from recognizing that an increase of hour in the morning from 6
to 8 should have a strong positive impact on the number of bike rentals while
an increase of similar magnitude in the evening from 18 to 20 should have a
strong negative impact on the predicted number of bike rentals.
Time-steps as categories#
Since the time features are encoded in a discrete manner using integers (24 unique values in the “hours” feature), we could decide to treat those as categorical variables using a one-hot encoding and thereby ignore any assumption implied by the ordering of the hour values.
Using one-hot encoding for the time features gives the linear model a lot more flexibility as we introduce one additional feature per discrete time level.
one_hot_linear_pipeline = make_pipeline(
ColumnTransformer(
transformers=[
("categorical", one_hot_encoder, categorical_columns),
("one_hot_time", one_hot_encoder, ["hour", "weekday", "month"]),
],
remainder=MinMaxScaler(),
verbose_feature_names_out=False,
),
RidgeCV(alphas=alphas),
)
evaluate(one_hot_linear_pipeline, X, y, cv=ts_cv)
Mean Absolute Error: 0.099 +/- 0.011
Root Mean Squared Error: 0.131 +/- 0.011
Pipeline(steps=[('columntransformer',
ColumnTransformer(remainder=MinMaxScaler(),
transformers=[('categorical',
OneHotEncoder(handle_unknown='ignore',
sparse_output=False),
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')),
('one_hot_time',
OneHotEncoder(handle_unknown='ignore',
sparse_output=False),
['hour', 'weekday',
'month'])],
verbose_featur...
RidgeCV(alphas=array([1.00000000e-06, 3.16227766e-06, 1.00000000e-05, 3.16227766e-05,
1.00000000e-04, 3.16227766e-04, 1.00000000e-03, 3.16227766e-03,
1.00000000e-02, 3.16227766e-02, 1.00000000e-01, 3.16227766e-01,
1.00000000e+00, 3.16227766e+00, 1.00000000e+01, 3.16227766e+01,
1.00000000e+02, 3.16227766e+02, 1.00000000e+03, 3.16227766e+03,
1.00000000e+04, 3.16227766e+04, 1.00000000e+05, 3.16227766e+05,
1.00000000e+06])))])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Only defined if the underlying estimator exposes such an attribute when fit. .. versionadded:: 1.0 |
ndarray[object](12,) | ['season','year','month',...,'feel_temp','humidity','windspeed'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. Only defined if the underlying first estimator in `steps` exposes such an attribute when fit. .. versionadded:: 0.24 |
int | 12 |
Parameters
Fitted attributes
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')
Parameters
Fitted attributes
11 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
['hour', 'weekday', 'month']
Parameters
Fitted attributes
43 features
| hour_0 |
| hour_1 |
| hour_2 |
| hour_3 |
| hour_4 |
| hour_5 |
| hour_6 |
| hour_7 |
| hour_8 |
| hour_9 |
| hour_10 |
| hour_11 |
| hour_12 |
| hour_13 |
| hour_14 |
| hour_15 |
| hour_16 |
| hour_17 |
| hour_18 |
| hour_19 |
| hour_20 |
| hour_21 |
| hour_22 |
| hour_23 |
| weekday_0 |
| weekday_1 |
| weekday_2 |
| weekday_3 |
| weekday_4 |
| weekday_5 |
| weekday_6 |
| month_1 |
| month_2 |
| month_3 |
| month_4 |
| month_5 |
| month_6 |
| month_7 |
| month_8 |
| month_9 |
| month_10 |
| month_11 |
| month_12 |
['year', 'temp', 'feel_temp', 'humidity', 'windspeed']
Parameters
Fitted attributes
5 features
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
59 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
| hour_0 |
| hour_1 |
| hour_2 |
| hour_3 |
| hour_4 |
| hour_5 |
| hour_6 |
| hour_7 |
| hour_8 |
| hour_9 |
| hour_10 |
| hour_11 |
| hour_12 |
| hour_13 |
| hour_14 |
| hour_15 |
| hour_16 |
| hour_17 |
| hour_18 |
| hour_19 |
| hour_20 |
| hour_21 |
| hour_22 |
| hour_23 |
| weekday_0 |
| weekday_1 |
| weekday_2 |
| weekday_3 |
| weekday_4 |
| weekday_5 |
| weekday_6 |
| month_1 |
| month_2 |
| month_3 |
| month_4 |
| month_5 |
| month_6 |
| month_7 |
| month_8 |
| month_9 |
| month_10 |
| month_11 |
| month_12 |
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
Parameters
Fitted attributes
The average error rate of this model is 10% which is much better than using the original (ordinal) encoding of the time feature, confirming our intuition that the linear regression model benefits from the added flexibility to not treat time progression in a monotonic manner.
However, this introduces a very large number of new features. If the time of
the day was represented in minutes since the start of the day instead of
hours, one-hot encoding would have introduced 1440 features instead of 24.
This could cause some significant overfitting. To avoid this we could use
sklearn.preprocessing.KBinsDiscretizer instead to re-bin the number
of levels of fine-grained ordinal or numerical variables while still
benefitting from the non-monotonic expressivity advantages of one-hot
encoding.
Finally, we also observe that one-hot encoding completely ignores the ordering of the hour levels while this could be an interesting inductive bias to preserve to some level. In the following we try to explore smooth, non-monotonic encoding that locally preserves the relative ordering of time features.
Trigonometric features#
As a first attempt, we can try to encode each of those periodic features using a sine and cosine transformation with the matching period.
Each ordinal time feature is transformed into 2 features that together encode equivalent information in a non-monotonic way, and more importantly without any jump between the first and the last value of the periodic range.
from sklearn.preprocessing import FunctionTransformer
def sin_transformer(period):
return FunctionTransformer(
lambda x: np.sin(x / period * 2 * np.pi), feature_names_out="one-to-one"
)
def cos_transformer(period):
return FunctionTransformer(
lambda x: np.cos(x / period * 2 * np.pi), feature_names_out="one-to-one"
)
Let us visualize the effect of this feature expansion on some synthetic hour data with a bit of extrapolation beyond hour=23:
import pandas as pd
hour_df = pd.DataFrame(
np.arange(26).reshape(-1, 1),
columns=["hour"],
)
hour_df["hour_sin"] = sin_transformer(24).fit_transform(hour_df)["hour"]
hour_df["hour_cos"] = cos_transformer(24).fit_transform(hour_df)["hour"]
hour_df.plot(x="hour")
_ = plt.title("Trigonometric encoding for the 'hour' feature")
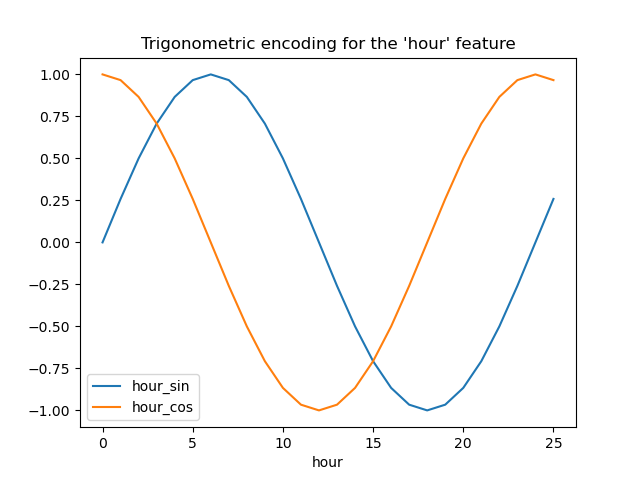
Let’s use a 2D scatter plot with the hours encoded as colors to better see how this representation maps the 24 hours of the day to a 2D space, akin to some sort of a 24 hour version of an analog clock. Note that the “25th” hour is mapped back to the 1st hour because of the periodic nature of the sine/cosine representation.
fig, ax = plt.subplots(figsize=(7, 5))
sp = ax.scatter(hour_df["hour_sin"], hour_df["hour_cos"], c=hour_df["hour"])
ax.set(
xlabel="sin(hour)",
ylabel="cos(hour)",
)
_ = fig.colorbar(sp)

We can now build a feature extraction pipeline using this strategy:
cyclic_cossin_transformer = ColumnTransformer(
transformers=[
("categorical", one_hot_encoder, categorical_columns),
("month_sin", sin_transformer(12), ["month"]),
("month_cos", cos_transformer(12), ["month"]),
("weekday_sin", sin_transformer(7), ["weekday"]),
("weekday_cos", cos_transformer(7), ["weekday"]),
("hour_sin", sin_transformer(24), ["hour"]),
("hour_cos", cos_transformer(24), ["hour"]),
],
remainder=MinMaxScaler(),
verbose_feature_names_out=True,
)
cyclic_cossin_linear_pipeline = make_pipeline(
cyclic_cossin_transformer,
RidgeCV(alphas=alphas),
)
evaluate(cyclic_cossin_linear_pipeline, X, y, cv=ts_cv)
Mean Absolute Error: 0.125 +/- 0.014
Root Mean Squared Error: 0.166 +/- 0.020
Pipeline(steps=[('columntransformer',
ColumnTransformer(remainder=MinMaxScaler(),
transformers=[('categorical',
OneHotEncoder(handle_unknown='ignore',
sparse_output=False),
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')),
('month_sin',
FunctionTransformer(feature_names_out='one-to-one',
func=<function sin_transformer.<locals>.<lambda> at 0x758...
RidgeCV(alphas=array([1.00000000e-06, 3.16227766e-06, 1.00000000e-05, 3.16227766e-05,
1.00000000e-04, 3.16227766e-04, 1.00000000e-03, 3.16227766e-03,
1.00000000e-02, 3.16227766e-02, 1.00000000e-01, 3.16227766e-01,
1.00000000e+00, 3.16227766e+00, 1.00000000e+01, 3.16227766e+01,
1.00000000e+02, 3.16227766e+02, 1.00000000e+03, 3.16227766e+03,
1.00000000e+04, 3.16227766e+04, 1.00000000e+05, 3.16227766e+05,
1.00000000e+06])))])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Only defined if the underlying estimator exposes such an attribute when fit. .. versionadded:: 1.0 |
ndarray[object](12,) | ['season','year','month',...,'feel_temp','humidity','windspeed'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. Only defined if the underlying first estimator in `steps` exposes such an attribute when fit. .. versionadded:: 0.24 |
int | 12 |
Parameters
Fitted attributes
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')
Parameters
Fitted attributes
11 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
['month']
Parameters
Fitted attributes
1 feature
| month |
['month']
Parameters
Fitted attributes
1 feature
| month |
['weekday']
Parameters
Fitted attributes
1 feature
| weekday |
['weekday']
Parameters
Fitted attributes
1 feature
| weekday |
['hour']
Parameters
Fitted attributes
1 feature
| hour |
['hour']
Parameters
Fitted attributes
1 feature
| hour |
['year', 'temp', 'feel_temp', 'humidity', 'windspeed']
Parameters
Fitted attributes
5 features
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
22 features
| categorical__season_fall |
| categorical__season_spring |
| categorical__season_summer |
| categorical__season_winter |
| categorical__holiday_False |
| categorical__holiday_True |
| categorical__workingday_False |
| categorical__workingday_True |
| categorical__weather_clear |
| categorical__weather_misty |
| categorical__weather_rain |
| month_sin__month |
| month_cos__month |
| weekday_sin__weekday |
| weekday_cos__weekday |
| hour_sin__hour |
| hour_cos__hour |
| remainder__year |
| remainder__temp |
| remainder__feel_temp |
| remainder__humidity |
| remainder__windspeed |
Parameters
Fitted attributes
The performance of our linear regression model with this simple feature engineering is a bit better than using the original ordinal time features but worse than using the one-hot encoded time features. We will further analyze possible reasons for this disappointing outcome at the end of this notebook.
Periodic spline features#
We can try an alternative encoding of the periodic time-related features using spline transformations with a large enough number of splines, and as a result a larger number of expanded features compared to the sine/cosine transformation:
from sklearn.preprocessing import SplineTransformer
def periodic_spline_transformer(period, n_splines=None, degree=3):
if n_splines is None:
n_splines = period
n_knots = n_splines + 1 # periodic and include_bias is True
return SplineTransformer(
degree=degree,
n_knots=n_knots,
knots=np.linspace(0, period, n_knots).reshape(n_knots, 1),
extrapolation="periodic",
include_bias=True,
)
Again, let us visualize the effect of this feature expansion on some synthetic hour data with a bit of extrapolation beyond hour=23:
hour_df = pd.DataFrame(
np.linspace(0, 26, 1000).reshape(-1, 1),
columns=["hour"],
)
splines = periodic_spline_transformer(24, n_splines=12).fit_transform(hour_df)
splines_df = pd.DataFrame(
splines,
columns=[f"spline_{i}" for i in range(splines.shape[1])],
)
pd.concat([hour_df, splines_df], axis="columns").plot(x="hour", cmap=plt.cm.tab20b)
_ = plt.title("Periodic spline-based encoding for the 'hour' feature")
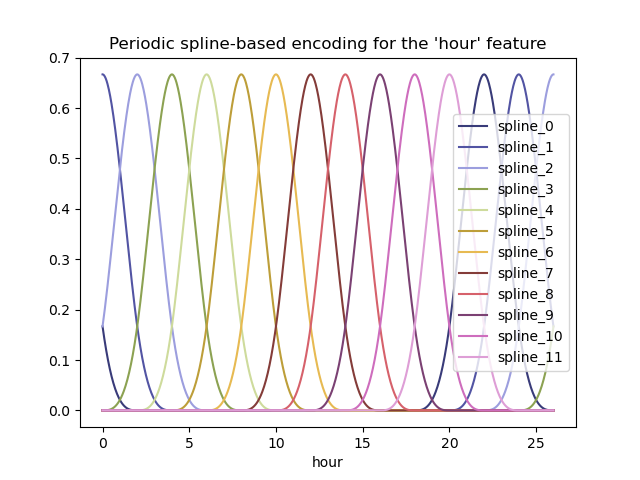
Thanks to the use of the extrapolation="periodic" parameter, we observe
that the feature encoding stays smooth when extrapolating beyond midnight.
We can now build a predictive pipeline using this alternative periodic feature engineering strategy.
It is possible to use fewer splines than discrete levels for those ordinal values. This makes spline-based encoding more efficient than one-hot encoding while preserving most of the expressivity:
cyclic_spline_transformer = ColumnTransformer(
transformers=[
("categorical", one_hot_encoder, categorical_columns),
("cyclic_month", periodic_spline_transformer(12, n_splines=6), ["month"]),
("cyclic_weekday", periodic_spline_transformer(7, n_splines=3), ["weekday"]),
("cyclic_hour", periodic_spline_transformer(24, n_splines=12), ["hour"]),
],
remainder=MinMaxScaler(),
verbose_feature_names_out=False,
)
cyclic_spline_linear_pipeline = make_pipeline(
cyclic_spline_transformer,
RidgeCV(alphas=alphas),
)
evaluate(cyclic_spline_linear_pipeline, X, y, cv=ts_cv)
Mean Absolute Error: 0.097 +/- 0.011
Root Mean Squared Error: 0.132 +/- 0.013
Pipeline(steps=[('columntransformer',
ColumnTransformer(remainder=MinMaxScaler(),
transformers=[('categorical',
OneHotEncoder(handle_unknown='ignore',
sparse_output=False),
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')),
('cyclic_month',
SplineTransformer(extrapolation='periodic',
knots=array([[ 0.],
[ 2.],
[ 4.],
[ 6.],
[ 8.],
[10.],
[12.]]),
n_knots...
RidgeCV(alphas=array([1.00000000e-06, 3.16227766e-06, 1.00000000e-05, 3.16227766e-05,
1.00000000e-04, 3.16227766e-04, 1.00000000e-03, 3.16227766e-03,
1.00000000e-02, 3.16227766e-02, 1.00000000e-01, 3.16227766e-01,
1.00000000e+00, 3.16227766e+00, 1.00000000e+01, 3.16227766e+01,
1.00000000e+02, 3.16227766e+02, 1.00000000e+03, 3.16227766e+03,
1.00000000e+04, 3.16227766e+04, 1.00000000e+05, 3.16227766e+05,
1.00000000e+06])))])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Only defined if the underlying estimator exposes such an attribute when fit. .. versionadded:: 1.0 |
ndarray[object](12,) | ['season','year','month',...,'feel_temp','humidity','windspeed'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. Only defined if the underlying first estimator in `steps` exposes such an attribute when fit. .. versionadded:: 0.24 |
int | 12 |
Parameters
Fitted attributes
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')
Parameters
Fitted attributes
11 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
['month']
Parameters
Fitted attributes
6 features
| month_sp_0 |
| month_sp_1 |
| month_sp_2 |
| month_sp_3 |
| month_sp_4 |
| month_sp_5 |
['weekday']
Parameters
Fitted attributes
3 features
| weekday_sp_0 |
| weekday_sp_1 |
| weekday_sp_2 |
['hour']
Parameters
Fitted attributes
12 features
| hour_sp_0 |
| hour_sp_1 |
| hour_sp_2 |
| hour_sp_3 |
| hour_sp_4 |
| hour_sp_5 |
| hour_sp_6 |
| hour_sp_7 |
| hour_sp_8 |
| hour_sp_9 |
| hour_sp_10 |
| hour_sp_11 |
['year', 'temp', 'feel_temp', 'humidity', 'windspeed']
Parameters
Fitted attributes
5 features
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
37 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
| month_sp_0 |
| month_sp_1 |
| month_sp_2 |
| month_sp_3 |
| month_sp_4 |
| month_sp_5 |
| weekday_sp_0 |
| weekday_sp_1 |
| weekday_sp_2 |
| hour_sp_0 |
| hour_sp_1 |
| hour_sp_2 |
| hour_sp_3 |
| hour_sp_4 |
| hour_sp_5 |
| hour_sp_6 |
| hour_sp_7 |
| hour_sp_8 |
| hour_sp_9 |
| hour_sp_10 |
| hour_sp_11 |
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
Parameters
Fitted attributes
Spline features make it possible for the linear model to successfully leverage the periodic time-related features and reduce the error from ~14% to ~10% of the maximum demand, which is similar to what we observed with the one-hot encoded features.
Qualitative analysis of the impact of features on linear model predictions#
Here, we want to visualize the impact of the feature engineering choices on the time related shape of the predictions.
To do so we consider an arbitrary time-based split to compare the predictions on a range of held out data points.
naive_linear_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
naive_linear_predictions = naive_linear_pipeline.predict(X.iloc[test_0])
one_hot_linear_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
one_hot_linear_predictions = one_hot_linear_pipeline.predict(X.iloc[test_0])
cyclic_cossin_linear_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
cyclic_cossin_linear_predictions = cyclic_cossin_linear_pipeline.predict(X.iloc[test_0])
cyclic_spline_linear_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
cyclic_spline_linear_predictions = cyclic_spline_linear_pipeline.predict(X.iloc[test_0])
We visualize those predictions by zooming on the last 96 hours (4 days) of the test set to get some qualitative insights:
last_hours = slice(-96, None)
fig, ax = plt.subplots(figsize=(12, 4))
fig.suptitle("Predictions by linear models")
ax.plot(
y.iloc[test_0].values[last_hours],
"x-",
alpha=0.2,
label="Actual demand",
color="black",
)
ax.plot(naive_linear_predictions[last_hours], "x-", label="Ordinal time features")
ax.plot(
cyclic_cossin_linear_predictions[last_hours],
"x-",
label="Trigonometric time features",
)
ax.plot(
cyclic_spline_linear_predictions[last_hours],
"x-",
label="Spline-based time features",
)
ax.plot(
one_hot_linear_predictions[last_hours],
"x-",
label="One-hot time features",
)
_ = ax.legend()
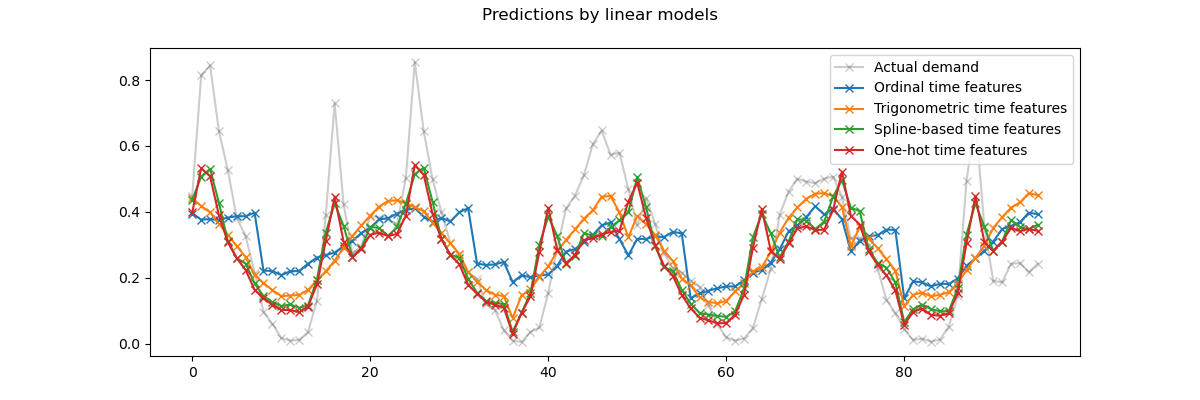
We can draw the following conclusions from the above plot:
The raw ordinal time-related features are problematic because they do not capture the natural periodicity: we observe a big jump in the predictions at the end of each day when the hour features goes from 23 back to 0. We can expect similar artifacts at the end of each week or each year.
As expected, the trigonometric features (sine and cosine) do not have these discontinuities at midnight, but the linear regression model fails to leverage those features to properly model intra-day variations. Using trigonometric features for higher harmonics or additional trigonometric features for the natural period with different phases could potentially fix this problem.
the periodic spline-based features fix those two problems at once: they give more expressivity to the linear model by making it possible to focus on specific hours thanks to the use of 12 splines. Furthermore the
extrapolation="periodic"option enforces a smooth representation betweenhour=23andhour=0.The one-hot encoded features behave similarly to the periodic spline-based features but are more spiky: for instance they can better model the morning peak during the week days since this peak lasts shorter than an hour. However, we will see in the following that what can be an advantage for linear models is not necessarily one for more expressive models.
We can also compare the number of features extracted by each feature engineering pipeline:
naive_linear_pipeline[:-1].transform(X).shape
(17379, 19)
one_hot_linear_pipeline[:-1].transform(X).shape
(17379, 59)
cyclic_cossin_linear_pipeline[:-1].transform(X).shape
(17379, 22)
cyclic_spline_linear_pipeline[:-1].transform(X).shape
(17379, 37)
This confirms that the one-hot encoding and the spline encoding strategies create a lot more features for the time representation than the alternatives, which in turn gives the downstream linear model more flexibility (degrees of freedom) to avoid underfitting.
Finally, we observe that none of the linear models can approximate the true bike rentals demand, especially for the peaks that can be very sharp at rush hours during the working days but much flatter during the week-ends: the most accurate linear models based on splines or one-hot encoding tend to forecast peaks of commuting-related bike rentals even on the week-ends and under-estimate the commuting-related events during the working days.
These systematic prediction errors reveal a form of under-fitting and can be explained by the lack of interactions terms between features, e.g. “workingday” and features derived from “hours”. This issue will be addressed in the following section.
Modeling pairwise interactions with splines and polynomial features#
Linear models do not automatically capture interaction effects between input
features. It does not help that some features are marginally non-linear as is
the case with features constructed by SplineTransformer (or one-hot
encoding or binning).
However, it is possible to use the PolynomialFeatures class on coarse
grained spline encoded hours to model the “workingday”/”hours” interaction
explicitly without introducing too many new variables:
from sklearn.pipeline import FeatureUnion
from sklearn.preprocessing import PolynomialFeatures
hour_workday_interaction = make_pipeline(
ColumnTransformer(
[
("cyclic_hour", periodic_spline_transformer(24, n_splines=8), ["hour"]),
(
"workingday",
FunctionTransformer(
lambda x: x == "True", feature_names_out="one-to-one"
),
["workingday"],
),
],
verbose_feature_names_out=False,
),
PolynomialFeatures(degree=2, interaction_only=True, include_bias=False),
)
Those features are then combined with the ones already computed in the previous spline-base pipeline. We can observe a nice performance improvement by modeling this pairwise interaction explicitly:
cyclic_spline_interactions_pipeline = make_pipeline(
FeatureUnion(
[
("marginal", cyclic_spline_transformer),
("interactions", hour_workday_interaction),
],
verbose_feature_names_out=True,
).set_output(transform="pandas"),
RidgeCV(alphas=alphas),
)
evaluate(cyclic_spline_interactions_pipeline, X, y, cv=ts_cv)
Mean Absolute Error: 0.078 +/- 0.009
Root Mean Squared Error: 0.104 +/- 0.009
Pipeline(steps=[('featureunion',
FeatureUnion(transformer_list=[('marginal',
ColumnTransformer(remainder=MinMaxScaler(),
transformers=[('categorical',
OneHotEncoder(handle_unknown='ignore',
sparse_output=False),
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')),
('cyclic_month',
SplineTransformer(extrapolation='periodic',
knots=array([[ 0.],
[ 2....
RidgeCV(alphas=array([1.00000000e-06, 3.16227766e-06, 1.00000000e-05, 3.16227766e-05,
1.00000000e-04, 3.16227766e-04, 1.00000000e-03, 3.16227766e-03,
1.00000000e-02, 3.16227766e-02, 1.00000000e-01, 3.16227766e-01,
1.00000000e+00, 3.16227766e+00, 1.00000000e+01, 3.16227766e+01,
1.00000000e+02, 3.16227766e+02, 1.00000000e+03, 3.16227766e+03,
1.00000000e+04, 3.16227766e+04, 1.00000000e+05, 3.16227766e+05,
1.00000000e+06])))])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Only defined if the underlying estimator exposes such an attribute when fit. .. versionadded:: 1.0 |
ndarray[object](12,) | ['season','year','month',...,'feel_temp','humidity','windspeed'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. Only defined if the underlying first estimator in `steps` exposes such an attribute when fit. .. versionadded:: 0.24 |
int | 12 |
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Defined only when `X` has feature names that are all strings. .. versionadded:: 1.3 |
ndarray[object](12,) | ['season','year','month',...,'feel_temp','humidity','windspeed'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. Only defined if the underlying first transformer in `transformer_list` exposes such an attribute when fit. .. versionadded:: 0.24 |
int | 12 |
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')
Parameters
Fitted attributes
11 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
['month']
Parameters
Fitted attributes
6 features
| month_sp_0 |
| month_sp_1 |
| month_sp_2 |
| month_sp_3 |
| month_sp_4 |
| month_sp_5 |
['weekday']
Parameters
Fitted attributes
3 features
| weekday_sp_0 |
| weekday_sp_1 |
| weekday_sp_2 |
['hour']
Parameters
Fitted attributes
12 features
| hour_sp_0 |
| hour_sp_1 |
| hour_sp_2 |
| hour_sp_3 |
| hour_sp_4 |
| hour_sp_5 |
| hour_sp_6 |
| hour_sp_7 |
| hour_sp_8 |
| hour_sp_9 |
| hour_sp_10 |
| hour_sp_11 |
['year', 'temp', 'feel_temp', 'humidity', 'windspeed']
Parameters
Fitted attributes
5 features
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
37 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
| month_sp_0 |
| month_sp_1 |
| month_sp_2 |
| month_sp_3 |
| month_sp_4 |
| month_sp_5 |
| weekday_sp_0 |
| weekday_sp_1 |
| weekday_sp_2 |
| hour_sp_0 |
| hour_sp_1 |
| hour_sp_2 |
| hour_sp_3 |
| hour_sp_4 |
| hour_sp_5 |
| hour_sp_6 |
| hour_sp_7 |
| hour_sp_8 |
| hour_sp_9 |
| hour_sp_10 |
| hour_sp_11 |
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
Parameters
Fitted attributes
['hour']
Parameters
Fitted attributes
8 features
| hour_sp_0 |
| hour_sp_1 |
| hour_sp_2 |
| hour_sp_3 |
| hour_sp_4 |
| hour_sp_5 |
| hour_sp_6 |
| hour_sp_7 |
['workingday']
Parameters
Fitted attributes
1 feature
| workingday |
9 features
| hour_sp_0 |
| hour_sp_1 |
| hour_sp_2 |
| hour_sp_3 |
| hour_sp_4 |
| hour_sp_5 |
| hour_sp_6 |
| hour_sp_7 |
| workingday |
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Defined only when `X` has feature names that are all strings. .. versionadded:: 1.0 |
ndarray[object](9,) | ['hour_sp_0','hour_sp_1','hour_sp_2',...,'hour_sp_6','hour_sp_7', 'workingday'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. .. versionadded:: 0.24 |
int | 9 |
|
n_output_features_
n_output_features_: int The total number of polynomial output features. The number of output features is computed by iterating over all suitably sized combinations of input features. |
int | 45 |
|
powers_
powers_: ndarray of shape (`n_output_features_`, `n_features_in_`) `powers_[i, j]` is the exponent of the jth input in the ith output. |
ndarray[int64](45, 9) | [[1,0,0,...,0,0,0], [0,1,0,...,0,0,0], [0,0,1,...,0,0,0], ..., [0,0,0,...,1,1,0], [0,0,0,...,1,0,1], [0,0,0,...,0,1,1]] |
45 features
| hour_sp_0 |
| hour_sp_1 |
| hour_sp_2 |
| hour_sp_3 |
| hour_sp_4 |
| hour_sp_5 |
| hour_sp_6 |
| hour_sp_7 |
| workingday |
| hour_sp_0 hour_sp_1 |
| hour_sp_0 hour_sp_2 |
| hour_sp_0 hour_sp_3 |
| hour_sp_0 hour_sp_4 |
| hour_sp_0 hour_sp_5 |
| hour_sp_0 hour_sp_6 |
| hour_sp_0 hour_sp_7 |
| hour_sp_0 workingday |
| hour_sp_1 hour_sp_2 |
| hour_sp_1 hour_sp_3 |
| hour_sp_1 hour_sp_4 |
| hour_sp_1 hour_sp_5 |
| hour_sp_1 hour_sp_6 |
| hour_sp_1 hour_sp_7 |
| hour_sp_1 workingday |
| hour_sp_2 hour_sp_3 |
| hour_sp_2 hour_sp_4 |
| hour_sp_2 hour_sp_5 |
| hour_sp_2 hour_sp_6 |
| hour_sp_2 hour_sp_7 |
| hour_sp_2 workingday |
| hour_sp_3 hour_sp_4 |
| hour_sp_3 hour_sp_5 |
| hour_sp_3 hour_sp_6 |
| hour_sp_3 hour_sp_7 |
| hour_sp_3 workingday |
| hour_sp_4 hour_sp_5 |
| hour_sp_4 hour_sp_6 |
| hour_sp_4 hour_sp_7 |
| hour_sp_4 workingday |
| hour_sp_5 hour_sp_6 |
| hour_sp_5 hour_sp_7 |
| hour_sp_5 workingday |
| hour_sp_6 hour_sp_7 |
| hour_sp_6 workingday |
| hour_sp_7 workingday |
82 features
| marginal__season_fall |
| marginal__season_spring |
| marginal__season_summer |
| marginal__season_winter |
| marginal__holiday_False |
| marginal__holiday_True |
| marginal__workingday_False |
| marginal__workingday_True |
| marginal__weather_clear |
| marginal__weather_misty |
| marginal__weather_rain |
| marginal__month_sp_0 |
| marginal__month_sp_1 |
| marginal__month_sp_2 |
| marginal__month_sp_3 |
| marginal__month_sp_4 |
| marginal__month_sp_5 |
| marginal__weekday_sp_0 |
| marginal__weekday_sp_1 |
| marginal__weekday_sp_2 |
| marginal__hour_sp_0 |
| marginal__hour_sp_1 |
| marginal__hour_sp_2 |
| marginal__hour_sp_3 |
| marginal__hour_sp_4 |
| marginal__hour_sp_5 |
| marginal__hour_sp_6 |
| marginal__hour_sp_7 |
| marginal__hour_sp_8 |
| marginal__hour_sp_9 |
| marginal__hour_sp_10 |
| marginal__hour_sp_11 |
| marginal__year |
| marginal__temp |
| marginal__feel_temp |
| marginal__humidity |
| marginal__windspeed |
| interactions__hour_sp_0 |
| interactions__hour_sp_1 |
| interactions__hour_sp_2 |
| interactions__hour_sp_3 |
| interactions__hour_sp_4 |
| interactions__hour_sp_5 |
| interactions__hour_sp_6 |
| interactions__hour_sp_7 |
| interactions__workingday |
| interactions__hour_sp_0 hour_sp_1 |
| interactions__hour_sp_0 hour_sp_2 |
| interactions__hour_sp_0 hour_sp_3 |
| interactions__hour_sp_0 hour_sp_4 |
| interactions__hour_sp_0 hour_sp_5 |
| interactions__hour_sp_0 hour_sp_6 |
| interactions__hour_sp_0 hour_sp_7 |
| interactions__hour_sp_0 workingday |
| interactions__hour_sp_1 hour_sp_2 |
| interactions__hour_sp_1 hour_sp_3 |
| interactions__hour_sp_1 hour_sp_4 |
| interactions__hour_sp_1 hour_sp_5 |
| interactions__hour_sp_1 hour_sp_6 |
| interactions__hour_sp_1 hour_sp_7 |
| interactions__hour_sp_1 workingday |
| interactions__hour_sp_2 hour_sp_3 |
| interactions__hour_sp_2 hour_sp_4 |
| interactions__hour_sp_2 hour_sp_5 |
| interactions__hour_sp_2 hour_sp_6 |
| interactions__hour_sp_2 hour_sp_7 |
| interactions__hour_sp_2 workingday |
| interactions__hour_sp_3 hour_sp_4 |
| interactions__hour_sp_3 hour_sp_5 |
| interactions__hour_sp_3 hour_sp_6 |
| interactions__hour_sp_3 hour_sp_7 |
| interactions__hour_sp_3 workingday |
| interactions__hour_sp_4 hour_sp_5 |
| interactions__hour_sp_4 hour_sp_6 |
| interactions__hour_sp_4 hour_sp_7 |
| interactions__hour_sp_4 workingday |
| interactions__hour_sp_5 hour_sp_6 |
| interactions__hour_sp_5 hour_sp_7 |
| interactions__hour_sp_5 workingday |
| interactions__hour_sp_6 hour_sp_7 |
| interactions__hour_sp_6 workingday |
| interactions__hour_sp_7 workingday |
Parameters
Fitted attributes
Modeling non-linear feature interactions with kernels#
The previous analysis highlighted the need to model the interactions between
"workingday" and "hours". Another example of a such a non-linear
interaction that we would like to model could be the impact of the rain that
might not be the same during the working days and the week-ends and holidays
for instance.
To model all such interactions, we could either use a polynomial expansion on all marginal features at once, after their spline-based expansion. However, this would create a quadratic number of features which can cause overfitting and computational tractability issues.
Alternatively, we can use the Nyström method to compute an approximate polynomial kernel expansion. Let us try the latter:
from sklearn.kernel_approximation import Nystroem
cyclic_spline_poly_pipeline = make_pipeline(
cyclic_spline_transformer,
Nystroem(kernel="poly", degree=2, n_components=300, random_state=0),
RidgeCV(alphas=alphas),
)
evaluate(cyclic_spline_poly_pipeline, X, y, cv=ts_cv)
Mean Absolute Error: 0.053 +/- 0.002
Root Mean Squared Error: 0.076 +/- 0.004
Pipeline(steps=[('columntransformer',
ColumnTransformer(remainder=MinMaxScaler(),
transformers=[('categorical',
OneHotEncoder(handle_unknown='ignore',
sparse_output=False),
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')),
('cyclic_month',
SplineTransformer(extrapolation='periodic',
knots=array([[ 0.],
[ 2.],
[ 4.],
[ 6.],
[ 8.],
[10.],
[12.]]),
n_knots...
RidgeCV(alphas=array([1.00000000e-06, 3.16227766e-06, 1.00000000e-05, 3.16227766e-05,
1.00000000e-04, 3.16227766e-04, 1.00000000e-03, 3.16227766e-03,
1.00000000e-02, 3.16227766e-02, 1.00000000e-01, 3.16227766e-01,
1.00000000e+00, 3.16227766e+00, 1.00000000e+01, 3.16227766e+01,
1.00000000e+02, 3.16227766e+02, 1.00000000e+03, 3.16227766e+03,
1.00000000e+04, 3.16227766e+04, 1.00000000e+05, 3.16227766e+05,
1.00000000e+06])))])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Only defined if the underlying estimator exposes such an attribute when fit. .. versionadded:: 1.0 |
ndarray[object](12,) | ['season','year','month',...,'feel_temp','humidity','windspeed'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. Only defined if the underlying first estimator in `steps` exposes such an attribute when fit. .. versionadded:: 0.24 |
int | 12 |
Parameters
Fitted attributes
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')
Parameters
Fitted attributes
11 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
['month']
Parameters
Fitted attributes
6 features
| month_sp_0 |
| month_sp_1 |
| month_sp_2 |
| month_sp_3 |
| month_sp_4 |
| month_sp_5 |
['weekday']
Parameters
Fitted attributes
3 features
| weekday_sp_0 |
| weekday_sp_1 |
| weekday_sp_2 |
['hour']
Parameters
Fitted attributes
12 features
| hour_sp_0 |
| hour_sp_1 |
| hour_sp_2 |
| hour_sp_3 |
| hour_sp_4 |
| hour_sp_5 |
| hour_sp_6 |
| hour_sp_7 |
| hour_sp_8 |
| hour_sp_9 |
| hour_sp_10 |
| hour_sp_11 |
['year', 'temp', 'feel_temp', 'humidity', 'windspeed']
Parameters
Fitted attributes
5 features
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
37 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
| month_sp_0 |
| month_sp_1 |
| month_sp_2 |
| month_sp_3 |
| month_sp_4 |
| month_sp_5 |
| weekday_sp_0 |
| weekday_sp_1 |
| weekday_sp_2 |
| hour_sp_0 |
| hour_sp_1 |
| hour_sp_2 |
| hour_sp_3 |
| hour_sp_4 |
| hour_sp_5 |
| hour_sp_6 |
| hour_sp_7 |
| hour_sp_8 |
| hour_sp_9 |
| hour_sp_10 |
| hour_sp_11 |
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
component_indices_
component_indices_: ndarray of shape (n_components) Indices of ``components_`` in the training set. |
ndarray[int64](300,) | [9394, 898,2398,...,2685,5725,8051] |
|
components_
components_: ndarray of shape (n_components, n_features) Subset of training points used to construct the feature map. |
ndarray[float64](300, 37) | [[0. ,0. ,1. ,...,0.6 ,0.63,0.33], [0. ,0. ,1. ,...,0.51,0.86,0.19], [1. ,0. ,0. ,...,0.74,0.7 ,0.3 ], ..., [1. ,0. ,0. ,...,0.65,0.64,0.11], [0. ,0. ,0. ,...,0.43,1. ,0. ], [0. ,1. ,0. ,...,0.63,0.25,0.12]] |
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Defined only when `X` has feature names that are all strings. .. versionadded:: 1.0 |
ndarray[object](37,) | ['season_fall','season_spring','season_summer',...,'feel_temp','humidity', 'windspeed'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. .. versionadded:: 0.24 |
int | 37 |
|
normalization_
normalization_: ndarray of shape (n_components, n_components) Normalization matrix needed for embedding. Square root of the kernel matrix on ``components_``. |
ndarray[float64](300, 300) | [[ 71.07, 2.07, 11.92,..., -3.79, -1.28, -4.85], [ 2.07, 42.81, 2.42,..., 1.85, -1.69, -0.97], [ 11.92, 2.42,149.76,...,-20.1 , 0.77, -2.76], ..., [ -3.79, 1.85,-20.1 ,...,112.03, -0.52, -2.54], [ -1.28, -1.69, 0.77,..., -0.52, 27.07, 1.13], [ -4.85, -0.97, -2.76,..., -2.54, 1.13, 82.79]] |
300 features
| nystroem0 |
| nystroem1 |
| nystroem2 |
| nystroem3 |
| nystroem4 |
| nystroem5 |
| nystroem6 |
| nystroem7 |
| nystroem8 |
| nystroem9 |
| nystroem10 |
| nystroem11 |
| nystroem12 |
| nystroem13 |
| nystroem14 |
| nystroem15 |
| nystroem16 |
| nystroem17 |
| nystroem18 |
| nystroem19 |
| nystroem20 |
| nystroem21 |
| nystroem22 |
| nystroem23 |
| nystroem24 |
| nystroem25 |
| nystroem26 |
| nystroem27 |
| nystroem28 |
| nystroem29 |
| nystroem30 |
| nystroem31 |
| nystroem32 |
| nystroem33 |
| nystroem34 |
| nystroem35 |
| nystroem36 |
| nystroem37 |
| nystroem38 |
| nystroem39 |
| nystroem40 |
| nystroem41 |
| nystroem42 |
| nystroem43 |
| nystroem44 |
| nystroem45 |
| nystroem46 |
| nystroem47 |
| nystroem48 |
| nystroem49 |
| nystroem50 |
| nystroem51 |
| nystroem52 |
| nystroem53 |
| nystroem54 |
| nystroem55 |
| nystroem56 |
| nystroem57 |
| nystroem58 |
| nystroem59 |
| nystroem60 |
| nystroem61 |
| nystroem62 |
| nystroem63 |
| nystroem64 |
| nystroem65 |
| nystroem66 |
| nystroem67 |
| nystroem68 |
| nystroem69 |
| nystroem70 |
| nystroem71 |
| nystroem72 |
| nystroem73 |
| nystroem74 |
| nystroem75 |
| nystroem76 |
| nystroem77 |
| nystroem78 |
| nystroem79 |
| nystroem80 |
| nystroem81 |
| nystroem82 |
| nystroem83 |
| nystroem84 |
| nystroem85 |
| nystroem86 |
| nystroem87 |
| nystroem88 |
| nystroem89 |
| nystroem90 |
| nystroem91 |
| nystroem92 |
| nystroem93 |
| nystroem94 |
| nystroem95 |
| nystroem96 |
| nystroem97 |
| nystroem98 |
| nystroem99 |
| nystroem100 |
| nystroem101 |
| nystroem102 |
| nystroem103 |
| nystroem104 |
| nystroem105 |
| nystroem106 |
| nystroem107 |
| nystroem108 |
| nystroem109 |
| nystroem110 |
| nystroem111 |
| nystroem112 |
| nystroem113 |
| nystroem114 |
| nystroem115 |
| nystroem116 |
| nystroem117 |
| nystroem118 |
| nystroem119 |
| nystroem120 |
| nystroem121 |
| nystroem122 |
| nystroem123 |
| nystroem124 |
| nystroem125 |
| nystroem126 |
| nystroem127 |
| nystroem128 |
| nystroem129 |
| nystroem130 |
| nystroem131 |
| nystroem132 |
| nystroem133 |
| nystroem134 |
| nystroem135 |
| nystroem136 |
| nystroem137 |
| nystroem138 |
| nystroem139 |
| nystroem140 |
| nystroem141 |
| nystroem142 |
| nystroem143 |
| nystroem144 |
| nystroem145 |
| nystroem146 |
| nystroem147 |
| nystroem148 |
| nystroem149 |
| nystroem150 |
| nystroem151 |
| nystroem152 |
| nystroem153 |
| nystroem154 |
| nystroem155 |
| nystroem156 |
| nystroem157 |
| nystroem158 |
| nystroem159 |
| nystroem160 |
| nystroem161 |
| nystroem162 |
| nystroem163 |
| nystroem164 |
| nystroem165 |
| nystroem166 |
| nystroem167 |
| nystroem168 |
| nystroem169 |
| nystroem170 |
| nystroem171 |
| nystroem172 |
| nystroem173 |
| nystroem174 |
| nystroem175 |
| nystroem176 |
| nystroem177 |
| nystroem178 |
| nystroem179 |
| nystroem180 |
| nystroem181 |
| nystroem182 |
| nystroem183 |
| nystroem184 |
| nystroem185 |
| nystroem186 |
| nystroem187 |
| nystroem188 |
| nystroem189 |
| nystroem190 |
| nystroem191 |
| nystroem192 |
| nystroem193 |
| nystroem194 |
| nystroem195 |
| nystroem196 |
| nystroem197 |
| nystroem198 |
| nystroem199 |
| nystroem200 |
| nystroem201 |
| nystroem202 |
| nystroem203 |
| nystroem204 |
| nystroem205 |
| nystroem206 |
| nystroem207 |
| nystroem208 |
| nystroem209 |
| nystroem210 |
| nystroem211 |
| nystroem212 |
| nystroem213 |
| nystroem214 |
| nystroem215 |
| nystroem216 |
| nystroem217 |
| nystroem218 |
| nystroem219 |
| nystroem220 |
| nystroem221 |
| nystroem222 |
| nystroem223 |
| nystroem224 |
| nystroem225 |
| nystroem226 |
| nystroem227 |
| nystroem228 |
| nystroem229 |
| nystroem230 |
| nystroem231 |
| nystroem232 |
| nystroem233 |
| nystroem234 |
| nystroem235 |
| nystroem236 |
| nystroem237 |
| nystroem238 |
| nystroem239 |
| nystroem240 |
| nystroem241 |
| nystroem242 |
| nystroem243 |
| nystroem244 |
| nystroem245 |
| nystroem246 |
| nystroem247 |
| nystroem248 |
| nystroem249 |
| nystroem250 |
| nystroem251 |
| nystroem252 |
| nystroem253 |
| nystroem254 |
| nystroem255 |
| nystroem256 |
| nystroem257 |
| nystroem258 |
| nystroem259 |
| nystroem260 |
| nystroem261 |
| nystroem262 |
| nystroem263 |
| nystroem264 |
| nystroem265 |
| nystroem266 |
| nystroem267 |
| nystroem268 |
| nystroem269 |
| nystroem270 |
| nystroem271 |
| nystroem272 |
| nystroem273 |
| nystroem274 |
| nystroem275 |
| nystroem276 |
| nystroem277 |
| nystroem278 |
| nystroem279 |
| nystroem280 |
| nystroem281 |
| nystroem282 |
| nystroem283 |
| nystroem284 |
| nystroem285 |
| nystroem286 |
| nystroem287 |
| nystroem288 |
| nystroem289 |
| nystroem290 |
| nystroem291 |
| nystroem292 |
| nystroem293 |
| nystroem294 |
| nystroem295 |
| nystroem296 |
| nystroem297 |
| nystroem298 |
| nystroem299 |
Parameters
Fitted attributes
We observe that this model can almost rival the performance of the gradient boosted trees with an average error around 5% of the maximum demand.
Note that while the final step of this pipeline is a linear regression model, the intermediate steps such as the spline feature extraction and the Nyström kernel approximation are highly non-linear. As a result the compound pipeline is much more expressive than a simple linear regression model with raw features.
For the sake of completeness, we also evaluate the combination of one-hot encoding and kernel approximation:
one_hot_poly_pipeline = make_pipeline(
ColumnTransformer(
transformers=[
("categorical", one_hot_encoder, categorical_columns),
("one_hot_time", one_hot_encoder, ["hour", "weekday", "month"]),
],
remainder="passthrough",
verbose_feature_names_out=False,
),
Nystroem(kernel="poly", degree=2, n_components=300, random_state=0),
RidgeCV(alphas=alphas),
).set_output(transform="pandas")
evaluate(one_hot_poly_pipeline, X, y, cv=ts_cv)
Mean Absolute Error: 0.082 +/- 0.006
Root Mean Squared Error: 0.111 +/- 0.011
Pipeline(steps=[('columntransformer',
ColumnTransformer(remainder='passthrough',
transformers=[('categorical',
OneHotEncoder(handle_unknown='ignore',
sparse_output=False),
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')),
('one_hot_time',
OneHotEncoder(handle_unknown='ignore',
sparse_output=False),
['hour', 'weekday',
'month'])],
verbose_feature...
RidgeCV(alphas=array([1.00000000e-06, 3.16227766e-06, 1.00000000e-05, 3.16227766e-05,
1.00000000e-04, 3.16227766e-04, 1.00000000e-03, 3.16227766e-03,
1.00000000e-02, 3.16227766e-02, 1.00000000e-01, 3.16227766e-01,
1.00000000e+00, 3.16227766e+00, 1.00000000e+01, 3.16227766e+01,
1.00000000e+02, 3.16227766e+02, 1.00000000e+03, 3.16227766e+03,
1.00000000e+04, 3.16227766e+04, 1.00000000e+05, 3.16227766e+05,
1.00000000e+06])))])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Only defined if the underlying estimator exposes such an attribute when fit. .. versionadded:: 1.0 |
ndarray[object](12,) | ['season','year','month',...,'feel_temp','humidity','windspeed'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. Only defined if the underlying first estimator in `steps` exposes such an attribute when fit. .. versionadded:: 0.24 |
int | 12 |
Parameters
Fitted attributes
Index(['season', 'holiday', 'workingday', 'weather'], dtype='str')
Parameters
Fitted attributes
11 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
['hour', 'weekday', 'month']
Parameters
Fitted attributes
43 features
| hour_0 |
| hour_1 |
| hour_2 |
| hour_3 |
| hour_4 |
| hour_5 |
| hour_6 |
| hour_7 |
| hour_8 |
| hour_9 |
| hour_10 |
| hour_11 |
| hour_12 |
| hour_13 |
| hour_14 |
| hour_15 |
| hour_16 |
| hour_17 |
| hour_18 |
| hour_19 |
| hour_20 |
| hour_21 |
| hour_22 |
| hour_23 |
| weekday_0 |
| weekday_1 |
| weekday_2 |
| weekday_3 |
| weekday_4 |
| weekday_5 |
| weekday_6 |
| month_1 |
| month_2 |
| month_3 |
| month_4 |
| month_5 |
| month_6 |
| month_7 |
| month_8 |
| month_9 |
| month_10 |
| month_11 |
| month_12 |
['year', 'temp', 'feel_temp', 'humidity', 'windspeed']
5 features
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
59 features
| season_fall |
| season_spring |
| season_summer |
| season_winter |
| holiday_False |
| holiday_True |
| workingday_False |
| workingday_True |
| weather_clear |
| weather_misty |
| weather_rain |
| hour_0 |
| hour_1 |
| hour_2 |
| hour_3 |
| hour_4 |
| hour_5 |
| hour_6 |
| hour_7 |
| hour_8 |
| hour_9 |
| hour_10 |
| hour_11 |
| hour_12 |
| hour_13 |
| hour_14 |
| hour_15 |
| hour_16 |
| hour_17 |
| hour_18 |
| hour_19 |
| hour_20 |
| hour_21 |
| hour_22 |
| hour_23 |
| weekday_0 |
| weekday_1 |
| weekday_2 |
| weekday_3 |
| weekday_4 |
| weekday_5 |
| weekday_6 |
| month_1 |
| month_2 |
| month_3 |
| month_4 |
| month_5 |
| month_6 |
| month_7 |
| month_8 |
| month_9 |
| month_10 |
| month_11 |
| month_12 |
| year |
| temp |
| feel_temp |
| humidity |
| windspeed |
Parameters
Fitted attributes
| Name | Type | Value |
|---|---|---|
|
component_indices_
component_indices_: ndarray of shape (n_components) Indices of ``components_`` in the training set. |
ndarray[int64](300,) | [9394, 898,2398,...,2685,5725,8051] |
|
components_
components_: ndarray of shape (n_components, n_features) Subset of training points used to construct the feature map. |
ndarray[float64](300, 59) | [[ 0. , 0. , 1. ,...,30.3 , 0.69,19. ], [ 0. , 0. , 1. ,...,25.76, 0.88,11. ], [ 1. , 0. , 0. ,...,37.12, 0.75,17. ], ..., [ 1. , 0. , 0. ,...,32.58, 0.7 , 6. ], [ 0. , 0. , 0. ,...,21.97, 1. , 0. ], [ 0. , 1. , 0. ,...,31.82, 0.37, 7. ]] |
|
feature_names_in_
feature_names_in_: ndarray of shape (`n_features_in_`,) Names of features seen during :term:`fit`. Defined only when `X` has feature names that are all strings. .. versionadded:: 1.0 |
ndarray[object](59,) | ['season_fall','season_spring','season_summer',...,'feel_temp','humidity', 'windspeed'] |
|
n_features_in_
n_features_in_: int Number of features seen during :term:`fit`. .. versionadded:: 0.24 |
int | 59 |
|
normalization_
normalization_: ndarray of shape (n_components, n_components) Normalization matrix needed for embedding. Square root of the kernel matrix on ``components_``. |
ndarray[float64](300, 300) | [[14.11,-0.59, 0.34,..., 0.2 ,-0.49,-0.07], [-0.59,11.24, 0.12,...,-0.03, 0.18,-0.09], [ 0.34, 0.12,11.45,...,-0.43,-0.15, 0.04], ..., [ 0.2 ,-0.03,-0.43,...,11.77,-0.74, 0.35], [-0.49, 0.18,-0.15,...,-0.74,10.65, 0.24], [-0.07,-0.09, 0.04,..., 0.35, 0.24, 8.52]] |
300 features
| nystroem0 |
| nystroem1 |
| nystroem2 |
| nystroem3 |
| nystroem4 |
| nystroem5 |
| nystroem6 |
| nystroem7 |
| nystroem8 |
| nystroem9 |
| nystroem10 |
| nystroem11 |
| nystroem12 |
| nystroem13 |
| nystroem14 |
| nystroem15 |
| nystroem16 |
| nystroem17 |
| nystroem18 |
| nystroem19 |
| nystroem20 |
| nystroem21 |
| nystroem22 |
| nystroem23 |
| nystroem24 |
| nystroem25 |
| nystroem26 |
| nystroem27 |
| nystroem28 |
| nystroem29 |
| nystroem30 |
| nystroem31 |
| nystroem32 |
| nystroem33 |
| nystroem34 |
| nystroem35 |
| nystroem36 |
| nystroem37 |
| nystroem38 |
| nystroem39 |
| nystroem40 |
| nystroem41 |
| nystroem42 |
| nystroem43 |
| nystroem44 |
| nystroem45 |
| nystroem46 |
| nystroem47 |
| nystroem48 |
| nystroem49 |
| nystroem50 |
| nystroem51 |
| nystroem52 |
| nystroem53 |
| nystroem54 |
| nystroem55 |
| nystroem56 |
| nystroem57 |
| nystroem58 |
| nystroem59 |
| nystroem60 |
| nystroem61 |
| nystroem62 |
| nystroem63 |
| nystroem64 |
| nystroem65 |
| nystroem66 |
| nystroem67 |
| nystroem68 |
| nystroem69 |
| nystroem70 |
| nystroem71 |
| nystroem72 |
| nystroem73 |
| nystroem74 |
| nystroem75 |
| nystroem76 |
| nystroem77 |
| nystroem78 |
| nystroem79 |
| nystroem80 |
| nystroem81 |
| nystroem82 |
| nystroem83 |
| nystroem84 |
| nystroem85 |
| nystroem86 |
| nystroem87 |
| nystroem88 |
| nystroem89 |
| nystroem90 |
| nystroem91 |
| nystroem92 |
| nystroem93 |
| nystroem94 |
| nystroem95 |
| nystroem96 |
| nystroem97 |
| nystroem98 |
| nystroem99 |
| nystroem100 |
| nystroem101 |
| nystroem102 |
| nystroem103 |
| nystroem104 |
| nystroem105 |
| nystroem106 |
| nystroem107 |
| nystroem108 |
| nystroem109 |
| nystroem110 |
| nystroem111 |
| nystroem112 |
| nystroem113 |
| nystroem114 |
| nystroem115 |
| nystroem116 |
| nystroem117 |
| nystroem118 |
| nystroem119 |
| nystroem120 |
| nystroem121 |
| nystroem122 |
| nystroem123 |
| nystroem124 |
| nystroem125 |
| nystroem126 |
| nystroem127 |
| nystroem128 |
| nystroem129 |
| nystroem130 |
| nystroem131 |
| nystroem132 |
| nystroem133 |
| nystroem134 |
| nystroem135 |
| nystroem136 |
| nystroem137 |
| nystroem138 |
| nystroem139 |
| nystroem140 |
| nystroem141 |
| nystroem142 |
| nystroem143 |
| nystroem144 |
| nystroem145 |
| nystroem146 |
| nystroem147 |
| nystroem148 |
| nystroem149 |
| nystroem150 |
| nystroem151 |
| nystroem152 |
| nystroem153 |
| nystroem154 |
| nystroem155 |
| nystroem156 |
| nystroem157 |
| nystroem158 |
| nystroem159 |
| nystroem160 |
| nystroem161 |
| nystroem162 |
| nystroem163 |
| nystroem164 |
| nystroem165 |
| nystroem166 |
| nystroem167 |
| nystroem168 |
| nystroem169 |
| nystroem170 |
| nystroem171 |
| nystroem172 |
| nystroem173 |
| nystroem174 |
| nystroem175 |
| nystroem176 |
| nystroem177 |
| nystroem178 |
| nystroem179 |
| nystroem180 |
| nystroem181 |
| nystroem182 |
| nystroem183 |
| nystroem184 |
| nystroem185 |
| nystroem186 |
| nystroem187 |
| nystroem188 |
| nystroem189 |
| nystroem190 |
| nystroem191 |
| nystroem192 |
| nystroem193 |
| nystroem194 |
| nystroem195 |
| nystroem196 |
| nystroem197 |
| nystroem198 |
| nystroem199 |
| nystroem200 |
| nystroem201 |
| nystroem202 |
| nystroem203 |
| nystroem204 |
| nystroem205 |
| nystroem206 |
| nystroem207 |
| nystroem208 |
| nystroem209 |
| nystroem210 |
| nystroem211 |
| nystroem212 |
| nystroem213 |
| nystroem214 |
| nystroem215 |
| nystroem216 |
| nystroem217 |
| nystroem218 |
| nystroem219 |
| nystroem220 |
| nystroem221 |
| nystroem222 |
| nystroem223 |
| nystroem224 |
| nystroem225 |
| nystroem226 |
| nystroem227 |
| nystroem228 |
| nystroem229 |
| nystroem230 |
| nystroem231 |
| nystroem232 |
| nystroem233 |
| nystroem234 |
| nystroem235 |
| nystroem236 |
| nystroem237 |
| nystroem238 |
| nystroem239 |
| nystroem240 |
| nystroem241 |
| nystroem242 |
| nystroem243 |
| nystroem244 |
| nystroem245 |
| nystroem246 |
| nystroem247 |
| nystroem248 |
| nystroem249 |
| nystroem250 |
| nystroem251 |
| nystroem252 |
| nystroem253 |
| nystroem254 |
| nystroem255 |
| nystroem256 |
| nystroem257 |
| nystroem258 |
| nystroem259 |
| nystroem260 |
| nystroem261 |
| nystroem262 |
| nystroem263 |
| nystroem264 |
| nystroem265 |
| nystroem266 |
| nystroem267 |
| nystroem268 |
| nystroem269 |
| nystroem270 |
| nystroem271 |
| nystroem272 |
| nystroem273 |
| nystroem274 |
| nystroem275 |
| nystroem276 |
| nystroem277 |
| nystroem278 |
| nystroem279 |
| nystroem280 |
| nystroem281 |
| nystroem282 |
| nystroem283 |
| nystroem284 |
| nystroem285 |
| nystroem286 |
| nystroem287 |
| nystroem288 |
| nystroem289 |
| nystroem290 |
| nystroem291 |
| nystroem292 |
| nystroem293 |
| nystroem294 |
| nystroem295 |
| nystroem296 |
| nystroem297 |
| nystroem298 |
| nystroem299 |
Parameters
Fitted attributes
While one-hot encoded features were competitive with spline-based features when using linear models, this is no longer the case when using a low-rank approximation of a non-linear kernel: this can be explained by the fact that spline features are smoother and allow the kernel approximation to find a more expressive decision function.
Let us now have a qualitative look at the predictions of the kernel models and of the gradient boosted trees that should be able to better model non-linear interactions between features:
gbrt.fit(X.iloc[train_0], y.iloc[train_0])
gbrt_predictions = gbrt.predict(X.iloc[test_0])
one_hot_poly_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
one_hot_poly_predictions = one_hot_poly_pipeline.predict(X.iloc[test_0])
cyclic_spline_poly_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
cyclic_spline_poly_predictions = cyclic_spline_poly_pipeline.predict(X.iloc[test_0])
Again we zoom on the last 4 days of the test set:
last_hours = slice(-96, None)
fig, ax = plt.subplots(figsize=(12, 4))
fig.suptitle("Predictions by non-linear regression models")
ax.plot(
y.iloc[test_0].values[last_hours],
"x-",
alpha=0.2,
label="Actual demand",
color="black",
)
ax.plot(
gbrt_predictions[last_hours],
"x-",
label="Gradient Boosted Trees",
)
ax.plot(
one_hot_poly_predictions[last_hours],
"x-",
label="One-hot + polynomial kernel",
)
ax.plot(
cyclic_spline_poly_predictions[last_hours],
"x-",
label="Splines + polynomial kernel",
)
_ = ax.legend()
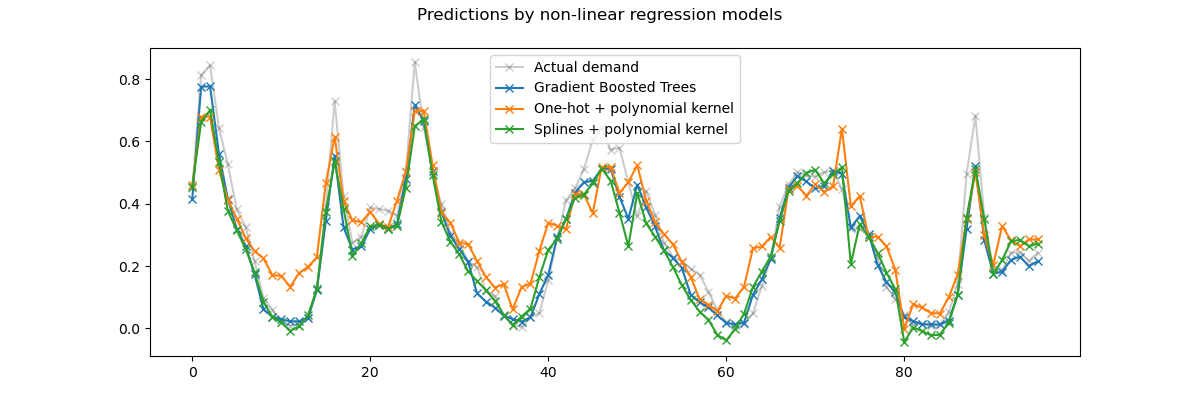
First, note that trees can naturally model non-linear feature interactions since, by default, decision trees are allowed to grow beyond a depth of 2 levels.
Here, we can observe that the combinations of spline features and non-linear kernels works quite well and can almost rival the accuracy of the gradient boosting regression trees.
On the contrary, one-hot encoded time features do not perform that well with the low rank kernel model. In particular, they significantly over-estimate the low demand hours more than the competing models.
We also observe that none of the models can successfully predict some of the peak rentals at the rush hours during the working days. It is possible that access to additional features would be required to further improve the accuracy of the predictions. For instance, it could be useful to have access to the geographical repartition of the fleet at any point in time or the fraction of bikes that are immobilized because they need servicing.
Let us finally get a more quantitative look at the prediction errors of those three models using the true vs predicted demand scatter plots:
from sklearn.metrics import PredictionErrorDisplay
fig, axes = plt.subplots(nrows=2, ncols=3, figsize=(13, 7), sharex=True, sharey="row")
fig.suptitle("Non-linear regression models", y=1.0)
predictions = [
one_hot_poly_predictions,
cyclic_spline_poly_predictions,
gbrt_predictions,
]
labels = [
"One hot +\npolynomial kernel",
"Splines +\npolynomial kernel",
"Gradient Boosted\nTrees",
]
plot_kinds = ["actual_vs_predicted", "residual_vs_predicted"]
for axis_idx, kind in enumerate(plot_kinds):
for ax, pred, label in zip(axes[axis_idx], predictions, labels):
disp = PredictionErrorDisplay.from_predictions(
y_true=y.iloc[test_0],
y_pred=pred,
kind=kind,
scatter_kwargs={"alpha": 0.3},
ax=ax,
)
ax.set_xticks(np.linspace(0, 1, num=5))
if axis_idx == 0:
ax.set_yticks(np.linspace(0, 1, num=5))
ax.legend(
["Best model", label],
loc="upper center",
bbox_to_anchor=(0.5, 1.3),
ncol=2,
)
ax.set_aspect("equal", adjustable="box")
plt.show()
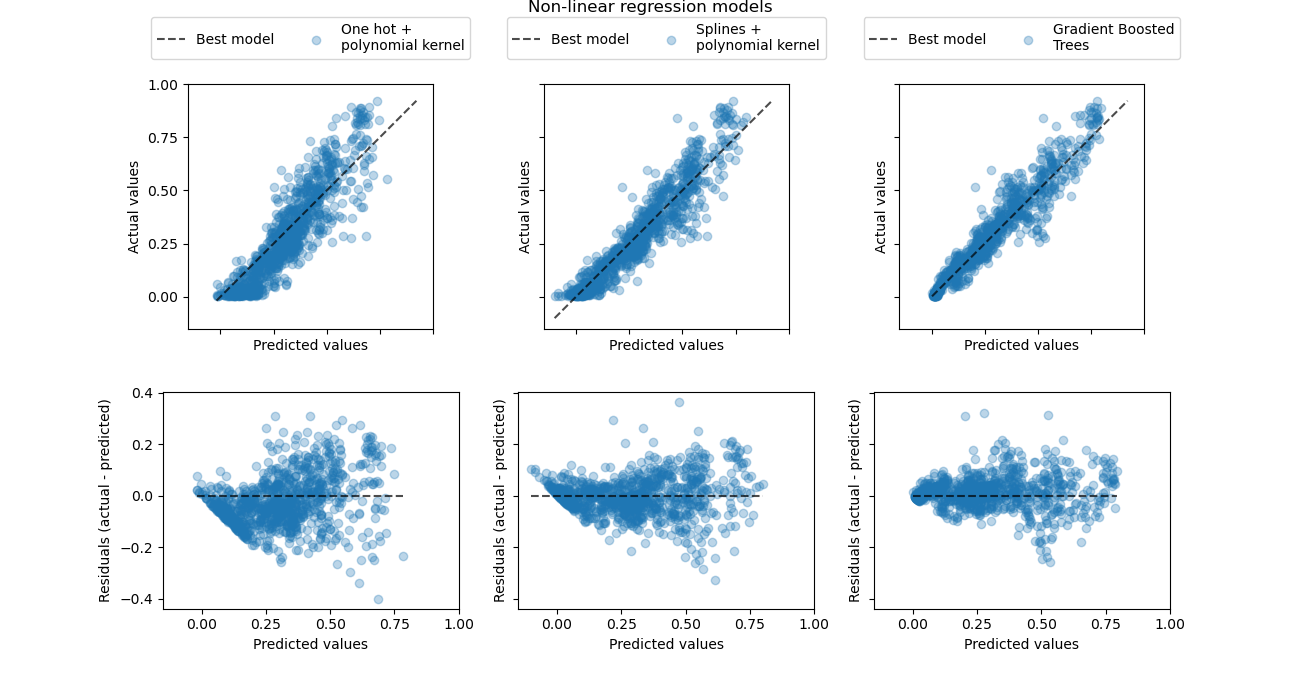
This visualization confirms the conclusions we draw on the previous plot.
All models under-estimate the high demand events (working day rush hours), but gradient boosting a bit less so. The low demand events are well predicted on average by gradient boosting while the one-hot polynomial regression pipeline seems to systematically over-estimate demand in that regime. Overall the predictions of the gradient boosted trees are closer to the diagonal than for the kernel models.
Concluding remarks#
We note that we could have obtained slightly better results for kernel models
by using more components (higher rank kernel approximation) at the cost of
longer fit and prediction durations. For large values of n_components, the
performance of the one-hot encoded features would even match the spline
features.
The Nystroem + RidgeCV regressor could also have been replaced by
MLPRegressor with one or two hidden layers
and we would have obtained quite similar results.
The dataset we used in this case study is sampled on an hourly basis. However cyclic spline-based features could model time-within-day or time-within-week very efficiently with finer-grained time resolutions (for instance with measurements taken every minute instead of every hour) without introducing more features. One-hot encoding time representations would not offer this flexibility.
Finally, in this notebook we used RidgeCV because it is very efficient from
a computational point of view. However, it models the target variable as a
Gaussian random variable with constant variance. For positive regression
problems, it is likely that using a Poisson or Gamma distribution would make
more sense. This could be achieved by using
GridSearchCV(TweedieRegressor(power=2), param_grid({"alpha": alphas}))
instead of RidgeCV.
Total running time of the script: (0 minutes 14.944 seconds)
Related examples