Note
Go to the end to download the full example code or to run this example in your browser via JupyterLite or Binder.
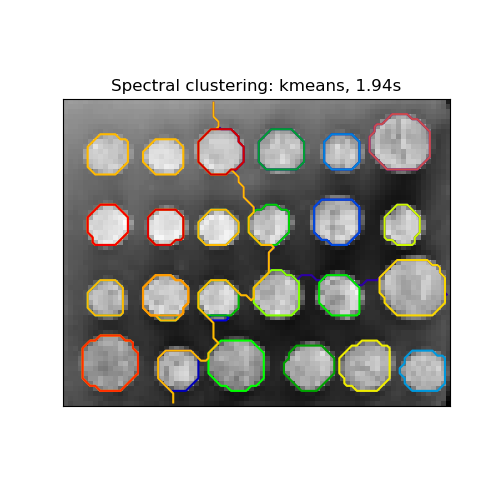
Segmenting the picture of greek coins in regions#
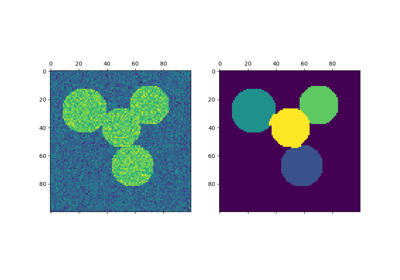
This example uses Spectral clustering on a graph created from voxel-to-voxel difference on an image to break this image into multiple partly-homogeneous regions.
This procedure (spectral clustering on an image) is an efficient approximate solution for finding normalized graph cuts.
There are three options to assign labels:
‘kmeans’ spectral clustering clusters samples in the embedding space using a kmeans algorithm
‘discrete’ iteratively searches for the closest partition space to the embedding space of spectral clustering.
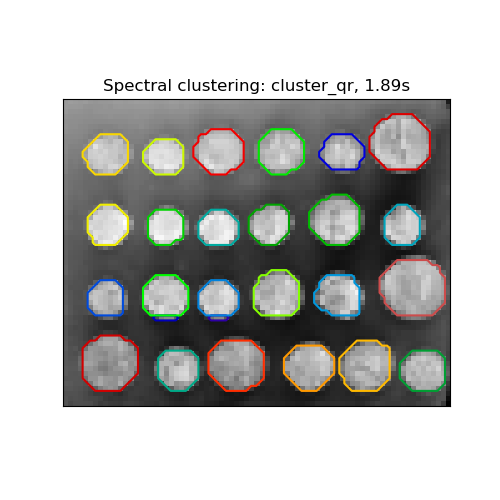
‘cluster_qr’ assigns labels using the QR factorization with pivoting that directly determines the partition in the embedding space.
# Authors: The scikit-learn developers
# SPDX-License-Identifier: BSD-3-Clause
import time
import matplotlib.pyplot as plt
import numpy as np
from scipy.ndimage import gaussian_filter
from skimage.data import coins
from skimage.transform import rescale
from sklearn.cluster import spectral_clustering
from sklearn.feature_extraction import image
# load the coins as a numpy array
orig_coins = coins()
# Resize it to 20% of the original size to speed up the processing
# Applying a Gaussian filter for smoothing prior to down-scaling
# reduces aliasing artifacts.
smoothened_coins = gaussian_filter(orig_coins, sigma=2)
rescaled_coins = rescale(smoothened_coins, 0.2, mode="reflect", anti_aliasing=False)
# Convert the image into a graph with the value of the gradient on the
# edges.
graph = image.img_to_graph(rescaled_coins)
# Take a decreasing function of the gradient: an exponential
# The smaller beta is, the more independent the segmentation is of the
# actual image. For beta=1, the segmentation is close to a voronoi
beta = 10
eps = 1e-6
graph.data = np.exp(-beta * graph.data / graph.data.std()) + eps
# The number of segmented regions to display needs to be chosen manually.
# The current version of 'spectral_clustering' does not support determining
# the number of good quality clusters automatically.
n_regions = 26
Compute and visualize the resulting regions
# Computing a few extra eigenvectors may speed up the eigen_solver.
# The spectral clustering quality may also benefit from requesting
# extra regions for segmentation.
n_regions_plus = 3
# Apply spectral clustering using the default eigen_solver='arpack'.
# Any implemented solver can be used: eigen_solver='arpack', 'lobpcg', or 'amg'.
# Choosing eigen_solver='amg' requires an extra package called 'pyamg'.
# The quality of segmentation and the speed of calculations is mostly determined
# by the choice of the solver and the value of the tolerance 'eigen_tol'.
# TODO: varying eigen_tol seems to have no effect for 'lobpcg' and 'amg' #21243.
for assign_labels in ("kmeans", "discretize", "cluster_qr"):
t0 = time.time()
labels = spectral_clustering(
graph,
n_clusters=(n_regions + n_regions_plus),
eigen_tol=1e-7,
assign_labels=assign_labels,
random_state=42,
)
t1 = time.time()
labels = labels.reshape(rescaled_coins.shape)
plt.figure(figsize=(5, 5))
plt.imshow(rescaled_coins, cmap=plt.cm.gray)
plt.xticks(())
plt.yticks(())
title = "Spectral clustering: %s, %.2fs" % (assign_labels, (t1 - t0))
print(title)
plt.title(title)
for l in range(n_regions):
colors = [plt.cm.nipy_spectral((l + 4) / float(n_regions + 4))]
plt.contour(labels == l, colors=colors)
# To view individual segments as appear comment in plt.pause(0.5)
plt.show()
# TODO: After #21194 is merged and #21243 is fixed, check which eigen_solver
# is the best and set eigen_solver='arpack', 'lobpcg', or 'amg' and eigen_tol
# explicitly in this example.
Spectral clustering: kmeans, 0.19s
Spectral clustering: discretize, 0.08s
Spectral clustering: cluster_qr, 0.06s
Total running time of the script: (0 minutes 0.686 seconds)
Related examples

A demo of structured Ward hierarchical clustering on an image of coins