Note
Go to the end to download the full example code. or to run this example in your browser via JupyterLite or Binder
SVM Exercise#
A tutorial exercise for using different SVM kernels.
This exercise is used in the using_kernels_tut part of the supervised_learning_tut section of the stat_learn_tut_index.
# Authors: The scikit-learn developers
# SPDX-License-Identifier: BSD-3-Clause
import matplotlib.pyplot as plt
import numpy as np
from sklearn import datasets, svm
iris = datasets.load_iris()
X = iris.data
y = iris.target
X = X[y != 0, :2]
y = y[y != 0]
n_sample = len(X)
np.random.seed(0)
order = np.random.permutation(n_sample)
X = X[order]
y = y[order].astype(float)
X_train = X[: int(0.9 * n_sample)]
y_train = y[: int(0.9 * n_sample)]
X_test = X[int(0.9 * n_sample) :]
y_test = y[int(0.9 * n_sample) :]
# fit the model
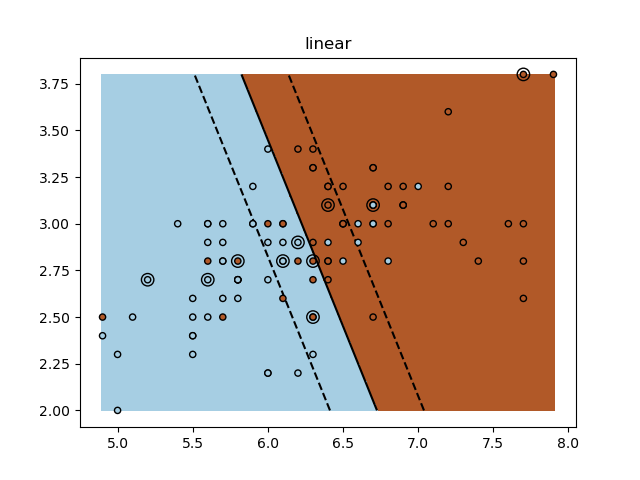
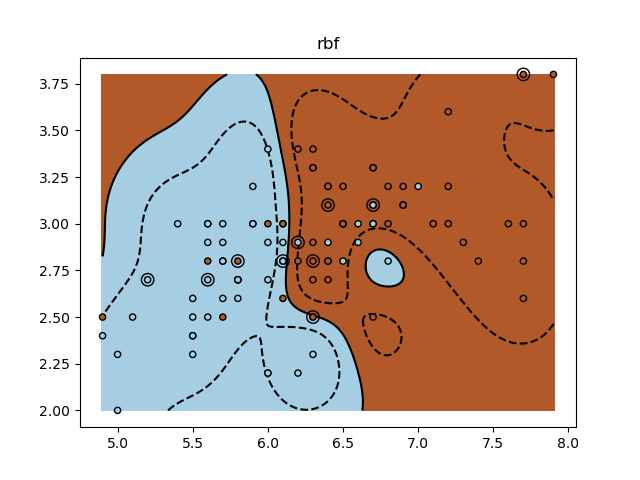
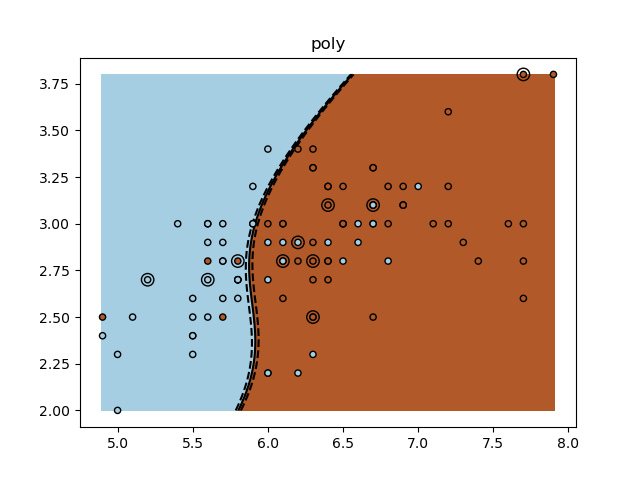
for kernel in ("linear", "rbf", "poly"):
clf = svm.SVC(kernel=kernel, gamma=10)
clf.fit(X_train, y_train)
plt.figure()
plt.clf()
plt.scatter(
X[:, 0], X[:, 1], c=y, zorder=10, cmap=plt.cm.Paired, edgecolor="k", s=20
)
# Circle out the test data
plt.scatter(
X_test[:, 0], X_test[:, 1], s=80, facecolors="none", zorder=10, edgecolor="k"
)
plt.axis("tight")
x_min = X[:, 0].min()
x_max = X[:, 0].max()
y_min = X[:, 1].min()
y_max = X[:, 1].max()
XX, YY = np.mgrid[x_min:x_max:200j, y_min:y_max:200j]
Z = clf.decision_function(np.c_[XX.ravel(), YY.ravel()])
# Put the result into a color plot
Z = Z.reshape(XX.shape)
plt.pcolormesh(XX, YY, Z > 0, cmap=plt.cm.Paired)
plt.contour(
XX,
YY,
Z,
colors=["k", "k", "k"],
linestyles=["--", "-", "--"],
levels=[-0.5, 0, 0.5],
)
plt.title(kernel)
plt.show()
Total running time of the script: (0 minutes 5.711 seconds)
Related examples
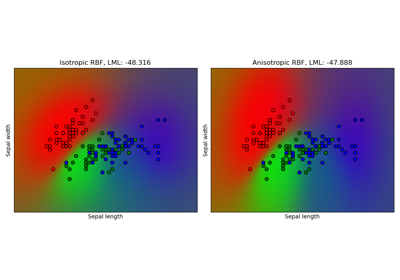
Gaussian process classification (GPC) on iris dataset
Gaussian process classification (GPC) on iris dataset
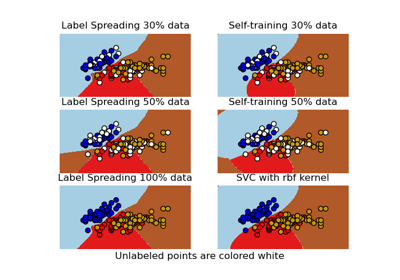
Decision boundary of semi-supervised classifiers versus SVM on the Iris dataset
Decision boundary of semi-supervised classifiers versus SVM on the Iris dataset