SVC#
- class sklearn.svm.SVC(*, C=1.0, kernel='rbf', degree=3, gamma='scale', coef0=0.0, shrinking=True, probability=False, tol=0.001, cache_size=200, class_weight=None, verbose=False, max_iter=-1, decision_function_shape='ovr', break_ties=False, random_state=None)[source]#
C-Support Vector Classification.
The implementation is based on libsvm. The fit time scales at least quadratically with the number of samples and may be impractical beyond tens of thousands of samples. For large datasets consider using
LinearSVCorSGDClassifierinstead, possibly after aNystroemtransformer or other Kernel Approximation.The multiclass support is handled according to a one-vs-one scheme.
For details on the precise mathematical formulation of the provided kernel functions and how
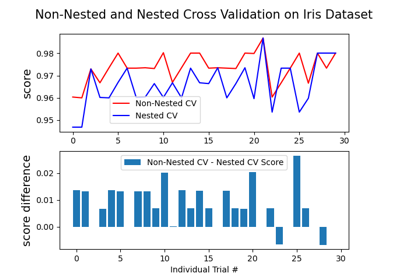
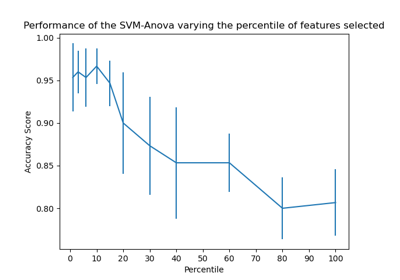
gamma,coef0anddegreeaffect each other, see the corresponding section in the narrative documentation: Kernel functions.To learn how to tune SVC’s hyperparameters, see the following example: Nested versus non-nested cross-validation
Read more in the User Guide.
- Parameters:
- Cfloat, default=1.0
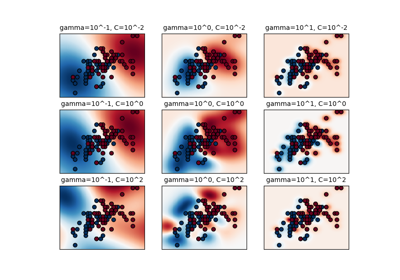
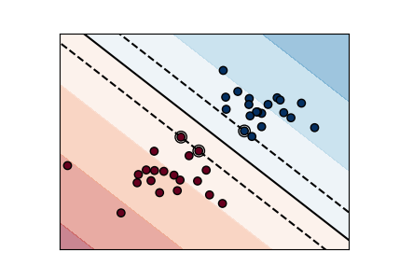
Regularization parameter. The strength of the regularization is inversely proportional to C. Must be strictly positive. The penalty is a squared l2 penalty. For an intuitive visualization of the effects of scaling the regularization parameter C, see Scaling the regularization parameter for SVCs.
- kernel{‘linear’, ‘poly’, ‘rbf’, ‘sigmoid’, ‘precomputed’} or callable, default=’rbf’
Specifies the kernel type to be used in the algorithm. If none is given, ‘rbf’ will be used. If a callable is given it is used to pre-compute the kernel matrix from data matrices; that matrix should be an array of shape
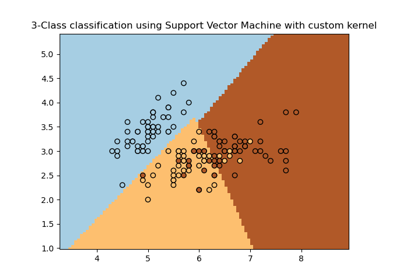
(n_samples, n_samples). For an intuitive visualization of different kernel types see Plot classification boundaries with different SVM Kernels.- degreeint, default=3
Degree of the polynomial kernel function (‘poly’). Must be non-negative. Ignored by all other kernels.
- gamma{‘scale’, ‘auto’} or float, default=’scale’
Kernel coefficient for ‘rbf’, ‘poly’ and ‘sigmoid’.
if
gamma='scale'(default) is passed then it uses 1 / (n_features * X.var()) as value of gamma,if ‘auto’, uses 1 / n_features
if float, must be non-negative.
Changed in version 0.22: The default value of
gammachanged from ‘auto’ to ‘scale’.- coef0float, default=0.0
Independent term in kernel function. It is only significant in ‘poly’ and ‘sigmoid’.
- shrinkingbool, default=True
Whether to use the shrinking heuristic. See the User Guide.
- probabilitybool, default=False
Whether to enable probability estimates. This must be enabled prior to calling
fit, will slow down that method as it internally uses 5-fold cross-validation, andpredict_probamay be inconsistent withpredict. Read more in the User Guide.- tolfloat, default=1e-3
Tolerance for stopping criterion.
- cache_sizefloat, default=200
Specify the size of the kernel cache (in MB).
- class_weightdict or ‘balanced’, default=None
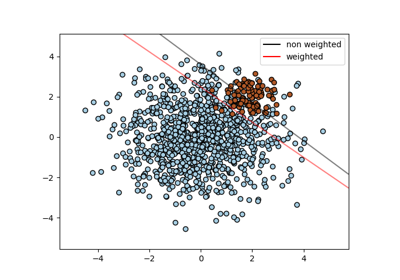
Set the parameter C of class i to class_weight[i]*C for SVC. If not given, all classes are supposed to have weight one. The “balanced” mode uses the values of y to automatically adjust weights inversely proportional to class frequencies in the input data as
n_samples / (n_classes * np.bincount(y)).- verbosebool, default=False
Enable verbose output. Note that this setting takes advantage of a per-process runtime setting in libsvm that, if enabled, may not work properly in a multithreaded context.
- max_iterint, default=-1
Hard limit on iterations within solver, or -1 for no limit.
- decision_function_shape{‘ovo’, ‘ovr’}, default=’ovr’
Whether to return a one-vs-rest (‘ovr’) decision function of shape (n_samples, n_classes) as all other classifiers, or the original one-vs-one (‘ovo’) decision function of libsvm which has shape (n_samples, n_classes * (n_classes - 1) / 2). However, note that internally, one-vs-one (‘ovo’) is always used as a multi-class strategy to train models; an ovr matrix is only constructed from the ovo matrix. The parameter is ignored for binary classification.
Changed in version 0.19: decision_function_shape is ‘ovr’ by default.
Added in version 0.17: decision_function_shape=’ovr’ is recommended.
Changed in version 0.17: Deprecated decision_function_shape=’ovo’ and None.
- break_tiesbool, default=False
If true,
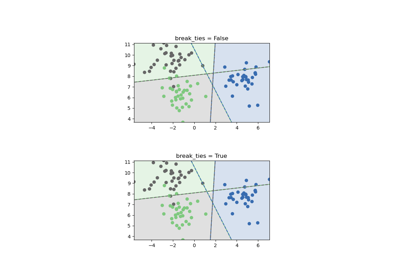
decision_function_shape='ovr', and number of classes > 2, predict will break ties according to the confidence values of decision_function; otherwise the first class among the tied classes is returned. Please note that breaking ties comes at a relatively high computational cost compared to a simple predict. See SVM Tie Breaking Example for an example of its usage withdecision_function_shape='ovr'.Added in version 0.22.
- random_stateint, RandomState instance or None, default=None
Controls the pseudo random number generation for shuffling the data for probability estimates. Ignored when
probabilityis False. Pass an int for reproducible output across multiple function calls. See Glossary.
- Attributes:
- class_weight_ndarray of shape (n_classes,)
Multipliers of parameter C for each class. Computed based on the
class_weightparameter.- classes_ndarray of shape (n_classes,)
The classes labels.
coef_ndarray of shape (n_classes * (n_classes - 1) / 2, n_features)Weights assigned to the features when
kernel="linear".- dual_coef_ndarray of shape (n_classes -1, n_SV)
Dual coefficients of the support vector in the decision function (see Mathematical formulation), multiplied by their targets. For multiclass, coefficient for all 1-vs-1 classifiers. The layout of the coefficients in the multiclass case is somewhat non-trivial. See the multi-class section of the User Guide for details.
- fit_status_int
0 if correctly fitted, 1 otherwise (will raise warning)
- intercept_ndarray of shape (n_classes * (n_classes - 1) / 2,)
Constants in decision function.
- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
- n_iter_ndarray of shape (n_classes * (n_classes - 1) // 2,)
Number of iterations run by the optimization routine to fit the model. The shape of this attribute depends on the number of models optimized which in turn depends on the number of classes.
Added in version 1.1.
- support_ndarray of shape (n_SV)
Indices of support vectors.
- support_vectors_ndarray of shape (n_SV, n_features)
Support vectors. An empty array if kernel is precomputed.
n_support_ndarray of shape (n_classes,), dtype=int32Number of support vectors for each class.
probA_ndarray of shape (n_classes * (n_classes - 1) / 2)Parameter learned in Platt scaling when
probability=True.probB_ndarray of shape (n_classes * (n_classes - 1) / 2)Parameter learned in Platt scaling when
probability=True.- shape_fit_tuple of int of shape (n_dimensions_of_X,)
Array dimensions of training vector
X.
See also
References
Examples
>>> import numpy as np >>> from sklearn.pipeline import make_pipeline >>> from sklearn.preprocessing import StandardScaler >>> X = np.array([[-1, -1], [-2, -1], [1, 1], [2, 1]]) >>> y = np.array([1, 1, 2, 2]) >>> from sklearn.svm import SVC >>> clf = make_pipeline(StandardScaler(), SVC(gamma='auto')) >>> clf.fit(X, y) Pipeline(steps=[('standardscaler', StandardScaler()), ('svc', SVC(gamma='auto'))])
>>> print(clf.predict([[-0.8, -1]])) [1]
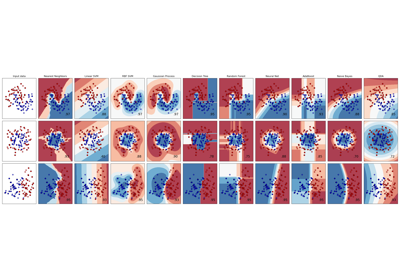
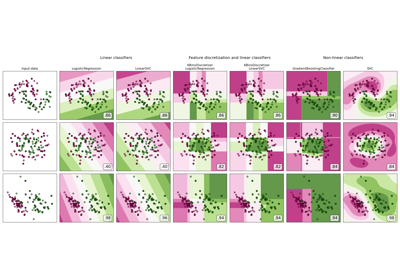
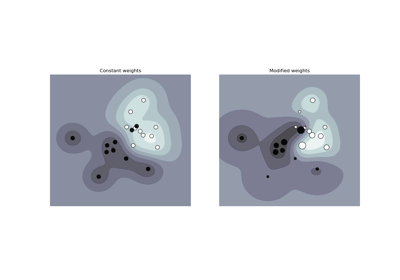
For a comparison of the SVC with other classifiers see: Plot classification probability.
- decision_function(X)[source]#
Evaluate the decision function for the samples in X.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
The input samples.
- Returns:
- Xndarray of shape (n_samples, n_classes * (n_classes-1) / 2)
Returns the decision function of the sample for each class in the model. If decision_function_shape=’ovr’, the shape is (n_samples, n_classes).
Notes
If decision_function_shape=’ovo’, the function values are proportional to the distance of the samples X to the separating hyperplane. If the exact distances are required, divide the function values by the norm of the weight vector (
coef_). See also this question for further details. If decision_function_shape=’ovr’, the decision function is a monotonic transformation of ovo decision function.
- fit(X, y, sample_weight=None)[source]#
Fit the SVM model according to the given training data.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features) or (n_samples, n_samples)
Training vectors, where
n_samplesis the number of samples andn_featuresis the number of features. For kernel=”precomputed”, the expected shape of X is (n_samples, n_samples).- yarray-like of shape (n_samples,)
Target values (class labels in classification, real numbers in regression).
- sample_weightarray-like of shape (n_samples,), default=None
Per-sample weights. Rescale C per sample. Higher weights force the classifier to put more emphasis on these points.
- Returns:
- selfobject
Fitted estimator.
Notes
If X and y are not C-ordered and contiguous arrays of np.float64 and X is not a scipy.sparse.csr_matrix, X and/or y may be copied.
If X is a dense array, then the other methods will not support sparse matrices as input.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- predict(X)[source]#
Perform classification on samples in X.
For a one-class model, +1 or -1 is returned.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features) or (n_samples_test, n_samples_train)
For kernel=”precomputed”, the expected shape of X is (n_samples_test, n_samples_train).
- Returns:
- y_predndarray of shape (n_samples,)
Class labels for samples in X.
- predict_log_proba(X)[source]#
Compute log probabilities of possible outcomes for samples in X.
The model need to have probability information computed at training time: fit with attribute
probabilityset to True.- Parameters:
- Xarray-like of shape (n_samples, n_features) or (n_samples_test, n_samples_train)
For kernel=”precomputed”, the expected shape of X is (n_samples_test, n_samples_train).
- Returns:
- Tndarray of shape (n_samples, n_classes)
Returns the log-probabilities of the sample for each class in the model. The columns correspond to the classes in sorted order, as they appear in the attribute classes_.
Notes
The probability model is created using cross validation, so the results can be slightly different than those obtained by predict. Also, it will produce meaningless results on very small datasets.
- predict_proba(X)[source]#
Compute probabilities of possible outcomes for samples in X.
The model needs to have probability information computed at training time: fit with attribute
probabilityset to True.- Parameters:
- Xarray-like of shape (n_samples, n_features)
For kernel=”precomputed”, the expected shape of X is (n_samples_test, n_samples_train).
- Returns:
- Tndarray of shape (n_samples, n_classes)
Returns the probability of the sample for each class in the model. The columns correspond to the classes in sorted order, as they appear in the attribute classes_.
Notes
The probability model is created using cross validation, so the results can be slightly different than those obtained by predict. Also, it will produce meaningless results on very small datasets.
- score(X, y, sample_weight=None)[source]#
Return accuracy on provided data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True labels for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
Mean accuracy of
self.predict(X)w.r.t.y.
- set_fit_request(*, sample_weight: bool | None | str = '$UNCHANGED$') SVC[source]#
Configure whether metadata should be requested to be passed to the
fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter infit.
- Returns:
- selfobject
The updated object.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') SVC[source]#
Configure whether metadata should be requested to be passed to the
scoremethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.
Gallery examples#
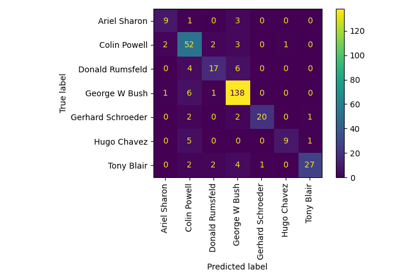
Faces recognition example using eigenfaces and SVMs
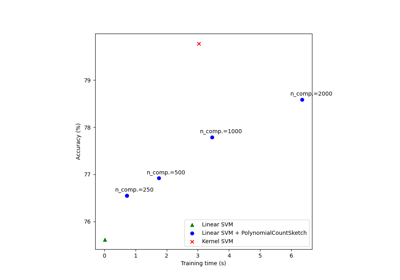
Scalable learning with polynomial kernel approximation
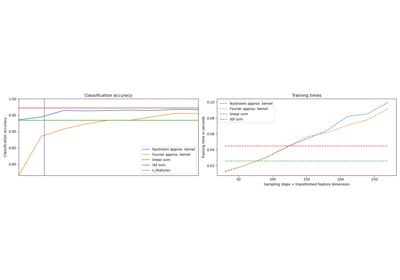
Explicit feature map approximation for RBF kernels
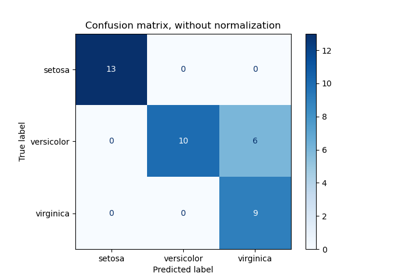
Evaluate the performance of a classifier with Confusion Matrix

Custom refit strategy of a grid search with cross-validation
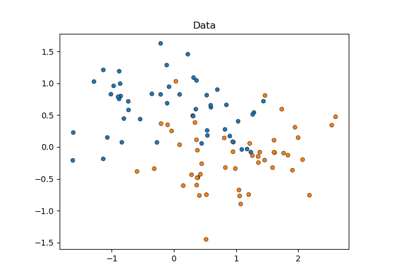
Statistical comparison of models using grid search
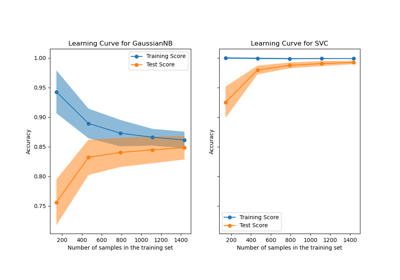
Plotting Learning Curves and Checking Models’ Scalability
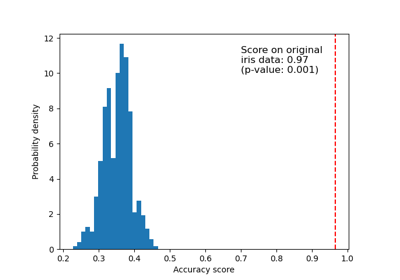
Test with permutations the significance of a classification score
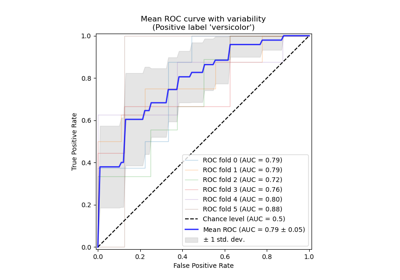
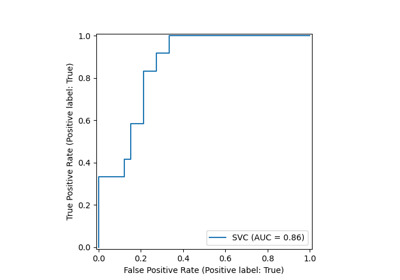
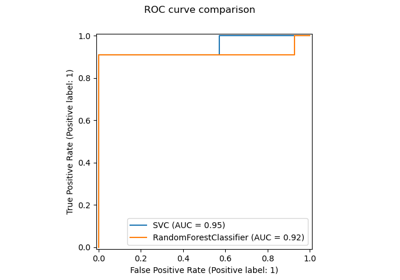
Receiver Operating Characteristic (ROC) with cross validation
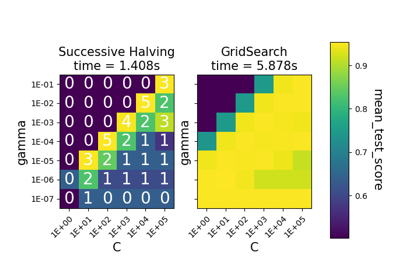
Comparison between grid search and successive halving
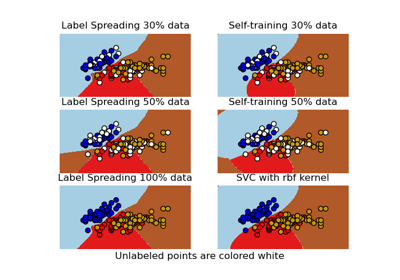
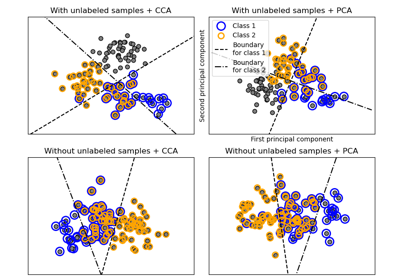
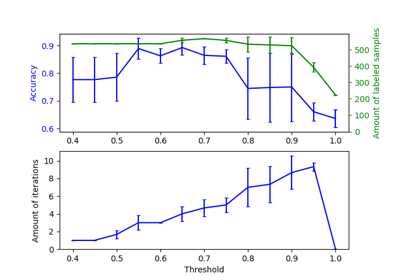
Decision boundary of semi-supervised classifiers versus SVM on the Iris dataset
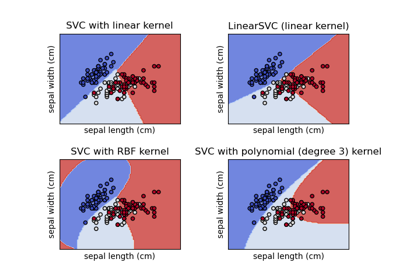
Plot different SVM classifiers in the iris dataset
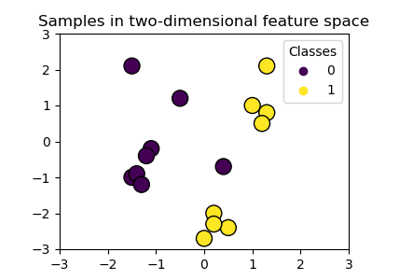
Plot classification boundaries with different SVM Kernels