StratifiedShuffleSplit#
- class sklearn.model_selection.StratifiedShuffleSplit(n_splits=10, *, test_size=None, train_size=None, random_state=None)[source]#
Class-wise stratified ShuffleSplit cross-validator.
Provides train/test indices to split data in train/test sets.
This cross-validation object is a merge of
StratifiedKFoldandShuffleSplit, which returns stratified randomized folds. The folds are made by preserving the percentage of samples for each class inyin a binary or multiclass classification setting.Note: like the
ShuffleSplitstrategy, stratified random splits do not guarantee that test sets across all folds will be mutually exclusive, and might include overlapping samples. However, this is still very likely for sizeable datasets.Read more in the User Guide.
For visualisation of cross-validation behaviour and comparison between common scikit-learn split methods refer to Visualizing cross-validation behavior in scikit-learn
Note
Stratification on the class label solves an engineering problem rather than a statistical one. See Cross-validation iterators with stratification based on class labels for more details.
- Parameters:
- n_splitsint, default=10
Number of re-shuffling & splitting iterations.
- test_sizefloat or int, default=None
If float, should be between 0.0 and 1.0 and represent the proportion of the dataset to include in the test split. If int, represents the absolute number of test samples. If None, the value is set to the complement of the train size. If
train_sizeis also None, it will be set to 0.1.- train_sizefloat or int, default=None
If float, should be between 0.0 and 1.0 and represent the proportion of the dataset to include in the train split. If int, represents the absolute number of train samples. If None, the value is automatically set to the complement of the test size.
- random_stateint, RandomState instance or None, default=None
Controls the randomness of the training and testing indices produced. Pass an int for reproducible output across multiple function calls. See Glossary.
Examples
>>> import numpy as np >>> from sklearn.model_selection import StratifiedShuffleSplit >>> X = np.array([[1, 2], [3, 4], [1, 2], [3, 4], [1, 2], [3, 4]]) >>> y = np.array([0, 0, 0, 1, 1, 1]) >>> sss = StratifiedShuffleSplit(n_splits=5, test_size=0.5, random_state=0) >>> sss.get_n_splits() 5 >>> print(sss) StratifiedShuffleSplit(n_splits=5, random_state=0, ...) >>> for i, (train_index, test_index) in enumerate(sss.split(X, y)): ... print(f"Fold {i}:") ... print(f" Train: index={train_index}") ... print(f" Test: index={test_index}") Fold 0: Train: index=[5 2 3] Test: index=[4 1 0] Fold 1: Train: index=[5 1 4] Test: index=[0 2 3] Fold 2: Train: index=[5 0 2] Test: index=[4 3 1] Fold 3: Train: index=[4 1 0] Test: index=[2 3 5] Fold 4: Train: index=[0 5 1] Test: index=[3 4 2]
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_n_splits(X=None, y=None, groups=None)[source]#
Returns the number of splitting iterations as set with the
n_splitsparam when instantiating the cross-validator.- Parameters:
- Xarray-like of shape (n_samples, n_features), default=None
Always ignored, exists for API compatibility.
- yarray-like of shape (n_samples,), default=None
Always ignored, exists for API compatibility.
- groupsarray-like of shape (n_samples,), default=None
Always ignored, exists for API compatibility.
- Returns:
- n_splitsint
Returns the number of splitting iterations in the cross-validator.
- split(X, y, groups=None)[source]#
Generate indices to split data into training and test set.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Training data, where
n_samplesis the number of samples andn_featuresis the number of features.Note that providing
yis sufficient to generate the splits and hencenp.zeros(n_samples)may be used as a placeholder forXinstead of actual training data.- yarray-like of shape (n_samples,) or (n_samples, n_labels)
The target variable for supervised learning problems. Stratification is done based on the y labels.
- groupsarray-like of shape (n_samples,), default=None
Always ignored, exists for API compatibility.
- Yields:
- trainndarray
The training set indices for that split.
- testndarray
The testing set indices for that split.
Notes
Randomized CV splitters may return different results for each call of split. You can make the results identical by setting
random_stateto an integer.
Gallery examples#
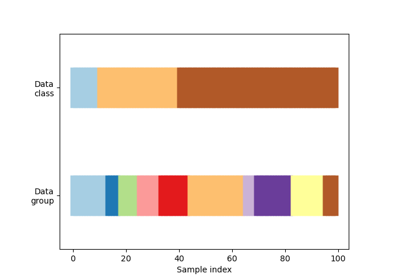
Visualizing cross-validation behavior in scikit-learn