AgglomerativeClustering#
- class sklearn.cluster.AgglomerativeClustering(n_clusters=2, *, metric='euclidean', memory=None, connectivity=None, compute_full_tree='auto', linkage='ward', distance_threshold=None, compute_distances=False)[source]#
Agglomerative Clustering.
Recursively merges pair of clusters of sample data; uses linkage distance.
Read more in the User Guide.
- Parameters:
- n_clustersint or None, default=2
The number of clusters to find. It must be
Noneifdistance_thresholdis notNone.- metricstr or callable, default=”euclidean”
Metric used to compute the linkage. Can be “euclidean”, “l1”, “l2”, “manhattan”, “cosine”, or “precomputed”. If linkage is “ward”, only “euclidean” is accepted. If “precomputed”, a distance matrix is needed as input for the fit method. If connectivity is None, linkage is “single” and affinity is not “precomputed” any valid pairwise distance metric can be assigned.
For an example of agglomerative clustering with different metrics, see Agglomerative clustering with different metrics.
Added in version 1.2.
- memorystr or object with the joblib.Memory interface, default=None
Used to cache the output of the computation of the tree. By default, no caching is done. If a string is given, it is the path to the caching directory.
- connectivityarray-like, sparse matrix, or callable, default=None
Connectivity matrix. Defines for each sample the neighboring samples following a given structure of the data. This can be a connectivity matrix itself or a callable that transforms the data into a connectivity matrix, such as derived from
kneighbors_graph. Default isNone, i.e, the hierarchical clustering algorithm is unstructured.For an example of connectivity matrix using
kneighbors_graph, see Hierarchical clustering with and without structure.- compute_full_tree‘auto’ or bool, default=’auto’
Stop early the construction of the tree at
n_clusters. This is useful to decrease computation time if the number of clusters is not small compared to the number of samples. This option is useful only when specifying a connectivity matrix. Note also that when varying the number of clusters and using caching, it may be advantageous to compute the full tree. It must beTrueifdistance_thresholdis notNone. By defaultcompute_full_treeis “auto”, which is equivalent toTruewhendistance_thresholdis notNoneor thatn_clustersis inferior to the maximum between 100 or0.02 * n_samples. Otherwise, “auto” is equivalent toFalse.- linkage{‘ward’, ‘complete’, ‘average’, ‘single’}, default=’ward’
Which linkage criterion to use. The linkage criterion determines which distance to use between sets of observation. The algorithm will merge the pairs of cluster that minimize this criterion.
‘ward’ minimizes the variance of the clusters being merged.
‘average’ uses the average of the distances of each observation of the two sets.
‘complete’ or ‘maximum’ linkage uses the maximum distances between all observations of the two sets.
‘single’ uses the minimum of the distances between all observations of the two sets.
Added in version 0.20: Added the ‘single’ option
For examples comparing different
linkagecriteria, see Comparing different hierarchical linkage methods on toy datasets.- distance_thresholdfloat, default=None
The linkage distance threshold at or above which clusters will not be merged. If not
None,n_clustersmust beNoneandcompute_full_treemust beTrue.Added in version 0.21.
- compute_distancesbool, default=False
Computes distances between clusters even if
distance_thresholdis not used. This can be used to make dendrogram visualization, but introduces a computational and memory overhead.Added in version 0.24.
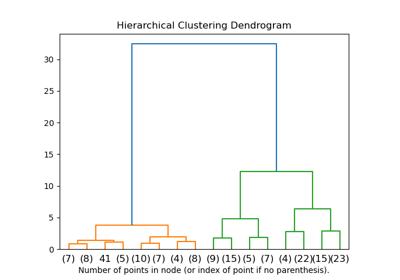
For an example of dendrogram visualization, see Plot Hierarchical Clustering Dendrogram.
- Attributes:
- n_clusters_int
The number of clusters found by the algorithm. If
distance_threshold=None, it will be equal to the givenn_clusters.- labels_ndarray of shape (n_samples)
Cluster labels for each point.
- n_leaves_int
Number of leaves in the hierarchical tree.
- n_connected_components_int
The estimated number of connected components in the graph.
Added in version 0.21:
n_connected_components_was added to replacen_components_.- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
- children_array-like of shape (n_samples-1, 2)
The children of each non-leaf node. Values less than
n_samplescorrespond to leaves of the tree which are the original samples. A nodeigreater than or equal ton_samplesis a non-leaf node and has childrenchildren_[i - n_samples]. Alternatively at the i-th iteration, children[i][0] and children[i][1] are merged to form noden_samples + i.- distances_array-like of shape (n_nodes-1,)
Distances between nodes in the corresponding place in
children_. Only computed ifdistance_thresholdis used orcompute_distancesis set toTrue.
See also
FeatureAgglomerationAgglomerative clustering but for features instead of samples.
ward_treeHierarchical clustering with ward linkage.
Examples
>>> from sklearn.cluster import AgglomerativeClustering >>> import numpy as np >>> X = np.array([[1, 2], [1, 4], [1, 0], ... [4, 2], [4, 4], [4, 0]]) >>> clustering = AgglomerativeClustering().fit(X) >>> clustering AgglomerativeClustering() >>> clustering.labels_ array([1, 1, 1, 0, 0, 0])
For a comparison of Agglomerative clustering with other clustering algorithms, see Comparing different clustering algorithms on toy datasets
- fit(X, y=None)[source]#
Fit the hierarchical clustering from features, or distance matrix.
- Parameters:
- Xarray-like, shape (n_samples, n_features) or (n_samples, n_samples)
Training instances to cluster, or distances between instances if
metric='precomputed'.- yIgnored
Not used, present here for API consistency by convention.
- Returns:
- selfobject
Returns the fitted instance.
- fit_predict(X, y=None)[source]#
Fit and return the result of each sample’s clustering assignment.
In addition to fitting, this method also return the result of the clustering assignment for each sample in the training set.
- Parameters:
- Xarray-like of shape (n_samples, n_features) or (n_samples, n_samples)
Training instances to cluster, or distances between instances if
affinity='precomputed'.- yIgnored
Not used, present here for API consistency by convention.
- Returns:
- labelsndarray of shape (n_samples,)
Cluster labels.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
Gallery examples#
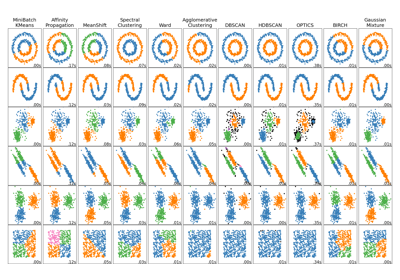
Comparing different clustering algorithms on toy datasets

A demo of structured Ward hierarchical clustering on an image of coins
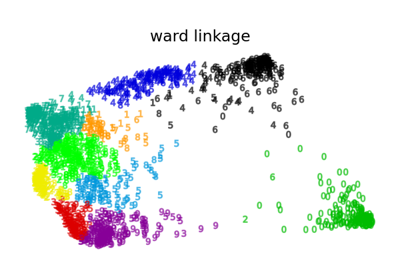
Various Agglomerative Clustering on a 2D embedding of digits
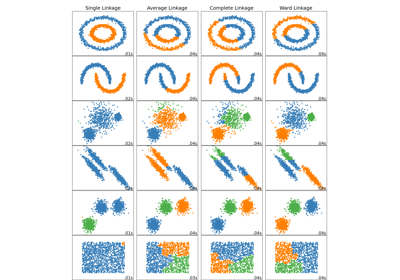
Comparing different hierarchical linkage methods on toy datasets
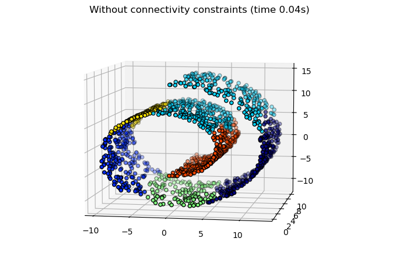
Hierarchical clustering with and without structure