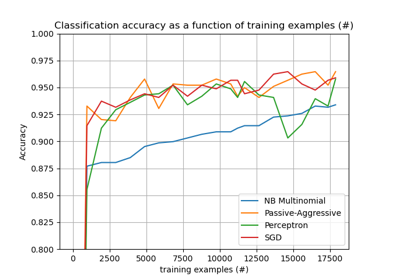
Perceptron#
- class sklearn.linear_model.Perceptron(*, penalty=None, alpha=0.0001, l1_ratio=0.15, fit_intercept=True, max_iter=1000, tol=0.001, shuffle=True, verbose=0, eta0=1.0, n_jobs=None, random_state=0, early_stopping=False, validation_fraction=0.1, n_iter_no_change=5, class_weight=None, warm_start=False)[source]#
Linear perceptron classifier.
The implementation is a wrapper around
SGDClassifierby fixing thelossandlearning_rateparameters as:SGDClassifier(loss="perceptron", learning_rate="constant")
Other available parameters are described below and are forwarded to
SGDClassifier.Read more in the User Guide.
- Parameters:
- penalty{‘l2’,’l1’,’elasticnet’}, default=None
The penalty (aka regularization term) to be used.
- alphafloat, default=0.0001
Constant that multiplies the regularization term if regularization is used.
- l1_ratiofloat, default=0.15
The Elastic Net mixing parameter, with
0 <= l1_ratio <= 1.l1_ratio=0corresponds to L2 penalty,l1_ratio=1to L1. Only used ifpenalty='elasticnet'.Added in version 0.24.
- fit_interceptbool, default=True
Whether the intercept should be estimated or not. If False, the data is assumed to be already centered.
- max_iterint, default=1000
The maximum number of passes over the training data (aka epochs). It only impacts the behavior in the
fitmethod, and not thepartial_fitmethod.Added in version 0.19.
- tolfloat or None, default=1e-3
The stopping criterion. If it is not None, the iterations will stop when (loss > previous_loss - tol).
Added in version 0.19.
- shufflebool, default=True
Whether or not the training data should be shuffled after each epoch.
- verboseint, default=0
The verbosity level.
- eta0float, default=1
Constant by which the updates are multiplied.
- n_jobsint, default=None
The number of CPUs to use to do the OVA (One Versus All, for multi-class problems) computation.
Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.- random_stateint, RandomState instance or None, default=0
Used to shuffle the training data, when
shuffleis set toTrue. Pass an int for reproducible output across multiple function calls. See Glossary.- early_stoppingbool, default=False
Whether to use early stopping to terminate training when validation score is not improving. If set to True, it will automatically set aside a stratified fraction of training data as validation and terminate training when validation score is not improving by at least
tolforn_iter_no_changeconsecutive epochs.Added in version 0.20.
- validation_fractionfloat, default=0.1
The proportion of training data to set aside as validation set for early stopping. Must be between 0 and 1. Only used if early_stopping is True.
Added in version 0.20.
- n_iter_no_changeint, default=5
Number of iterations with no improvement to wait before early stopping.
Added in version 0.20.
- class_weightdict, {class_label: weight} or “balanced”, default=None
Preset for the class_weight fit parameter.
Weights associated with classes. If not given, all classes are supposed to have weight one.
The “balanced” mode uses the values of y to automatically adjust weights inversely proportional to class frequencies in the input data as
n_samples / (n_classes * np.bincount(y)).- warm_startbool, default=False
When set to True, reuse the solution of the previous call to fit as initialization, otherwise, just erase the previous solution. See the Glossary.
- Attributes:
- classes_ndarray of shape (n_classes,)
The unique classes labels.
- coef_ndarray of shape (1, n_features) if n_classes == 2 else (n_classes, n_features)
Weights assigned to the features.
- intercept_ndarray of shape (1,) if n_classes == 2 else (n_classes,)
Constants in decision function.
- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
- n_iter_int
The actual number of iterations to reach the stopping criterion. For multiclass fits, it is the maximum over every binary fit.
- t_int
Number of weight updates performed during training. Same as
(n_iter_ * n_samples + 1).
See also
sklearn.linear_model.SGDClassifierLinear classifiers (SVM, logistic regression, etc.) with SGD training.
Notes
Perceptronis a classification algorithm which shares the same underlying implementation withSGDClassifier. In fact,Perceptron()is equivalent toSGDClassifier(loss="perceptron", eta0=1, learning_rate="constant", penalty=None).References
https://en.wikipedia.org/wiki/Perceptron and references therein.
Examples
>>> from sklearn.datasets import load_digits >>> from sklearn.linear_model import Perceptron >>> X, y = load_digits(return_X_y=True) >>> clf = Perceptron(tol=1e-3, random_state=0) >>> clf.fit(X, y) Perceptron() >>> clf.score(X, y) 0.939...
- decision_function(X)[source]#
Predict confidence scores for samples.
The confidence score for a sample is proportional to the signed distance of that sample to the hyperplane.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The data matrix for which we want to get the confidence scores.
- Returns:
- scoresndarray of shape (n_samples,) or (n_samples, n_classes)
Confidence scores per
(n_samples, n_classes)combination. In the binary case, confidence score forself.classes_[1]where >0 means this class would be predicted.
- densify()[source]#
Convert coefficient matrix to dense array format.
Converts the
coef_member (back) to a numpy.ndarray. This is the default format ofcoef_and is required for fitting, so calling this method is only required on models that have previously been sparsified; otherwise, it is a no-op.- Returns:
- self
Fitted estimator.
- fit(X, y, coef_init=None, intercept_init=None, sample_weight=None)[source]#
Fit linear model with Stochastic Gradient Descent.
- Parameters:
- X{array-like, sparse matrix}, shape (n_samples, n_features)
Training data.
- yndarray of shape (n_samples,)
Target values.
- coef_initndarray of shape (n_classes, n_features), default=None
The initial coefficients to warm-start the optimization.
- intercept_initndarray of shape (n_classes,), default=None
The initial intercept to warm-start the optimization.
- sample_weightarray-like, shape (n_samples,), default=None
Weights applied to individual samples. If not provided, uniform weights are assumed. These weights will be multiplied with class_weight (passed through the constructor) if class_weight is specified.
- Returns:
- selfobject
Returns an instance of self.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- partial_fit(X, y, classes=None, sample_weight=None)[source]#
Perform one epoch of stochastic gradient descent on given samples.
Internally, this method uses
max_iter = 1. Therefore, it is not guaranteed that a minimum of the cost function is reached after calling it once. Matters such as objective convergence, early stopping, and learning rate adjustments should be handled by the user.- Parameters:
- X{array-like, sparse matrix}, shape (n_samples, n_features)
Subset of the training data.
- yndarray of shape (n_samples,)
Subset of the target values.
- classesndarray of shape (n_classes,), default=None
Classes across all calls to partial_fit. Can be obtained by via
np.unique(y_all), where y_all is the target vector of the entire dataset. This argument is required for the first call to partial_fit and can be omitted in the subsequent calls. Note that y doesn’t need to contain all labels inclasses.- sample_weightarray-like, shape (n_samples,), default=None
Weights applied to individual samples. If not provided, uniform weights are assumed.
- Returns:
- selfobject
Returns an instance of self.
- predict(X)[source]#
Predict class labels for samples in X.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The data matrix for which we want to get the predictions.
- Returns:
- y_predndarray of shape (n_samples,)
Vector containing the class labels for each sample.
- score(X, y, sample_weight=None)[source]#
Return accuracy on provided data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True labels for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
Mean accuracy of
self.predict(X)w.r.t.y.
- set_fit_request(*, coef_init: bool | None | str = '$UNCHANGED$', intercept_init: bool | None | str = '$UNCHANGED$', sample_weight: bool | None | str = '$UNCHANGED$') Perceptron[source]#
Configure whether metadata should be requested to be passed to the
fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- coef_initstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
coef_initparameter infit.- intercept_initstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
intercept_initparameter infit.- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter infit.
- Returns:
- selfobject
The updated object.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_partial_fit_request(*, classes: bool | None | str = '$UNCHANGED$', sample_weight: bool | None | str = '$UNCHANGED$') Perceptron[source]#
Configure whether metadata should be requested to be passed to the
partial_fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed topartial_fitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it topartial_fit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- classesstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
classesparameter inpartial_fit.- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inpartial_fit.
- Returns:
- selfobject
The updated object.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') Perceptron[source]#
Configure whether metadata should be requested to be passed to the
scoremethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.
- sparsify()[source]#
Convert coefficient matrix to sparse format.
Converts the
coef_member to a scipy.sparse matrix, which for L1-regularized models can be much more memory- and storage-efficient than the usual numpy.ndarray representation.The
intercept_member is not converted.Warning
This method is not supported for estimators fitted with array API inputs (i.e. when
sklearn.config_contextis used witharray_api_dispatch=True). The call may succeed but subsequent calls topredictand other methods involving passing arrays may raise or return unexpected results.- Returns:
- self
Fitted estimator.
Notes
For non-sparse models, i.e. when there are not many zeros in
coef_, this may actually increase memory usage, so use this method with care. A rule of thumb is that the number of zero elements, which can be computed with(coef_ == 0).sum(), must be more than 50% for this to provide significant benefits.After calling this method, further fitting with the partial_fit method (if any) will not work until you call densify.