GroupKFold#
- class sklearn.model_selection.GroupKFold(n_splits=5, *, shuffle=False, random_state=None)[source]#
K-fold iterator variant with non-overlapping groups.
Each group will appear exactly once in the test set across all folds (the number of distinct groups has to be at least equal to the number of folds).
The folds are approximately balanced in the sense that the number of samples is approximately the same in each test fold when
shuffleis True.Read more in the User Guide.
For visualisation of cross-validation behaviour and comparison between common scikit-learn split methods refer to Visualizing cross-validation behavior in scikit-learn
- Parameters:
- n_splitsint, default=5
Number of folds. Must be at least 2.
Changed in version 0.22:
n_splitsdefault value changed from 3 to 5.- shufflebool, default=False
Whether to shuffle the groups before splitting into batches. Note that the samples within each split will not be shuffled.
Added in version 1.6.
- random_stateint, RandomState instance or None, default=None
When
shuffleis True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. Pass an int for reproducible output across multiple function calls. See Glossary.Added in version 1.6.
See also
LeaveOneGroupOutFor splitting the data according to explicit, domain-specific grouping of the dataset.
StratifiedKFoldTakes class information into account to avoid building folds with imbalanced class proportions (for binary or multiclass classification tasks).
Notes
Groups appear in an arbitrary order throughout the folds.
Examples
>>> import numpy as np >>> from sklearn.model_selection import GroupKFold >>> X = np.array([[1, 2], [3, 4], [5, 6], [7, 8], [9, 10], [11, 12]]) >>> y = np.array([1, 2, 3, 4, 5, 6]) >>> groups = np.array([0, 0, 2, 2, 3, 3]) >>> group_kfold = GroupKFold(n_splits=2) >>> group_kfold.get_n_splits() 2 >>> print(group_kfold) GroupKFold(n_splits=2, random_state=None, shuffle=False) >>> for i, (train_index, test_index) in enumerate(group_kfold.split(X, y, groups)): ... print(f"Fold {i}:") ... print(f" Train: index={train_index}, group={groups[train_index]}") ... print(f" Test: index={test_index}, group={groups[test_index]}") Fold 0: Train: index=[2 3], group=[2 2] Test: index=[0 1 4 5], group=[0 0 3 3] Fold 1: Train: index=[0 1 4 5], group=[0 0 3 3] Test: index=[2 3], group=[2 2]
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_n_splits(X=None, y=None, groups=None)[source]#
Returns the number of splitting iterations as set with the
n_splitsparam when instantiating the cross-validator.- Parameters:
- Xarray-like of shape (n_samples, n_features), default=None
Always ignored, exists for API compatibility.
- yarray-like of shape (n_samples,), default=None
Always ignored, exists for API compatibility.
- groupsarray-like of shape (n_samples,), default=None
Always ignored, exists for API compatibility.
- Returns:
- n_splitsint
Returns the number of splitting iterations in the cross-validator.
- set_split_request(*, groups: bool | None | str = '$UNCHANGED$') GroupKFold[source]#
Configure whether metadata should be requested to be passed to the
splitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tosplitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tosplit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- groupsstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
groupsparameter insplit.
- Returns:
- selfobject
The updated object.
- split(X, y=None, groups=None)[source]#
Generate indices to split data into training and test set.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Training data, where
n_samplesis the number of samples andn_featuresis the number of features.- yarray-like of shape (n_samples,), default=None
The target variable for supervised learning problems.
- groupsarray-like of shape (n_samples,)
Group labels for the samples used while splitting the dataset into train/test set.
- Yields:
- trainndarray
The training set indices for that split.
- testndarray
The testing set indices for that split.
Gallery examples#
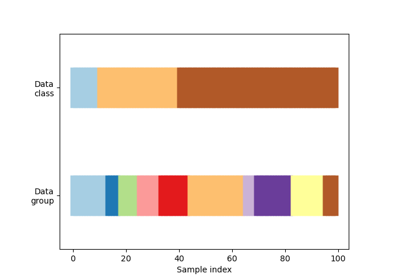
Visualizing cross-validation behavior in scikit-learn