ARDRegression#
- class sklearn.linear_model.ARDRegression(*, max_iter=300, tol=0.001, alpha_1=1e-06, alpha_2=1e-06, lambda_1=1e-06, lambda_2=1e-06, compute_score=False, threshold_lambda=10000.0, fit_intercept=True, copy_X=True, verbose=False)[source]#
Bayesian ARD regression.
Fit the weights of a regression model, using an ARD prior. The weights of the regression model are assumed to be in Gaussian distributions. Also estimate the parameters lambda (precisions of the distributions of the weights) and alpha (precision of the distribution of the noise). The estimation is done by an iterative procedures (Evidence Maximization)
Read more in the User Guide.
- Parameters:
- max_iterint, default=300
Maximum number of iterations.
Changed in version 1.3.
- tolfloat, default=1e-3
Stop the algorithm if w has converged.
- alpha_1float, default=1e-6
Hyper-parameter : shape parameter for the Gamma distribution prior over the alpha parameter.
- alpha_2float, default=1e-6
Hyper-parameter : inverse scale parameter (rate parameter) for the Gamma distribution prior over the alpha parameter.
- lambda_1float, default=1e-6
Hyper-parameter : shape parameter for the Gamma distribution prior over the lambda parameter.
- lambda_2float, default=1e-6
Hyper-parameter : inverse scale parameter (rate parameter) for the Gamma distribution prior over the lambda parameter.
- compute_scorebool, default=False
If True, compute the objective function at each step of the model.
- threshold_lambdafloat, default=10 000
Threshold for removing (pruning) weights with high precision from the computation.
- fit_interceptbool, default=True
Whether to calculate the intercept for this model. If set to false, no intercept will be used in calculations (i.e. data is expected to be centered).
- copy_Xbool, default=True
If True, X will be copied; else, it may be overwritten.
- verbosebool, default=False
Verbose mode when fitting the model.
- Attributes:
- coef_array-like of shape (n_features,)
Coefficients of the regression model (mean of distribution)
- alpha_float
estimated precision of the noise.
- lambda_array-like of shape (n_features,)
estimated precisions of the weights.
- sigma_array-like of shape (n_features, n_features)
estimated variance-covariance matrix of the weights
- scores_float
if computed, value of the objective function (to be maximized)
- n_iter_int
The actual number of iterations to reach the stopping criterion.
Added in version 1.3.
- intercept_float
Independent term in decision function. Set to 0.0 if
fit_intercept = False.- X_offset_float
If
fit_intercept=True, offset subtracted for centering data to a zero mean. Set to np.zeros(n_features) otherwise.- X_scale_float
Set to np.ones(n_features).
- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
See also
BayesianRidgeBayesian ridge regression.
References
D. J. C. MacKay, Bayesian nonlinear modeling for the prediction competition, ASHRAE Transactions, 1994.
R. Salakhutdinov, Lecture notes on Statistical Machine Learning, http://www.utstat.toronto.edu/~rsalakhu/sta4273/notes/Lecture2.pdf#page=15 Their beta is our
self.alpha_Their alpha is ourself.lambda_ARD is a little different than the slide: only dimensions/features for whichself.lambda_ < self.threshold_lambdaare kept and the rest are discarded.Examples
>>> from sklearn import linear_model >>> clf = linear_model.ARDRegression() >>> clf.fit([[0,0], [1, 1], [2, 2]], [0, 1, 2]) ARDRegression() >>> clf.predict([[1, 1]]) array([1.])
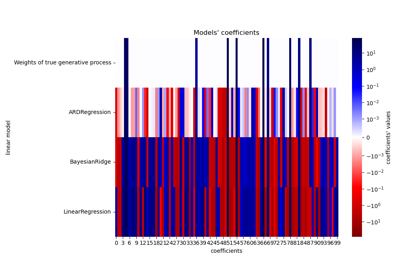
Comparing Linear Bayesian Regressors demonstrates ARD Regression.
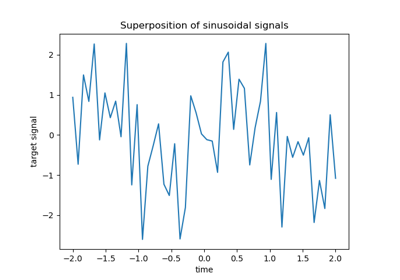
L1-based models for Sparse Signals showcases ARD Regression alongside Lasso and Elastic-Net for sparse, correlated signals, in the presence of noise.
- fit(X, y)[source]#
Fit the model according to the given training data and parameters.
Iterative procedure to maximize the evidence
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Training vector, where
n_samplesis the number of samples andn_featuresis the number of features.- yarray-like of shape (n_samples,)
Target values (integers). Will be cast to X’s dtype if necessary.
- Returns:
- selfobject
Fitted estimator.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- predict(X, return_std=False)[source]#
Predict using the linear model.
In addition to the mean of the predictive distribution, also its standard deviation can be returned.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Samples.
- return_stdbool, default=False
Whether to return the standard deviation of posterior prediction.
- Returns:
- y_meanarray-like of shape (n_samples,)
Mean of predictive distribution of query points.
- y_stdarray-like of shape (n_samples,)
Standard deviation of predictive distribution of query points.
- score(X, y, sample_weight=None)[source]#
Return coefficient of determination on test data.
The coefficient of determination, \(R^2\), is defined as \((1 - \frac{u}{v})\), where \(u\) is the residual sum of squares
((y_true - y_pred)** 2).sum()and \(v\) is the total sum of squares((y_true - y_true.mean()) ** 2).sum(). The best possible score is 1.0 and it can be negative (because the model can be arbitrarily worse). A constant model that always predicts the expected value ofy, disregarding the input features, would get a \(R^2\) score of 0.0.- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples. For some estimators this may be a precomputed kernel matrix or a list of generic objects instead with shape
(n_samples, n_samples_fitted), wheren_samples_fittedis the number of samples used in the fitting for the estimator.- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True values for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
\(R^2\) of
self.predict(X)w.r.t.y.
Notes
The \(R^2\) score used when calling
scoreon a regressor usesmultioutput='uniform_average'from version 0.23 to keep consistent with default value ofr2_score. This influences thescoremethod of all the multioutput regressors (except forMultiOutputRegressor).
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_predict_request(*, return_std: bool | None | str = '$UNCHANGED$') ARDRegression[source]#
Configure whether metadata should be requested to be passed to the
predictmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed topredictif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it topredict.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- return_stdstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
return_stdparameter inpredict.
- Returns:
- selfobject
The updated object.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') ARDRegression[source]#
Configure whether metadata should be requested to be passed to the
scoremethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.