GridSearchCV#
- class sklearn.model_selection.GridSearchCV(estimator, param_grid, *, scoring=None, n_jobs=None, refit=True, cv=None, verbose=0, pre_dispatch='2*n_jobs', error_score=nan, return_train_score=False)[source]#
Exhaustive search over specified parameter values for an estimator.
Important members are fit, predict.
GridSearchCV implements a “fit” and a “score” method. It also implements “score_samples”, “predict”, “predict_proba”, “decision_function”, “transform” and “inverse_transform” if they are implemented in the estimator used.
The parameters of the estimator used to apply these methods are optimized by cross-validated grid-search over a parameter grid.
Read more in the User Guide.
- Parameters:
- estimatorestimator object
This is assumed to implement the scikit-learn estimator interface. Either estimator needs to provide a
scorefunction, orscoringmust be passed.- param_griddict or list of dictionaries
Dictionary with parameters names (
str) as keys and lists of parameter settings to try as values, or a list of such dictionaries, in which case the grids spanned by each dictionary in the list are explored. This enables searching over any sequence of parameter settings.- scoringstr, callable, list, tuple or dict, default=None
Strategy to evaluate the performance of the cross-validated model on the test set.
If
scoringrepresents a single score, one can use:a single string (see String name scorers);
a callable (see Callable scorers) that returns a single value;
None, theestimator’s default evaluation criterion is used.
If
scoringrepresents multiple scores, one can use:a list or tuple of unique strings;
a callable returning a dictionary where the keys are the metric names and the values are the metric scores;
a dictionary with metric names as keys and callables as values.
See Specifying multiple metrics for evaluation for an example.
- n_jobsint, default=None
Number of jobs to run in parallel.
Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.Changed in version v0.20:
n_jobsdefault changed from 1 to None- refitbool, str, or callable, default=True
Refit an estimator using the best found parameters on the whole dataset.
For multiple metric evaluation, this needs to be a
strdenoting the scorer that would be used to find the best parameters for refitting the estimator at the end.Where there are considerations other than maximum score in choosing a best estimator,
refitcan be set to a function which returns the selectedbest_index_givencv_results_. In that case, thebest_estimator_andbest_params_will be set according to the returnedbest_index_while thebest_score_attribute will not be available.The refitted estimator is made available at the
best_estimator_attribute and permits usingpredictdirectly on thisGridSearchCVinstance.Also for multiple metric evaluation, the attributes
best_index_,best_score_andbest_params_will only be available ifrefitis set and all of them will be determined w.r.t this specific scorer.See
scoringparameter to know more about multiple metric evaluation.See Custom refit strategy of a grid search with cross-validation to see how to design a custom selection strategy using a callable via
refit.See this example for an example of how to use
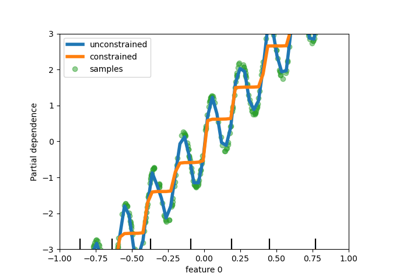
refit=callableto balance model complexity and cross-validated score.Changed in version 0.20: Support for callable added.
- cvint, cross-validation generator or an iterable, default=None
Determines the cross-validation splitting strategy. Possible inputs for cv are:
None, to use the default 5-fold cross validation,
integer, to specify the number of folds in a
(Stratified)KFold,an iterable yielding (train, test) splits as arrays of indices.
For integer/None inputs, if the estimator is a classifier and
yis either binary or multiclass,StratifiedKFoldis used. In all other cases,KFoldis used. These splitters are instantiated withshuffle=Falseso the splits will be the same across calls.Refer User Guide for the various cross-validation strategies that can be used here.
Changed in version 0.22:
cvdefault value if None changed from 3-fold to 5-fold.- verboseint, default=0
Controls the verbosity of information printed during fitting, with higher values yielding more detailed logging.
0 : no messages are printed;
>=1 : summary of the total number of fits;
>=2 : computation time for each fold and parameter candidate;
>=3 : fold indices and scores;
>=10 : parameter candidate indices and START messages before each fit.
- pre_dispatchint, or str, default=’2*n_jobs’
Controls the number of jobs that get dispatched during parallel execution. Reducing this number can be useful to avoid an explosion of memory consumption when more jobs get dispatched than CPUs can process. This parameter can be:
None, in which case all the jobs are immediately created and spawned. Use this for lightweight and fast-running jobs, to avoid delays due to on-demand spawning of the jobs
An int, giving the exact number of total jobs that are spawned
A str, giving an expression as a function of n_jobs, as in ‘2*n_jobs’
- error_score‘raise’ or numeric, default=np.nan
Value to assign to the score if an error occurs in estimator fitting. If set to ‘raise’, the error is raised. If a numeric value is given, FitFailedWarning is raised. This parameter does not affect the refit step, which will always raise the error.
- return_train_scorebool, default=False
If
False, thecv_results_attribute will not include training scores. Computing training scores is used to get insights on how different parameter settings impact the overfitting/underfitting trade-off. However computing the scores on the training set can be computationally expensive and is not strictly required to select the parameters that yield the best generalization performance.Added in version 0.19.
Changed in version 0.21: Default value was changed from
TruetoFalse
- Attributes:
- cv_results_dict of numpy (masked) ndarrays
A dict with keys as column headers and values as columns, that can be imported into a pandas
DataFrame.For instance the below given table
param_kernel
param_gamma
param_degree
split0_test_score
…
rank_t…
‘poly’
–
2
0.80
…
2
‘poly’
–
3
0.70
…
4
‘rbf’
0.1
–
0.80
…
3
‘rbf’
0.2
–
0.93
…
1
will be represented by a
cv_results_dict of:{ 'param_kernel': masked_array(data = ['poly', 'poly', 'rbf', 'rbf'], mask = [False False False False]...) 'param_gamma': masked_array(data = [-- -- 0.1 0.2], mask = [ True True False False]...), 'param_degree': masked_array(data = [2.0 3.0 -- --], mask = [False False True True]...), 'split0_test_score' : [0.80, 0.70, 0.80, 0.93], 'split1_test_score' : [0.82, 0.50, 0.70, 0.78], 'mean_test_score' : [0.81, 0.60, 0.75, 0.85], 'std_test_score' : [0.01, 0.10, 0.05, 0.08], 'rank_test_score' : [2, 4, 3, 1], 'split0_train_score' : [0.80, 0.92, 0.70, 0.93], 'split1_train_score' : [0.82, 0.55, 0.70, 0.87], 'mean_train_score' : [0.81, 0.74, 0.70, 0.90], 'std_train_score' : [0.01, 0.19, 0.00, 0.03], 'mean_fit_time' : [0.73, 0.63, 0.43, 0.49], 'std_fit_time' : [0.01, 0.02, 0.01, 0.01], 'mean_score_time' : [0.01, 0.06, 0.04, 0.04], 'std_score_time' : [0.00, 0.00, 0.00, 0.01], 'params' : [{'kernel': 'poly', 'degree': 2}, ...], }
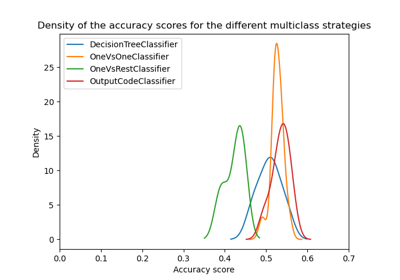
For an example of visualization and interpretation of GridSearch results, see Statistical comparison of models using grid search.
NOTE
The key
'params'is used to store a list of parameter settings dicts for all the parameter candidates.The
mean_fit_time,std_fit_time,mean_score_timeandstd_score_timeare all in seconds.For multi-metric evaluation, the scores for all the scorers are available in the
cv_results_dict at the keys ending with that scorer’s name ('_<scorer_name>') instead of'_score'shown above. (‘split0_test_precision’, ‘mean_train_precision’ etc.)- best_estimator_estimator
Estimator that was chosen by the search, i.e. estimator which gave highest score (or smallest loss if specified) on the left out data. Not available if
refit=False.See
refitparameter for more information on allowed values.- best_score_float
Mean cross-validated score of the best_estimator
For multi-metric evaluation, this is present only if
refitis specified.This attribute is not available if
refitis a function.- best_params_dict
Parameter setting that gave the best results on the hold out data.
For multi-metric evaluation, this is present only if
refitis specified.- best_index_int
The index (of the
cv_results_arrays) which corresponds to the best candidate parameter setting.The dict at
search.cv_results_['params'][search.best_index_]gives the parameter setting for the best model, that gives the highest mean score (search.best_score_).For multi-metric evaluation, this is present only if
refitis specified.- scorer_function or a dict
Scorer function used on the held out data to choose the best parameters for the model.
For multi-metric evaluation, this attribute holds the validated
scoringdict which maps the scorer key to the scorer callable.- n_splits_int
The number of cross-validation splits (folds/iterations).
- refit_time_float
Seconds used for refitting the best model on the whole dataset.
This is present only if
refitis not False.Added in version 0.20.
- multimetric_bool
Whether or not the scorers compute several metrics.
classes_ndarray of shape (n_classes,)Class labels.
n_features_in_intNumber of features seen during fit.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Only defined if
best_estimator_is defined (see the documentation for therefitparameter for more details) and thatbest_estimator_exposesfeature_names_in_when fit.Added in version 1.0.
See also
ParameterGridGenerates all the combinations of a hyperparameter grid.
train_test_splitUtility function to split the data into a development set usable for fitting a GridSearchCV instance and an evaluation set for its final evaluation.
sklearn.metrics.make_scorerMake a scorer from a performance metric or loss function.
Notes
The parameters selected are those that maximize the score of the left out data, unless an explicit score is passed in which case it is used instead.
If
n_jobswas set to a value higher than one, the data is copied for each point in the grid (and notn_jobstimes). This is done for efficiency reasons if individual jobs take very little time, but may raise errors if the dataset is large and not enough memory is available. A workaround in this case is to setpre_dispatch. Then, the memory is copied onlypre_dispatchmany times. A reasonable value forpre_dispatchis2 * n_jobs.Examples
>>> from sklearn import svm, datasets >>> from sklearn.model_selection import GridSearchCV >>> iris = datasets.load_iris() >>> parameters = {'kernel':('linear', 'rbf'), 'C':[1, 10]} >>> svc = svm.SVC() >>> clf = GridSearchCV(svc, parameters) >>> clf.fit(iris.data, iris.target) GridSearchCV(estimator=SVC(), param_grid={'C': [1, 10], 'kernel': ('linear', 'rbf')}) >>> sorted(clf.cv_results_.keys()) ['mean_fit_time', 'mean_score_time', 'mean_test_score',... 'param_C', 'param_kernel', 'params',... 'rank_test_score', 'split0_test_score',... 'split2_test_score', ... 'std_fit_time', 'std_score_time', 'std_test_score']
- decision_function(X)[source]#
Call decision_function on the estimator with the best found parameters.
Only available if
refit=Trueand the underlying estimator supportsdecision_function.- Parameters:
- Xindexable, length n_samples
Must fulfill the input assumptions of the underlying estimator.
- Returns:
- y_scorendarray of shape (n_samples,) or (n_samples, n_classes) or (n_samples, n_classes * (n_classes-1) / 2)
Result of the decision function for
Xbased on the estimator with the best found parameters.
- fit(X, y=None, **params)[source]#
Run fit with all sets of parameters.
- Parameters:
- Xarray-like of shape (n_samples, n_features) or (n_samples, n_samples)
Training vectors, where
n_samplesis the number of samples andn_featuresis the number of features. For precomputed kernel or distance matrix, the expected shape of X is (n_samples, n_samples).- yarray-like of shape (n_samples, n_output) or (n_samples,), default=None
Target relative to X for classification or regression; None for unsupervised learning.
- **paramsdict of str -> object
Parameters passed to the
fitmethod of the estimator, the scorer, and the CV splitter.If a fit parameter is an array-like whose length is equal to
num_samplesthen it will be split by cross-validation along withXandy. For example, the sample_weight parameter is split becauselen(sample_weights) = len(X). However, this behavior does not apply togroupswhich is passed to the splitter configured via thecvparameter of the constructor. Thus,groupsis used to perform the split and determines which samples are assigned to the each side of the a split.
- Returns:
- selfobject
Instance of fitted estimator.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
Added in version 1.4.
- Returns:
- routingMetadataRouter
A
MetadataRouterencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- inverse_transform(X)[source]#
Call inverse_transform on the estimator with the best found params.
Only available if the underlying estimator implements
inverse_transformandrefit=True.- Parameters:
- Xindexable, length n_samples
Must fulfill the input assumptions of the underlying estimator.
- Returns:
- X_original{ndarray, sparse matrix} of shape (n_samples, n_features)
Result of the
inverse_transformfunction forXbased on the estimator with the best found parameters.
- predict(X)[source]#
Call predict on the estimator with the best found parameters.
Only available if
refit=Trueand the underlying estimator supportspredict.- Parameters:
- Xindexable, length n_samples
Must fulfill the input assumptions of the underlying estimator.
- Returns:
- y_predndarray of shape (n_samples,)
The predicted labels or values for
Xbased on the estimator with the best found parameters.
- predict_log_proba(X)[source]#
Call predict_log_proba on the estimator with the best found parameters.
Only available if
refit=Trueand the underlying estimator supportspredict_log_proba.- Parameters:
- Xindexable, length n_samples
Must fulfill the input assumptions of the underlying estimator.
- Returns:
- y_predndarray of shape (n_samples,) or (n_samples, n_classes)
Predicted class log-probabilities for
Xbased on the estimator with the best found parameters. The order of the classes corresponds to that in the fitted attribute classes_.
- predict_proba(X)[source]#
Call predict_proba on the estimator with the best found parameters.
Only available if
refit=Trueand the underlying estimator supportspredict_proba.- Parameters:
- Xindexable, length n_samples
Must fulfill the input assumptions of the underlying estimator.
- Returns:
- y_predndarray of shape (n_samples,) or (n_samples, n_classes)
Predicted class probabilities for
Xbased on the estimator with the best found parameters. The order of the classes corresponds to that in the fitted attribute classes_.
- score(X, y=None, **params)[source]#
Return the score on the given data, if the estimator has been refit.
This uses the score defined by
scoringwhere provided, and thebest_estimator_.scoremethod otherwise.- Parameters:
- Xarray-like of shape (n_samples, n_features)
Input data, where
n_samplesis the number of samples andn_featuresis the number of features.- yarray-like of shape (n_samples, n_output) or (n_samples,), default=None
Target relative to X for classification or regression; None for unsupervised learning.
- **paramsdict
Parameters to be passed to the underlying scorer(s).
Added in version 1.4: Only available if
enable_metadata_routing=True. See Metadata Routing User Guide for more details.
- Returns:
- scorefloat
The score defined by
scoringif provided, and thebest_estimator_.scoremethod otherwise.
- score_samples(X)[source]#
Call score_samples on the estimator with the best found parameters.
Only available if
refit=Trueand the underlying estimator supportsscore_samples.Added in version 0.24.
- Parameters:
- Xiterable
Data to predict on. Must fulfill input requirements of the underlying estimator.
- Returns:
- y_scorendarray of shape (n_samples,)
The
best_estimator_.score_samplesmethod.
- set_callbacks(*callbacks)[source]#
Set callbacks for the estimator.
- Parameters:
- *callbackscallback instances
The callbacks to set.
- Returns:
- selfestimator instance
The estimator instance itself.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- transform(X)[source]#
Call transform on the estimator with the best found parameters.
Only available if the underlying estimator supports
transformandrefit=True.- Parameters:
- Xindexable, length n_samples
Must fulfill the input assumptions of the underlying estimator.
- Returns:
- Xt{ndarray, sparse matrix} of shape (n_samples, n_features)
Xtransformed in the new space based on the estimator with the best found parameters.
Gallery examples#

Analysis of the convergence of penalized logistic regression models
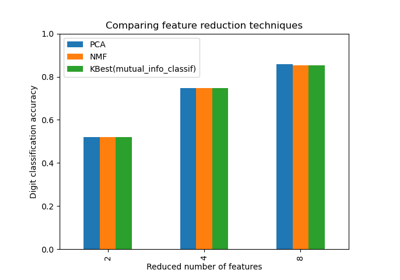
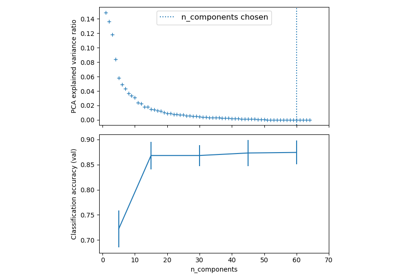
Selecting dimensionality reduction with Pipeline and GridSearchCV
Pipelining: chaining a PCA and a logistic regression
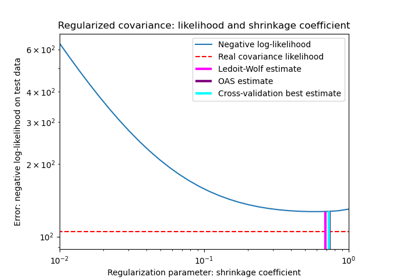
Shrinkage covariance estimation: LedoitWolf vs OAS and max-likelihood
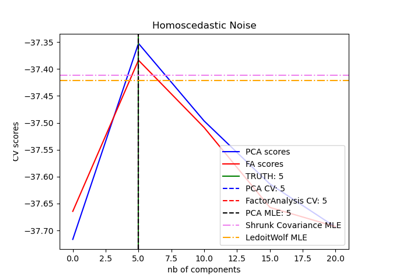
Model selection with Probabilistic PCA and Factor Analysis (FA)

Comparing Random Forests and Histogram Gradient Boosting models
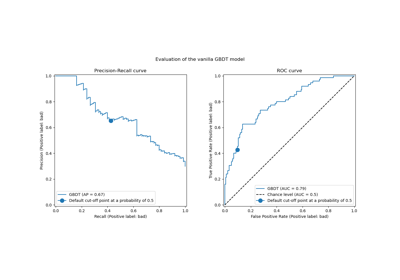
Post-tuning the decision threshold for cost-sensitive learning

Custom refit strategy of a grid search with cross-validation
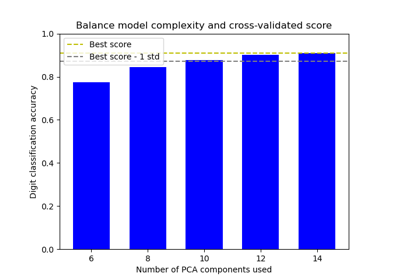
Balance model complexity and cross-validated score
Statistical comparison of models using grid search

Sample pipeline for text feature extraction and evaluation
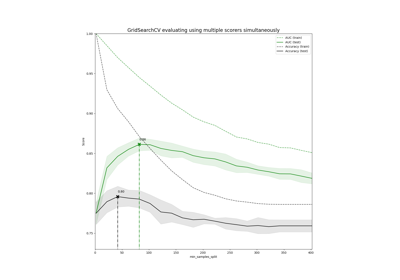
Demonstration of multi-metric evaluation on cross_val_score and GridSearchCV

Comparing randomized search and grid search for hyperparameter estimation
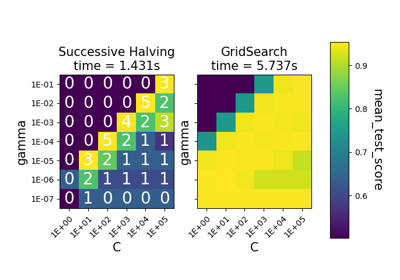
Comparison between grid search and successive halving
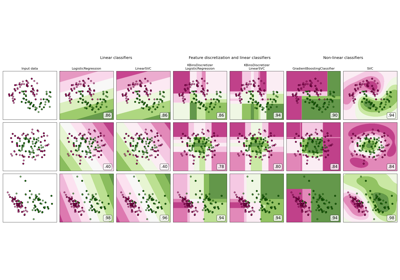
Plot classification boundaries with different SVM Kernels