BernoulliNB#
- class sklearn.naive_bayes.BernoulliNB(*, alpha=1.0, force_alpha=True, binarize=0.0, fit_prior=True, class_prior=None)[source]#
Naive Bayes classifier for multivariate Bernoulli models.
Like MultinomialNB, this classifier is suitable for discrete data. The difference is that while MultinomialNB works with occurrence counts, BernoulliNB is designed for binary/boolean features.
Read more in the User Guide.
- Parameters:
- alphafloat or array-like of shape (n_features,), default=1.0
Additive (Laplace/Lidstone) smoothing parameter (set alpha=0 and force_alpha=True, for no smoothing).
- force_alphabool, default=True
If False and alpha is less than 1e-10, it will set alpha to 1e-10. If True, alpha will remain unchanged. This may cause numerical errors if alpha is too close to 0.
Added in version 1.2.
Changed in version 1.4: The default value of
force_alphachanged toTrue.- binarizefloat or None, default=0.0
Threshold for binarizing (mapping to booleans) of sample features. If None, input is presumed to already consist of binary vectors.
- fit_priorbool, default=True
Whether to learn class prior probabilities or not. If false, a uniform prior will be used.
- class_priorarray-like of shape (n_classes,), default=None
Prior probabilities of the classes. If specified, the priors are not adjusted according to the data.
- Attributes:
- class_count_ndarray of shape (n_classes,)
Number of samples encountered for each class during fitting. This value is weighted by the sample weight when provided.
- class_log_prior_ndarray of shape (n_classes,)
Log probability of each class (smoothed).
- classes_ndarray of shape (n_classes,)
Class labels known to the classifier
- feature_count_ndarray of shape (n_classes, n_features)
Number of samples encountered for each (class, feature) during fitting. This value is weighted by the sample weight when provided.
- feature_log_prob_ndarray of shape (n_classes, n_features)
Empirical log probability of features given a class, P(x_i|y).
- n_features_in_int
Number of features seen during fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
See also
CategoricalNBNaive Bayes classifier for categorical features.
ComplementNBThe Complement Naive Bayes classifier described in Rennie et al. (2003).
GaussianNBGaussian Naive Bayes (GaussianNB).
MultinomialNBNaive Bayes classifier for multinomial models.
References
C.D. Manning, P. Raghavan and H. Schuetze (2008). Introduction to Information Retrieval. Cambridge University Press, pp. 234-265. https://nlp.stanford.edu/IR-book/html/htmledition/the-bernoulli-model-1.html
A. McCallum and K. Nigam (1998). A comparison of event models for naive Bayes text classification. Proc. AAAI/ICML-98 Workshop on Learning for Text Categorization, pp. 41-48.
V. Metsis, I. Androutsopoulos and G. Paliouras (2006). Spam filtering with naive Bayes – Which naive Bayes? 3rd Conf. on Email and Anti-Spam (CEAS).
Examples
>>> import numpy as np >>> rng = np.random.RandomState(1) >>> X = rng.randint(5, size=(6, 100)) >>> Y = np.array([1, 2, 3, 4, 4, 5]) >>> from sklearn.naive_bayes import BernoulliNB >>> clf = BernoulliNB() >>> clf.fit(X, Y) BernoulliNB() >>> print(clf.predict(X[2:3])) [3]
- fit(X, y, sample_weight=None)[source]#
Fit Naive Bayes classifier according to X, y.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Training vectors, where
n_samplesis the number of samples andn_featuresis the number of features.- yarray-like of shape (n_samples,)
Target values.
- sample_weightarray-like of shape (n_samples,), default=None
Weights applied to individual samples (1. for unweighted).
- Returns:
- selfobject
Returns the instance itself.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- partial_fit(X, y, classes=None, sample_weight=None)[source]#
Incremental fit on a batch of samples.
This method is expected to be called several times consecutively on different chunks of a dataset so as to implement out-of-core or online learning.
This is especially useful when the whole dataset is too big to fit in memory at once.
This method has some performance overhead hence it is better to call partial_fit on chunks of data that are as large as possible (as long as fitting in the memory budget) to hide the overhead.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Training vectors, where
n_samplesis the number of samples andn_featuresis the number of features.- yarray-like of shape (n_samples,)
Target values.
- classesarray-like of shape (n_classes,), default=None
List of all the classes that can possibly appear in the y vector.
Must be provided at the first call to partial_fit, can be omitted in subsequent calls.
- sample_weightarray-like of shape (n_samples,), default=None
Weights applied to individual samples (1. for unweighted).
- Returns:
- selfobject
Returns the instance itself.
- predict(X)[source]#
Perform classification on an array of test vectors X.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
The input samples.
- Returns:
- Cndarray of shape (n_samples,)
Predicted target values for X.
- predict_joint_log_proba(X)[source]#
Return joint log probability estimates for the test vector X.
For each row x of X and class y, the joint log probability is given by
log P(x, y) = log P(y) + log P(x|y),wherelog P(y)is the class prior probability andlog P(x|y)is the class-conditional probability.- Parameters:
- Xarray-like of shape (n_samples, n_features)
The input samples.
- Returns:
- Cndarray of shape (n_samples, n_classes)
Returns the joint log-probability of the samples for each class in the model. The columns correspond to the classes in sorted order, as they appear in the attribute classes_.
- predict_log_proba(X)[source]#
Return log-probability estimates for the test vector X.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
The input samples.
- Returns:
- Carray-like of shape (n_samples, n_classes)
Returns the log-probability of the samples for each class in the model. The columns correspond to the classes in sorted order, as they appear in the attribute classes_.
- predict_proba(X)[source]#
Return probability estimates for the test vector X.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
The input samples.
- Returns:
- Carray-like of shape (n_samples, n_classes)
Returns the probability of the samples for each class in the model. The columns correspond to the classes in sorted order, as they appear in the attribute classes_.
- score(X, y, sample_weight=None)[source]#
Return accuracy on provided data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True labels for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
Mean accuracy of
self.predict(X)w.r.t.y.
- set_fit_request(*, sample_weight: bool | None | str = '$UNCHANGED$') BernoulliNB[source]#
Configure whether metadata should be requested to be passed to the
fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter infit.
- Returns:
- selfobject
The updated object.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_partial_fit_request(*, classes: bool | None | str = '$UNCHANGED$', sample_weight: bool | None | str = '$UNCHANGED$') BernoulliNB[source]#
Configure whether metadata should be requested to be passed to the
partial_fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed topartial_fitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it topartial_fit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- classesstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
classesparameter inpartial_fit.- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inpartial_fit.
- Returns:
- selfobject
The updated object.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') BernoulliNB[source]#
Configure whether metadata should be requested to be passed to the
scoremethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.
Gallery examples#
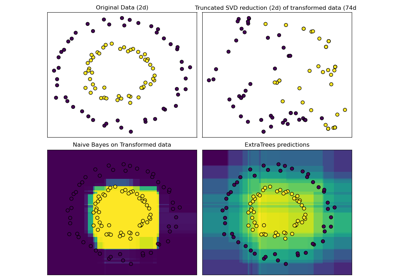
Hashing feature transformation using Totally Random Trees
