v_measure_score#
- sklearn.metrics.v_measure_score(labels_true, labels_pred, *, beta=1.0)[source]#
V-measure cluster labeling given a ground truth.
This score is identical to
normalized_mutual_info_scorewith the'arithmetic'option for averaging.The V-measure is the harmonic mean between homogeneity and completeness:
v = (1 + beta) * homogeneity * completeness / (beta * homogeneity + completeness)
This metric is independent of the absolute values of the labels: a permutation of the class or cluster label values won’t change the score value in any way.
This metric is furthermore symmetric: switching
label_truewithlabel_predwill return the same score value. This can be useful to measure the agreement of two independent label assignments strategies on the same dataset when the real ground truth is not known.Read more in the User Guide.
- Parameters:
- labels_truearray-like of shape (n_samples,)
Ground truth class labels to be used as a reference.
- labels_predarray-like of shape (n_samples,)
Cluster labels to evaluate.
- betafloat, default=1.0
Ratio of weight attributed to
homogeneityvscompleteness. Ifbetais greater than 1,completenessis weighted more strongly in the calculation. Ifbetais less than 1,homogeneityis weighted more strongly.
- Returns:
- v_measurefloat
Score between 0.0 and 1.0. 1.0 stands for perfectly complete labeling.
See also
homogeneity_scoreHomogeneity metric of cluster labeling.
completeness_scoreCompleteness metric of cluster labeling.
normalized_mutual_info_scoreNormalized Mutual Information.
References
Examples
Perfect labelings are both homogeneous and complete, hence have score 1.0:
>>> from sklearn.metrics.cluster import v_measure_score >>> v_measure_score([0, 0, 1, 1], [0, 0, 1, 1]) 1.0 >>> v_measure_score([0, 0, 1, 1], [1, 1, 0, 0]) 1.0
Labelings that assign all classes members to the same clusters are complete but not homogeneous, hence penalized:
>>> print("%.6f" % v_measure_score([0, 0, 1, 2], [0, 0, 1, 1])) 0.8 >>> print("%.6f" % v_measure_score([0, 1, 2, 3], [0, 0, 1, 1])) 0.67
Labelings that have pure clusters with members coming from the same classes are homogeneous but un-necessary splits harm completeness and thus penalize V-measure as well:
>>> print("%.6f" % v_measure_score([0, 0, 1, 1], [0, 0, 1, 2])) 0.8 >>> print("%.6f" % v_measure_score([0, 0, 1, 1], [0, 1, 2, 3])) 0.67
If classes members are completely split across different clusters, the assignment is totally incomplete, hence the V-Measure is null:
>>> print("%.6f" % v_measure_score([0, 0, 0, 0], [0, 1, 2, 3])) 0.0
Clusters that include samples from totally different classes totally destroy the homogeneity of the labeling, hence:
>>> print("%.6f" % v_measure_score([0, 0, 1, 1], [0, 0, 0, 0])) 0.0
Gallery examples#

Biclustering documents with the Spectral Co-clustering algorithm
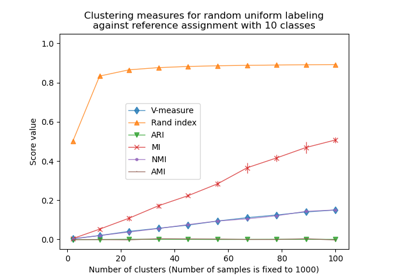
Adjustment for chance in clustering performance evaluation
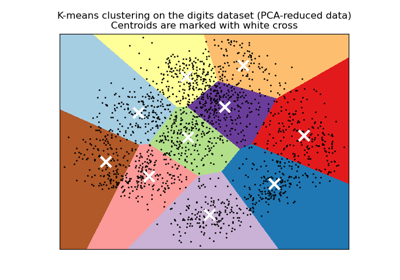
A demo of K-Means clustering on the handwritten digits data