mutual_info_score#
- sklearn.metrics.mutual_info_score(labels_true, labels_pred, *, contingency=None)[source]#
Mutual Information between two clusterings.
The Mutual Information is a measure of the similarity between two labels of the same data. Where \(|U_i|\) is the number of the samples in cluster \(U_i\) and \(|V_j|\) is the number of the samples in cluster \(V_j\), the Mutual Information between clusterings \(U\) and \(V\) is given as:
\[MI(U,V)=\sum_{i=1}^{|U|} \sum_{j=1}^{|V|} \frac{|U_i\cap V_j|}{N} \log\frac{N|U_i \cap V_j|}{|U_i||V_j|}\]This metric is independent of the absolute values of the labels: a permutation of the class or cluster label values won’t change the score value in any way.
This metric is furthermore symmetric: switching \(U\) (i.e
label_true) with \(V\) (i.e.label_pred) will return the same score value. This can be useful to measure the agreement of two independent label assignments strategies on the same dataset when the real ground truth is not known.Read more in the User Guide.
- Parameters:
- labels_truearray-like of shape (n_samples,)
A clustering of the data into disjoint subsets, called \(U\) in the above formula.
- labels_predarray-like of shape (n_samples,)
A clustering of the data into disjoint subsets, called \(V\) in the above formula.
- contingency{array-like, sparse matrix} of shape (n_classes_true, n_classes_pred), default=None
A contingency matrix given by the
contingency_matrixfunction. If value isNone, it will be computed, otherwise the given value is used, withlabels_trueandlabels_predignored.
- Returns:
- mifloat
Mutual information, a non-negative value, measured in nats using the natural logarithm.
See also
adjusted_mutual_info_scoreAdjusted against chance Mutual Information.
normalized_mutual_info_scoreNormalized Mutual Information.
Notes
The logarithm used is the natural logarithm (base-e).
Examples
>>> from sklearn.metrics import mutual_info_score >>> labels_true = [0, 1, 1, 0, 1, 0] >>> labels_pred = [0, 1, 0, 0, 1, 1] >>> mutual_info_score(labels_true, labels_pred) 0.0566
Gallery examples#
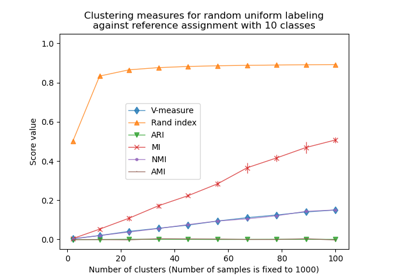
Adjustment for chance in clustering performance evaluation
