CalibratedClassifierCV#
- class sklearn.calibration.CalibratedClassifierCV(estimator=None, *, method='sigmoid', cv=None, n_jobs=None, ensemble='auto')[source]#
Calibrate probabilities using isotonic, sigmoid, or temperature scaling.
This class uses cross-validation to both estimate the parameters of a classifier and subsequently calibrate a classifier. With
ensemble=True, for each cv split it fits a copy of the base estimator to the training subset, and calibrates it using the testing subset. For prediction, predicted probabilities are averaged across these individual calibrated classifiers. Whenensemble=False, cross-validation is used to obtain unbiased predictions, viacross_val_predict, which are then used for calibration. For prediction, the base estimator, trained using all the data, is used. This is the prediction method implemented whenprobabilities=TrueforSVCandNuSVCestimators (see User Guide for details).Already fitted classifiers can be calibrated by wrapping the model in a
FrozenEstimator. In this case all provided data is used for calibration. The user has to take care manually that data for model fitting and calibration are disjoint.The calibration is based on the decision_function method of the
estimatorif it exists, else on predict_proba.Read more in the User Guide. In order to learn more on the CalibratedClassifierCV class, see the following calibration examples: Probability calibration of classifiers, Probability Calibration curves, and Probability Calibration for 3-class classification.
- Parameters:
- estimatorestimator instance, default=None
The classifier whose output need to be calibrated to provide more accurate
predict_probaoutputs. The default classifier is aLinearSVC.Added in version 1.2.
- method{‘sigmoid’, ‘isotonic’, ‘temperature’}, default=’sigmoid’
The method to use for calibration. Can be:
‘sigmoid’, which corresponds to Platt’s method (i.e. a binary logistic regression model).
‘isotonic’, which is a non-parametric approach.
‘temperature’, temperature scaling.
Sigmoid and isotonic calibration methods natively support only binary classifiers and extend to multi-class classification using a One-vs-Rest (OvR) strategy with post-hoc renormalization, i.e., adjusting the probabilities after calibration to ensure they sum up to 1.
In contrast, temperature scaling naturally supports multi-class calibration by applying
softmax(classifier_logits/T)with a value ofT(temperature) that optimizes the log loss.For very uncalibrated classifiers on very imbalanced datasets, sigmoid calibration might be preferred because it fits an additional intercept parameter. This helps shift decision boundaries appropriately when the classifier being calibrated is biased towards the majority class.
Isotonic calibration is not recommended when the number of calibration samples is too low
(≪1000)since it then tends to overfit.Changed in version 1.8: Added option ‘temperature’.
- cvint, cross-validation generator, or iterable, default=None
Determines the cross-validation splitting strategy. Possible inputs for cv are:
None, to use the default 5-fold cross-validation,
integer, to specify the number of folds.
An iterable yielding (train, test) splits as arrays of indices.
For integer/None inputs, if
yis binary or multiclass,StratifiedKFoldis used. Ifyis neither binary nor multiclass,KFoldis used.Refer to the User Guide for the various cross-validation strategies that can be used here.
Changed in version 0.22:
cvdefault value if None changed from 3-fold to 5-fold.- n_jobsint, default=None
Number of jobs to run in parallel.
Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors.Base estimator clones are fitted in parallel across cross-validation iterations.
See Glossary for more details.
Added in version 0.24.
- ensemblebool, or “auto”, default=”auto”
Determines how the calibrator is fitted.
“auto” will use
Falseif theestimatoris aFrozenEstimator, andTrueotherwise.If
True, theestimatoris fitted using training data, and calibrated using testing data, for eachcvfold. The final estimator is an ensemble ofn_cvfitted classifier and calibrator pairs, wheren_cvis the number of cross-validation folds. The output is the average predicted probabilities of all pairs.If
False,cvis used to compute unbiased predictions, viacross_val_predict, which are then used for calibration. At prediction time, the classifier used is theestimatortrained on all the data. Note that this method is also internally implemented insklearn.svmestimators with theprobabilities=Trueparameter.Added in version 0.24.
Changed in version 1.6:
"auto"option is added and is the default.
- Attributes:
- classes_ndarray of shape (n_classes,)
The class labels.
- n_features_in_int
Number of features seen during fit. Only defined if the underlying estimator exposes such an attribute when fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Only defined if the underlying estimator exposes such an attribute when fit.
Added in version 1.0.
- calibrated_classifiers_list (len() equal to cv or 1 if
ensemble=False) The list of classifier and calibrator pairs.
When
ensemble=True,n_cvfittedestimatorand calibrator pairs.n_cvis the number of cross-validation folds.When
ensemble=False, theestimator, fitted on all the data, and fitted calibrator.
Changed in version 0.24: Single calibrated classifier case when
ensemble=False.
See also
calibration_curveCompute true and predicted probabilities for a calibration curve.
References
[1]B. Zadrozny & C. Elkan. Obtaining calibrated probability estimates from decision trees and naive Bayesian classifiers, ICML 2001.
[2]B. Zadrozny & C. Elkan. Transforming Classifier Scores into Accurate Multiclass Probability Estimates, KDD 2002.
[3]J. Platt. Probabilistic Outputs for Support Vector Machines and Comparisons to Regularized Likelihood Methods, 1999.
[4]A. Niculescu-Mizil & R. Caruana. Predicting Good Probabilities with Supervised Learning, ICML 2005.
[5]Chuan Guo, Geoff Pleiss, Yu Sun, Kilian Q. Weinberger. On Calibration of Modern Neural Networks. Proceedings of the 34th International Conference on Machine Learning, PMLR 70:1321-1330, 2017.
Examples
>>> from sklearn.datasets import make_classification >>> from sklearn.naive_bayes import GaussianNB >>> from sklearn.calibration import CalibratedClassifierCV >>> X, y = make_classification(n_samples=100, n_features=2, ... n_redundant=0, random_state=42) >>> base_clf = GaussianNB() >>> calibrated_clf = CalibratedClassifierCV(base_clf, cv=3) >>> calibrated_clf.fit(X, y) CalibratedClassifierCV(...) >>> len(calibrated_clf.calibrated_classifiers_) 3 >>> calibrated_clf.predict_proba(X)[:5, :] array([[0.110, 0.889], [0.072, 0.927], [0.928, 0.072], [0.928, 0.072], [0.072, 0.928]]) >>> from sklearn.model_selection import train_test_split >>> X, y = make_classification(n_samples=100, n_features=2, ... n_redundant=0, random_state=42) >>> X_train, X_calib, y_train, y_calib = train_test_split( ... X, y, random_state=42 ... ) >>> base_clf = GaussianNB() >>> base_clf.fit(X_train, y_train) GaussianNB() >>> from sklearn.frozen import FrozenEstimator >>> calibrated_clf = CalibratedClassifierCV(FrozenEstimator(base_clf)) >>> calibrated_clf.fit(X_calib, y_calib) CalibratedClassifierCV(...) >>> len(calibrated_clf.calibrated_classifiers_) 1 >>> calibrated_clf.predict_proba([[-0.5, 0.5]]) array([[0.936, 0.063]])
- fit(X, y, sample_weight=None, **fit_params)[source]#
Fit the calibrated model.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Training data.
- yarray-like of shape (n_samples,)
Target values.
- sample_weightarray-like of shape (n_samples,), default=None
Sample weights. If None, then samples are equally weighted.
- **fit_paramsdict
Parameters to pass to the
fitmethod of the underlying classifier.
- Returns:
- selfobject
Returns an instance of self.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRouter
A
MetadataRouterencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- predict(X)[source]#
Predict the target of new samples.
The predicted class is the class that has the highest probability, and can thus be different from the prediction of the uncalibrated classifier.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
The samples, as accepted by
estimator.predict.
- Returns:
- Cndarray of shape (n_samples,)
The predicted class.
- predict_proba(X)[source]#
Calibrated probabilities of classification.
This function returns calibrated probabilities of classification according to each class on an array of test vectors X.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
The samples, as accepted by
estimator.predict_proba.
- Returns:
- Cndarray of shape (n_samples, n_classes)
The predicted probas.
- score(X, y, sample_weight=None)[source]#
Return accuracy on provided data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True labels for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
Mean accuracy of
self.predict(X)w.r.t.y.
- set_fit_request(*, sample_weight: bool | None | str = '$UNCHANGED$') CalibratedClassifierCV[source]#
Configure whether metadata should be requested to be passed to the
fitmethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter infit.
- Returns:
- selfobject
The updated object.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') CalibratedClassifierCV[source]#
Configure whether metadata should be requested to be passed to the
scoremethod.Note that this method is only relevant when this estimator is used as a sub-estimator within a meta-estimator and metadata routing is enabled with
enable_metadata_routing=True(seesklearn.set_config). Please check the User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.
Gallery examples#
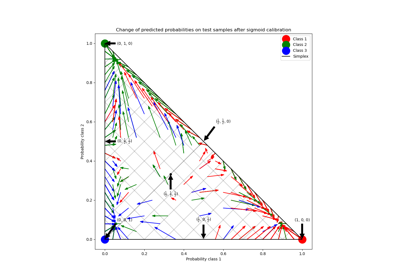
Probability Calibration for 3-class classification