accuracy_score#
- sklearn.metrics.accuracy_score(y_true, y_pred, *, normalize=True, sample_weight=None)[source]#
Accuracy classification score.
In multilabel classification, this function computes subset accuracy: the set of labels predicted for a sample must exactly match the corresponding set of labels in y_true.
Read more in the User Guide.
- Parameters:
- y_true1d array-like, or label indicator array / sparse matrix
Ground truth (correct) labels.
- y_pred1d array-like, or label indicator array / sparse matrix
Predicted labels, as returned by a classifier.
- normalizebool, default=True
If
False, return the number of correctly classified samples. Otherwise, return the fraction of correctly classified samples.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat or int
If
normalize == True, return the fraction of correctly classified samples (float), else returns the number of correctly classified samples (int).The best performance is 1 with
normalize == Trueand the number of samples withnormalize == False.
See also
balanced_accuracy_scoreCompute the balanced accuracy to deal with imbalanced datasets.
jaccard_scoreCompute the Jaccard similarity coefficient score.
hamming_lossCompute the average Hamming loss or Hamming distance between two sets of samples.
zero_one_lossCompute the Zero-one classification loss. By default, the function will return the percentage of imperfectly predicted subsets.
Examples
>>> from sklearn.metrics import accuracy_score >>> y_pred = [0, 2, 1, 3] >>> y_true = [0, 1, 2, 3] >>> accuracy_score(y_true, y_pred) 0.5 >>> accuracy_score(y_true, y_pred, normalize=False) 2.0
In the multilabel case with binary label indicators:
>>> import numpy as np >>> accuracy_score(np.array([[0, 1], [1, 1]]), np.ones((2, 2))) 0.5
Gallery examples#
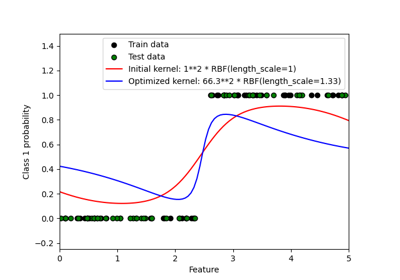
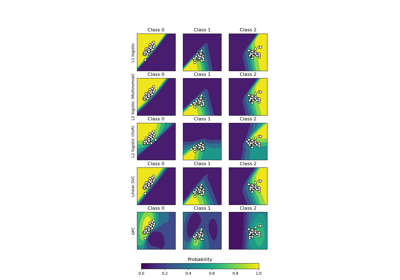
Probabilistic predictions with Gaussian process classification (GPC)
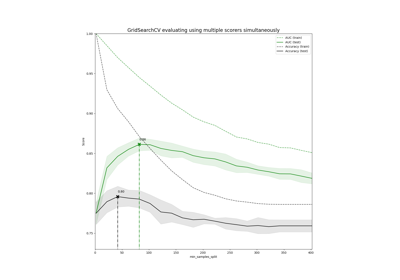
Demonstration of multi-metric evaluation on cross_val_score and GridSearchCV
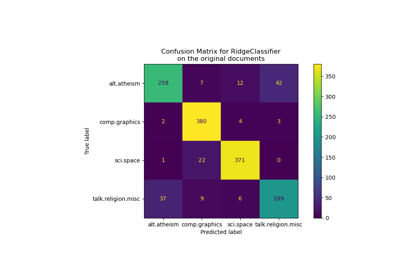
Classification of text documents using sparse features