sklearn.metrics.normalized_mutual_info_score¶
- sklearn.metrics.normalized_mutual_info_score(labels_true, labels_pred, *, average_method='arithmetic')[source]¶
Normalized Mutual Information between two clusterings.
Normalized Mutual Information (NMI) is a normalization of the Mutual Information (MI) score to scale the results between 0 (no mutual information) and 1 (perfect correlation). In this function, mutual information is normalized by some generalized mean of
H(labels_true)andH(labels_pred)), defined by theaverage_method.This measure is not adjusted for chance. Therefore
adjusted_mutual_info_scoremight be preferred.This metric is independent of the absolute values of the labels: a permutation of the class or cluster label values won’t change the score value in any way.
This metric is furthermore symmetric: switching
label_truewithlabel_predwill return the same score value. This can be useful to measure the agreement of two independent label assignments strategies on the same dataset when the real ground truth is not known.Read more in the User Guide.
- Parameters:
- labels_trueint array-like of shape (n_samples,)
A clustering of the data into disjoint subsets.
- labels_predint array-like of shape (n_samples,)
A clustering of the data into disjoint subsets.
- average_method{‘min’, ‘geometric’, ‘arithmetic’, ‘max’}, default=’arithmetic’
How to compute the normalizer in the denominator.
New in version 0.20.
Changed in version 0.22: The default value of
average_methodchanged from ‘geometric’ to ‘arithmetic’.
- Returns:
- nmifloat
Score between 0.0 and 1.0 in normalized nats (based on the natural logarithm). 1.0 stands for perfectly complete labeling.
See also
v_measure_scoreV-Measure (NMI with arithmetic mean option).
adjusted_rand_scoreAdjusted Rand Index.
adjusted_mutual_info_scoreAdjusted Mutual Information (adjusted against chance).
Examples
Perfect labelings are both homogeneous and complete, hence have score 1.0:
>>> from sklearn.metrics.cluster import normalized_mutual_info_score >>> normalized_mutual_info_score([0, 0, 1, 1], [0, 0, 1, 1]) ... 1.0 >>> normalized_mutual_info_score([0, 0, 1, 1], [1, 1, 0, 0]) ... 1.0
If classes members are completely split across different clusters, the assignment is totally in-complete, hence the NMI is null:
>>> normalized_mutual_info_score([0, 0, 0, 0], [0, 1, 2, 3]) ... 0.0
Examples using sklearn.metrics.normalized_mutual_info_score¶
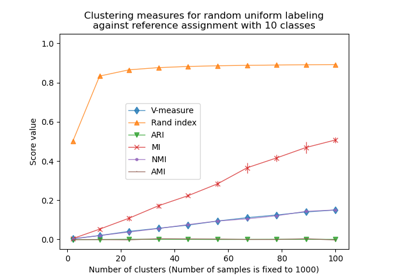
Adjustment for chance in clustering performance evaluation