sklearn.semi_supervised.LabelSpreading¶
- class sklearn.semi_supervised.LabelSpreading(kernel='rbf', *, gamma=20, n_neighbors=7, alpha=0.2, max_iter=30, tol=0.001, n_jobs=None)[source]¶
LabelSpreading model for semi-supervised learning.
This model is similar to the basic Label Propagation algorithm, but uses affinity matrix based on the normalized graph Laplacian and soft clamping across the labels.
Read more in the User Guide.
- Parameters:
- kernel{‘knn’, ‘rbf’} or callable, default=’rbf’
String identifier for kernel function to use or the kernel function itself. Only ‘rbf’ and ‘knn’ strings are valid inputs. The function passed should take two inputs, each of shape (n_samples, n_features), and return a (n_samples, n_samples) shaped weight matrix.
- gammafloat, default=20
Parameter for rbf kernel.
- n_neighborsint, default=7
Parameter for knn kernel which is a strictly positive integer.
- alphafloat, default=0.2
Clamping factor. A value in (0, 1) that specifies the relative amount that an instance should adopt the information from its neighbors as opposed to its initial label. alpha=0 means keeping the initial label information; alpha=1 means replacing all initial information.
- max_iterint, default=30
Maximum number of iterations allowed.
- tolfloat, default=1e-3
Convergence tolerance: threshold to consider the system at steady state.
- n_jobsint, default=None
The number of parallel jobs to run.
Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.
- Attributes:
- X_ndarray of shape (n_samples, n_features)
Input array.
- classes_ndarray of shape (n_classes,)
The distinct labels used in classifying instances.
- label_distributions_ndarray of shape (n_samples, n_classes)
Categorical distribution for each item.
- transduction_ndarray of shape (n_samples,)
Label assigned to each item during fit.
- n_features_in_int
Number of features seen during fit.
New in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.New in version 1.0.
- n_iter_int
Number of iterations run.
See also
LabelPropagationUnregularized graph based semi-supervised learning.
References
Examples
>>> import numpy as np >>> from sklearn import datasets >>> from sklearn.semi_supervised import LabelSpreading >>> label_prop_model = LabelSpreading() >>> iris = datasets.load_iris() >>> rng = np.random.RandomState(42) >>> random_unlabeled_points = rng.rand(len(iris.target)) < 0.3 >>> labels = np.copy(iris.target) >>> labels[random_unlabeled_points] = -1 >>> label_prop_model.fit(iris.data, labels) LabelSpreading(...)
Methods
fit(X, y)Fit a semi-supervised label propagation model to X.
Get metadata routing of this object.
get_params([deep])Get parameters for this estimator.
predict(X)Perform inductive inference across the model.
Predict probability for each possible outcome.
score(X, y[, sample_weight])Return the mean accuracy on the given test data and labels.
set_params(**params)Set the parameters of this estimator.
set_score_request(*[, sample_weight])Request metadata passed to the
scoremethod.- fit(X, y)[source]¶
Fit a semi-supervised label propagation model to X.
The input samples (labeled and unlabeled) are provided by matrix X, and target labels are provided by matrix y. We conventionally apply the label -1 to unlabeled samples in matrix y in a semi-supervised classification.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
Training data, where
n_samplesis the number of samples andn_featuresis the number of features.- yarray-like of shape (n_samples,)
Target class values with unlabeled points marked as -1. All unlabeled samples will be transductively assigned labels internally, which are stored in
transduction_.
- Returns:
- selfobject
Returns the instance itself.
- get_metadata_routing()[source]¶
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]¶
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- predict(X)[source]¶
Perform inductive inference across the model.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
The data matrix.
- Returns:
- yndarray of shape (n_samples,)
Predictions for input data.
- predict_proba(X)[source]¶
Predict probability for each possible outcome.
Compute the probability estimates for each single sample in X and each possible outcome seen during training (categorical distribution).
- Parameters:
- Xarray-like of shape (n_samples, n_features)
The data matrix.
- Returns:
- probabilitiesndarray of shape (n_samples, n_classes)
Normalized probability distributions across class labels.
- score(X, y, sample_weight=None)[source]¶
Return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True labels for
X.- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
Mean accuracy of
self.predict(X)w.r.t.y.
- set_params(**params)[source]¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- set_score_request(*, sample_weight: bool | None | str = '$UNCHANGED$') LabelSpreading[source]¶
Request metadata passed to the
scoremethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed toscoreif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it toscore.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.New in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.- Parameters:
- sample_weightstr, True, False, or None, default=sklearn.utils.metadata_routing.UNCHANGED
Metadata routing for
sample_weightparameter inscore.
- Returns:
- selfobject
The updated object.
Examples using sklearn.semi_supervised.LabelSpreading¶
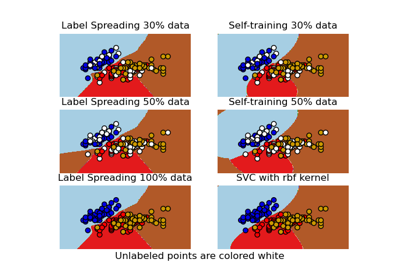
Decision boundary of semi-supervised classifiers versus SVM on the Iris dataset
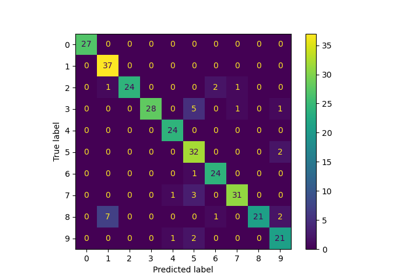
Label Propagation digits: Demonstrating performance