cross_validate#
- sklearn.model_selection.cross_validate(estimator, X, y=None, *, groups=None, scoring=None, cv=None, n_jobs=None, verbose=0, fit_params=None, params=None, pre_dispatch='2*n_jobs', return_train_score=False, return_estimator=False, return_indices=False, error_score=nan)[source]#
Evaluate metric(s) by cross-validation and also record fit/score times.
Read more in the User Guide.
- Parameters:
- estimatorestimator object implementing ‘fit’
The object to use to fit the data.
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The data to fit. Can be for example a list, or an array.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs), default=None
The target variable to try to predict in the case of supervised learning.
- groupsarray-like of shape (n_samples,), default=None
Group labels for the samples used while splitting the dataset into train/test set. Only used in conjunction with a “Group” cv instance (e.g.,
GroupKFold).Changed in version 1.4:
groupscan only be passed if metadata routing is not enabled viasklearn.set_config(enable_metadata_routing=True). When routing is enabled, passgroupsalongside other metadata via theparamsargument instead. E.g.:cross_validate(..., params={'groups': groups}).- scoringstr, callable, list, tuple, or dict, default=None
Strategy to evaluate the performance of the cross-validated model on the test set.
If
scoringrepresents a single score, one can use:a single string (see The scoring parameter: defining model evaluation rules);
a callable (see Defining your scoring strategy from metric functions) that returns a single value.
If
scoringrepresents multiple scores, one can use:a list or tuple of unique strings;
a callable returning a dictionary where the keys are the metric names and the values are the metric scores;
a dictionary with metric names as keys and callables a values.
See Specifying multiple metrics for evaluation for an example.
- cvint, cross-validation generator or an iterable, default=None
Determines the cross-validation splitting strategy. Possible inputs for cv are:
None, to use the default 5-fold cross validation,
int, to specify the number of folds in a
(Stratified)KFold,An iterable yielding (train, test) splits as arrays of indices.
For int/None inputs, if the estimator is a classifier and
yis either binary or multiclass,StratifiedKFoldis used. In all other cases,KFoldis used. These splitters are instantiated withshuffle=Falseso the splits will be the same across calls.Refer User Guide for the various cross-validation strategies that can be used here.
Changed in version 0.22:
cvdefault value if None changed from 3-fold to 5-fold.- n_jobsint, default=None
Number of jobs to run in parallel. Training the estimator and computing the score are parallelized over the cross-validation splits.
Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.- verboseint, default=0
The verbosity level.
- fit_paramsdict, default=None
Parameters to pass to the fit method of the estimator.
Deprecated since version 1.4: This parameter is deprecated and will be removed in version 1.6. Use
paramsinstead.- paramsdict, default=None
Parameters to pass to the underlying estimator’s
fit, the scorer, and the CV splitter.Added in version 1.4.
- pre_dispatchint or str, default=’2*n_jobs’
Controls the number of jobs that get dispatched during parallel execution. Reducing this number can be useful to avoid an explosion of memory consumption when more jobs get dispatched than CPUs can process. This parameter can be:
An int, giving the exact number of total jobs that are spawned
A str, giving an expression as a function of n_jobs, as in ‘2*n_jobs’
- return_train_scorebool, default=False
Whether to include train scores. Computing training scores is used to get insights on how different parameter settings impact the overfitting/underfitting trade-off. However computing the scores on the training set can be computationally expensive and is not strictly required to select the parameters that yield the best generalization performance.
Added in version 0.19.
Changed in version 0.21: Default value was changed from
TruetoFalse- return_estimatorbool, default=False
Whether to return the estimators fitted on each split.
Added in version 0.20.
- return_indicesbool, default=False
Whether to return the train-test indices selected for each split.
Added in version 1.3.
- error_score‘raise’ or numeric, default=np.nan
Value to assign to the score if an error occurs in estimator fitting. If set to ‘raise’, the error is raised. If a numeric value is given, FitFailedWarning is raised.
Added in version 0.20.
- Returns:
- scoresdict of float arrays of shape (n_splits,)
Array of scores of the estimator for each run of the cross validation.
A dict of arrays containing the score/time arrays for each scorer is returned. The possible keys for this
dictare:test_scoreThe score array for test scores on each cv split. Suffix
_scoreintest_scorechanges to a specific metric liketest_r2ortest_aucif there are multiple scoring metrics in the scoring parameter.train_scoreThe score array for train scores on each cv split. Suffix
_scoreintrain_scorechanges to a specific metric liketrain_r2ortrain_aucif there are multiple scoring metrics in the scoring parameter. This is available only ifreturn_train_scoreparameter isTrue.fit_timeThe time for fitting the estimator on the train set for each cv split.
score_timeThe time for scoring the estimator on the test set for each cv split. (Note time for scoring on the train set is not included even if
return_train_scoreis set toTrueestimatorThe estimator objects for each cv split. This is available only if
return_estimatorparameter is set toTrue.indicesThe train/test positional indices for each cv split. A dictionary is returned where the keys are either
"train"or"test"and the associated values are a list of integer-dtyped NumPy arrays with the indices. Available only ifreturn_indices=True.
See also
cross_val_scoreRun cross-validation for single metric evaluation.
cross_val_predictGet predictions from each split of cross-validation for diagnostic purposes.
sklearn.metrics.make_scorerMake a scorer from a performance metric or loss function.
Examples
>>> from sklearn import datasets, linear_model >>> from sklearn.model_selection import cross_validate >>> from sklearn.metrics import make_scorer >>> from sklearn.metrics import confusion_matrix >>> from sklearn.svm import LinearSVC >>> diabetes = datasets.load_diabetes() >>> X = diabetes.data[:150] >>> y = diabetes.target[:150] >>> lasso = linear_model.Lasso()
Single metric evaluation using
cross_validate>>> cv_results = cross_validate(lasso, X, y, cv=3) >>> sorted(cv_results.keys()) ['fit_time', 'score_time', 'test_score'] >>> cv_results['test_score'] array([0.3315057 , 0.08022103, 0.03531816])
Multiple metric evaluation using
cross_validate(please refer thescoringparameter doc for more information)>>> scores = cross_validate(lasso, X, y, cv=3, ... scoring=('r2', 'neg_mean_squared_error'), ... return_train_score=True) >>> print(scores['test_neg_mean_squared_error']) [-3635.5... -3573.3... -6114.7...] >>> print(scores['train_r2']) [0.28009951 0.3908844 0.22784907]
Gallery examples#
Common pitfalls in the interpretation of coefficients of linear models
Class Likelihood Ratios to measure classification performance
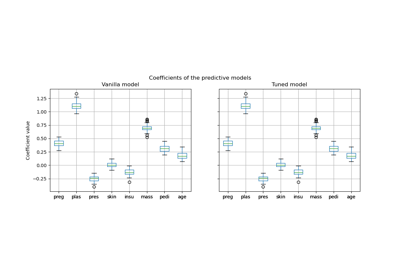
Post-hoc tuning the cut-off point of decision function