minmax_scale#
- sklearn.preprocessing.minmax_scale(X, feature_range=(0, 1), *, axis=0, copy=True)[source]#
Transform features by scaling each feature to a given range.
This estimator scales and translates each feature individually such that it is in the given range on the training set, i.e. between zero and one.
The transformation is given by (when
axis=0):X_std = (X - X.min(axis=0)) / (X.max(axis=0) - X.min(axis=0)) X_scaled = X_std * (max - min) + min
where min, max = feature_range.
The transformation is calculated as (when
axis=0):X_scaled = scale * X + min - X.min(axis=0) * scale where scale = (max - min) / (X.max(axis=0) - X.min(axis=0))
This transformation is often used as an alternative to zero mean, unit variance scaling.
Read more in the User Guide.
Added in version 0.17: minmax_scale function interface to
MinMaxScaler.- Parameters:
- Xarray-like of shape (n_samples, n_features)
The data.
- feature_rangetuple (min, max), default=(0, 1)
Desired range of transformed data.
- axis{0, 1}, default=0
Axis used to scale along. If 0, independently scale each feature, otherwise (if 1) scale each sample.
- copybool, default=True
If False, try to avoid a copy and scale in place. This is not guaranteed to always work in place; e.g. if the data is a numpy array with an int dtype, a copy will be returned even with copy=False.
- Returns:
- X_trndarray of shape (n_samples, n_features)
The transformed data.
Warning
Risk of data leak Do not use
minmax_scaleunless you know what you are doing. A common mistake is to apply it to the entire data before splitting into training and test sets. This will bias the model evaluation because information would have leaked from the test set to the training set. In general, we recommend usingMinMaxScalerwithin a Pipeline in order to prevent most risks of data leaking:pipe = make_pipeline(MinMaxScaler(), LogisticRegression()).
See also
MinMaxScalerPerforms scaling to a given range using the Transformer API (e.g. as part of a preprocessing
Pipeline).
Notes
For a comparison of the different scalers, transformers, and normalizers, see: Compare the effect of different scalers on data with outliers.
Examples
>>> from sklearn.preprocessing import minmax_scale >>> X = [[-2, 1, 2], [-1, 0, 1]] >>> minmax_scale(X, axis=0) # scale each column independently array([[0., 1., 1.], [1., 0., 0.]]) >>> minmax_scale(X, axis=1) # scale each row independently array([[0. , 0.75, 1. ], [0. , 0.5 , 1. ]])
Gallery examples#

Restricted Boltzmann Machine features for digit classification
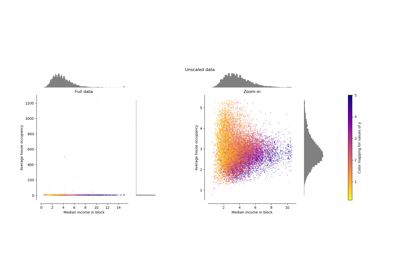
Compare the effect of different scalers on data with outliers
