Older Versions#
Version 0.12.1#
October 8, 2012
The 0.12.1 release is a bug-fix release with no additional features, but is instead a set of bug fixes
Changelog#
Improved numerical stability in spectral embedding by Gael Varoquaux
Doctest under windows 64bit by Gael Varoquaux
Documentation fixes for elastic net by Andreas Müller and Alexandre Gramfort
Proper behavior with fortran-ordered NumPy arrays by Gael Varoquaux
Make GridSearchCV work with non-CSR sparse matrix by Lars Buitinck
Fix parallel computing in MDS by Gael Varoquaux
Fix Unicode support in count vectorizer by Andreas Müller
Fix MinCovDet breaking with X.shape = (3, 1) by Virgile Fritsch
Fix clone of SGD objects by Peter Prettenhofer
Stabilize GMM by Virgile Fritsch
People#
Version 0.12#
September 4, 2012
Changelog#
Various speed improvements of the decision trees module, by Gilles Louppe.
GradientBoostingRegressorandGradientBoostingClassifiernow support feature subsampling via themax_featuresargument, by Peter Prettenhofer.Added Huber and Quantile loss functions to
GradientBoostingRegressor, by Peter Prettenhofer.Decision trees and forests of randomized trees now support multi-output classification and regression problems, by Gilles Louppe.
Added
LabelEncoder, a simple utility class to normalize labels or transform non-numerical labels, by Mathieu Blondel.Added the epsilon-insensitive loss and the ability to make probabilistic predictions with the modified huber loss in Stochastic Gradient Descent, by Mathieu Blondel.
Added Multi-dimensional Scaling (MDS), by Nelle Varoquaux.
SVMlight file format loader now detects compressed (gzip/bzip2) files and decompresses them on the fly, by Lars Buitinck.
SVMlight file format serializer now preserves double precision floating point values, by Olivier Grisel.
A common testing framework for all estimators was added, by Andreas Müller.
Understandable error messages for estimators that do not accept sparse input by Gael Varoquaux
Speedups in hierarchical clustering by Gael Varoquaux. In particular building the tree now supports early stopping. This is useful when the number of clusters is not small compared to the number of samples.
Add MultiTaskLasso and MultiTaskElasticNet for joint feature selection, by Alexandre Gramfort.
Added
metrics.auc_scoreandmetrics.average_precision_scoreconvenience functions by Andreas Müller.Improved sparse matrix support in the Feature selection module by Andreas Müller.
New word boundaries-aware character n-gram analyzer for the Text feature extraction module by @kernc.
Fixed bug in spectral clustering that led to single point clusters by Andreas Müller.
In
CountVectorizer, added an option to ignore infrequent words,min_dfby Andreas Müller.Add support for multiple targets in some linear models (ElasticNet, Lasso and OrthogonalMatchingPursuit) by Vlad Niculae and Alexandre Gramfort.
Fixes in
decomposition.ProbabilisticPCAscore function by Wei Li.Fixed feature importance computation in Gradient-boosted trees.
API changes summary#
The old
scikits.learnpackage has disappeared; all code should import fromsklearninstead, which was introduced in 0.9.In
metrics.roc_curve, thethresholdsarray is now returned with its order reversed, in order to keep it consistent with the order of the returnedfprandtpr.In
hmmobjects, likehmm.GaussianHMM,hmm.MultinomialHMM, etc., all parameters must be passed to the object when initialising it and not throughfit. Nowfitwill only accept the data as an input parameter.For all SVM classes, a faulty behavior of
gammawas fixed. Previously, the default gamma value was only computed the first timefitwas called and then stored. It is now recalculated on every call tofit.All
Baseclasses are now abstract meta classes so that they can not be instantiated.cluster.ward_treenow also returns the parent array. This is necessary for early-stopping in which case the tree is not completely built.In
CountVectorizerthe parametersmin_nandmax_nwere joined to the parametern_gram_rangeto enable grid-searching both at once.In
CountVectorizer, words that appear only in one document are now ignored by default. To reproduce the previous behavior, setmin_df=1.Fixed API inconsistency:
linear_model.SGDClassifier.predict_probanow returns 2d array when fit on two classes.Fixed API inconsistency:
discriminant_analysis.QuadraticDiscriminantAnalysis.decision_functionanddiscriminant_analysis.LinearDiscriminantAnalysis.decision_functionnow return 1d arrays when fit on two classes.Grid of alphas used for fitting
LassoCVandElasticNetCVis now stored in the attributealphas_rather than overriding the init parameteralphas.Linear models when alpha is estimated by cross-validation store the estimated value in the
alpha_attribute rather than justalphaorbest_alpha.GradientBoostingClassifiernow supportsstaged_predict_proba, andstaged_predict.svm.sparse.SVCand other sparse SVM classes are now deprecated. All classes in the Support Vector Machines module now automatically select the sparse or dense representation based on the input.All clustering algorithms now interpret the array
Xgiven tofitas input data, in particularSpectralClusteringandAffinityPropagationwhich previously expected affinity matrices.For clustering algorithms that take the desired number of clusters as a parameter, this parameter is now called
n_clusters.
People#
267 Andreas Müller
52 Vlad Niculae
44 Nelle Varoquaux
30 Alexis Mignon
30 Immanuel Bayer
16 Subhodeep Moitra
13 Yannick Schwartz
12 @kernc
9 Daniel Duckworth
8 John Benediktsson
7 Marko Burjek
4 Alexandre Abraham
3 Florian Hoenig
3 flyingimmidev
2 Francois Savard
2 Hannes Schulz
2 Peter Welinder
2 Wei Li
1 Alex Companioni
1 Brandyn A. White
1 Bussonnier Matthias
1 Charles-Pierre Astolfi
1 Dan O’Huiginn
1 David Cournapeau
1 Keith Goodman
1 Ludwig Schwardt
1 Olivier Hervieu
1 Sergio Medina
1 Shiqiao Du
1 Tim Sheerman-Chase
1 buguen
Version 0.11#
May 7, 2012
Changelog#
Highlights#
Gradient boosted regression trees (Gradient-boosted trees) for classification and regression by Peter Prettenhofer and Scott White .
Simple dict-based feature loader with support for categorical variables (
DictVectorizer) by Lars Buitinck.Added Matthews correlation coefficient (
metrics.matthews_corrcoef) and added macro and micro average options toprecision_score,metrics.recall_scoreandf1_scoreby Satrajit Ghosh.Out of Bag Estimates of generalization error for Ensembles: Gradient boosting, random forests, bagging, voting, stacking by Andreas Müller.
Randomized sparse linear models for feature selection, by Alexandre Gramfort and Gael Varoquaux
Label Propagation for semi-supervised learning, by Clay Woolam. Note the semi-supervised API is still work in progress, and may change.
Added BIC/AIC model selection to classical Gaussian mixture models and unified the API with the remainder of scikit-learn, by Bertrand Thirion
Added
sklearn.cross_validation.StratifiedShuffleSplit, which is asklearn.cross_validation.ShuffleSplitwith balanced splits, by Yannick Schwartz.NearestCentroidclassifier added, along with ashrink_thresholdparameter, which implements shrunken centroid classification, by Robert Layton.
Other changes#
Merged dense and sparse implementations of Stochastic Gradient Descent module and exposed utility extension types for sequential datasets
seq_datasetand weight vectorsweight_vectorby Peter Prettenhofer.Added
partial_fit(support for online/minibatch learning) and warm_start to the Stochastic Gradient Descent module by Mathieu Blondel.Dense and sparse implementations of Support Vector Machines classes and
LogisticRegressionmerged by Lars Buitinck.Regressors can now be used as base estimator in the Multiclass and multioutput algorithms module by Mathieu Blondel.
Added n_jobs option to
metrics.pairwise_distancesandmetrics.pairwise.pairwise_kernelsfor parallel computation, by Mathieu Blondel.K-means can now be run in parallel, using the
n_jobsargument to either K-means orcluster.KMeans, by Robert Layton.Improved Cross-validation: evaluating estimator performance and Tuning the hyper-parameters of an estimator documentation and introduced the new
cross_validation.train_test_splithelper function by Olivier GriselSVCmemberscoef_andintercept_changed sign for consistency withdecision_function; forkernel==linear,coef_was fixed in the one-vs-one case, by Andreas Müller.Performance improvements to efficient leave-one-out cross-validated Ridge regression, esp. for the
n_samples > n_featurescase, inRidgeCV, by Reuben Fletcher-Costin.Refactoring and simplification of the Text feature extraction API and fixed a bug that caused possible negative IDF, by Olivier Grisel.
Beam pruning option in
_BaseHMMmodule has been removed since it is difficult to Cythonize. If you are interested in contributing a Cython version, you can use the python version in the git history as a reference.Classes in Nearest Neighbors now support arbitrary Minkowski metric for nearest neighbors searches. The metric can be specified by argument
p.
API changes summary#
covariance.EllipticEnvelopis now deprecated. Please useEllipticEnvelopeinstead.NeighborsClassifierandNeighborsRegressorare gone in the module Nearest Neighbors. Use the classesKNeighborsClassifier,RadiusNeighborsClassifier,KNeighborsRegressorand/orRadiusNeighborsRegressorinstead.Sparse classes in the Stochastic Gradient Descent module are now deprecated.
In
mixture.GMM,mixture.DPGMMandmixture.VBGMM, parameters must be passed to an object when initialising it and not throughfit. Nowfitwill only accept the data as an input parameter.methods
rvsanddecodeinGMMmodule are now deprecated.sampleandscoreorpredictshould be used instead.attribute
_scoresand_pvaluesin univariate feature selection objects are now deprecated.scores_orpvalues_should be used instead.In
LogisticRegression,LinearSVC,SVCandNuSVC, theclass_weightparameter is now an initialization parameter, not a parameter to fit. This makes grid searches over this parameter possible.LFW
datais now always shape(n_samples, n_features)to be consistent with the Olivetti faces dataset. Useimagesandpairsattribute to access the natural images shapes instead.In
LinearSVC, the meaning of themulti_classparameter changed. Options now are'ovr'and'crammer_singer', with'ovr'being the default. This does not change the default behavior but hopefully is less confusing.Class
feature_selection.text.Vectorizeris deprecated and replaced byfeature_selection.text.TfidfVectorizer.The preprocessor / analyzer nested structure for text feature extraction has been removed. All those features are now directly passed as flat constructor arguments to
feature_selection.text.TfidfVectorizerandfeature_selection.text.CountVectorizer, in particular the following parameters are now used:analyzercan be'word'or'char'to switch the default analysis scheme, or use a specific python callable (as previously).tokenizerandpreprocessorhave been introduced to make it still possible to customize those steps with the new API.inputexplicitly control how to interpret the sequence passed tofitandpredict: filenames, file objects or direct (byte or Unicode) strings.charset decoding is explicit and strict by default.
the
vocabulary, fitted or not is now stored in thevocabulary_attribute to be consistent with the project conventions.Class
feature_selection.text.TfidfVectorizernow derives directly fromfeature_selection.text.CountVectorizerto make grid search trivial.methods
rvsin_BaseHMMmodule are now deprecated.sampleshould be used instead.Beam pruning option in
_BaseHMMmodule is removed since it is difficult to be Cythonized. If you are interested, you can look in the history codes by git.The SVMlight format loader now supports files with both zero-based and one-based column indices, since both occur “in the wild”.
Arguments in class
ShuffleSplitare now consistent withStratifiedShuffleSplit. Argumentstest_fractionandtrain_fractionare deprecated and renamed totest_sizeandtrain_sizeand can accept bothfloatandint.Arguments in class
Bootstrapare now consistent withStratifiedShuffleSplit. Argumentsn_testandn_trainare deprecated and renamed totest_sizeandtrain_sizeand can accept bothfloatandint.Argument
padded to classes in Nearest Neighbors to specify an arbitrary Minkowski metric for nearest neighbors searches.
People#
282 Andreas Müller
198 Gael Varoquaux
129 Olivier Grisel
114 Mathieu Blondel
103 Clay Woolam
28 flyingimmidev
26 Shiqiao Du
17 David Marek
14 Vlad Niculae
11 Yannick Schwartz
9 fcostin
7 Nick Wilson
5 Adrien Gaidon
5 Nelle Varoquaux
5 Emmanuelle Gouillart
3 Joonas Sillanpää
3 Paolo Losi
2 Charles McCarthy
2 Roy Hyunjin Han
2 Scott White
2 ibayer
1 Brandyn White
1 Carlos Scheidegger
1 Claire Revillet
1 Conrad Lee
1 Jan Hendrik Metzen
1 Meng Xinfan
1 Shiqiao
1 Udi Weinsberg
1 Virgile Fritsch
1 Xinfan Meng
1 Yaroslav Halchenko
1 jansoe
1 Leon Palafox
Version 0.10#
January 11, 2012
Changelog#
Python 2.5 compatibility was dropped; the minimum Python version needed to use scikit-learn is now 2.6.
Sparse inverse covariance estimation using the graph Lasso, with associated cross-validated estimator, by Gael Varoquaux
New Tree module by Brian Holt, Peter Prettenhofer, Satrajit Ghosh and Gilles Louppe. The module comes with complete documentation and examples.
Fixed a bug in the RFE module by Gilles Louppe (issue #378).
Fixed a memory leak in Support Vector Machines module by Brian Holt (issue #367).
Faster tests by Fabian Pedregosa and others.
Silhouette Coefficient cluster analysis evaluation metric added as
silhouette_scoreby Robert Layton.Fixed a bug in K-means in the handling of the
n_initparameter: the clustering algorithm used to be runn_inittimes but the last solution was retained instead of the best solution by Olivier Grisel.Minor refactoring in Stochastic Gradient Descent module; consolidated dense and sparse predict methods; Enhanced test time performance by converting model parameters to fortran-style arrays after fitting (only multi-class).
Adjusted Mutual Information metric added as
adjusted_mutual_info_scoreby Robert Layton.Models like SVC/SVR/LinearSVC/LogisticRegression from libsvm/liblinear now support scaling of C regularization parameter by the number of samples by Alexandre Gramfort.
New Ensemble Methods module by Gilles Louppe and Brian Holt. The module comes with the random forest algorithm and the extra-trees method, along with documentation and examples.
Novelty and Outlier Detection: outlier and novelty detection, by Virgile Fritsch.
Kernel Approximation: a transform implementing kernel approximation for fast SGD on non-linear kernels by Andreas Müller.
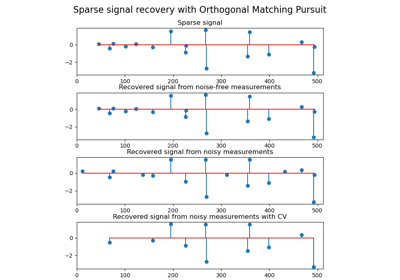
Fixed a bug due to atom swapping in Orthogonal Matching Pursuit (OMP) by Vlad Niculae.
Sparse coding with a precomputed dictionary by Vlad Niculae.
Mini Batch K-Means performance improvements by Olivier Grisel.
K-means support for sparse matrices by Mathieu Blondel.
Improved documentation for developers and for the
sklearn.utilsmodule, by Jake Vanderplas.Vectorized 20newsgroups dataset loader (
fetch_20newsgroups_vectorized) by Mathieu Blondel.Utilities for fast computation of mean and variance for sparse matrices by Mathieu Blondel.
Make
scaleandsklearn.preprocessing.Scalerwork on sparse matrices by Olivier GriselFeature importances using decision trees and/or forest of trees, by Gilles Louppe.
Parallel implementation of forests of randomized trees by Gilles Louppe.
sklearn.cross_validation.ShuffleSplitcan subsample the train sets as well as the test sets by Olivier Grisel.Errors in the build of the documentation fixed by Andreas Müller.
API changes summary#
Here are the code migration instructions when upgrading from scikit-learn version 0.9:
Some estimators that may overwrite their inputs to save memory previously had
overwrite_parameters; these have been replaced withcopy_parameters with exactly the opposite meaning.This particularly affects some of the estimators in
linear_model. The default behavior is still to copy everything passed in.The SVMlight dataset loader
load_svmlight_fileno longer supports loading two files at once; useload_svmlight_filesinstead. Also, the (unused)buffer_mbparameter is gone.Sparse estimators in the Stochastic Gradient Descent module use dense parameter vector
coef_instead ofsparse_coef_. This significantly improves test time performance.The Covariance estimation module now has a robust estimator of covariance, the Minimum Covariance Determinant estimator.
Cluster evaluation metrics in
clusterhave been refactored but the changes are backwards compatible. They have been moved to themetrics.cluster.supervised, along withmetrics.cluster.unsupervisedwhich contains the Silhouette Coefficient.The
permutation_test_scorefunction now behaves the same way ascross_val_score(i.e. uses the mean score across the folds.)Cross Validation generators now use integer indices (
indices=True) by default instead of boolean masks. This makes it more intuitive to use with sparse matrix data.The functions used for sparse coding,
sparse_encodeandsparse_encode_parallelhave been combined intosparse_encode, and the shapes of the arrays have been transposed for consistency with the matrix factorization setting, as opposed to the regression setting.Fixed an off-by-one error in the SVMlight/LibSVM file format handling; files generated using
dump_svmlight_fileshould be re-generated. (They should continue to work, but accidentally had one extra column of zeros prepended.)BaseDictionaryLearningclass replaced bySparseCodingMixin.sklearn.utils.extmath.fast_svdhas been renamedrandomized_svdand the default oversampling is now fixed to 10 additional random vectors instead of doubling the number of components to extract. The new behavior follows the reference paper.
People#
The following people contributed to scikit-learn since last release:
246 Andreas Müller
242 Olivier Grisel
220 Gilles Louppe
183 Brian Holt
166 Gael Varoquaux
144 Lars Buitinck
73 Vlad Niculae
60 Robert Layton
44 Noel Dawe
3 Jan Hendrik Metzen
3 Kenneth C. Arnold
3 Shiqiao Du
3 Tim Sheerman-Chase
2 Bala Subrahmanyam Varanasi
2 DraXus
2 Michael Eickenberg
1 Bogdan Trach
1 Félix-Antoine Fortin
1 Juan Manuel Caicedo Carvajal
1 Nelle Varoquaux
1 Tiziano Zito
1 Xinfan Meng
Version 0.9#
September 21, 2011
scikit-learn 0.9 was released on September 2011, three months after the 0.8 release and includes the new modules Manifold learning, The Dirichlet Process as well as several new algorithms and documentation improvements.
This release also includes the dictionary-learning work developed by Vlad Niculae as part of the Google Summer of Code program.
Changelog#
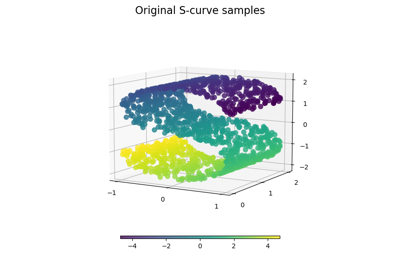
New Manifold learning module by Jake Vanderplas and Fabian Pedregosa.
New Dirichlet Process Gaussian Mixture Model by Alexandre Passos
Nearest Neighbors module refactoring by Jake Vanderplas : general refactoring, support for sparse matrices in input, speed and documentation improvements. See the next section for a full list of API changes.
Improvements on the Feature selection module by Gilles Louppe : refactoring of the RFE classes, documentation rewrite, increased efficiency and minor API changes.
Sparse principal components analysis (SparsePCA and MiniBatchSparsePCA) by Vlad Niculae, Gael Varoquaux and Alexandre Gramfort
Printing an estimator now behaves independently of architectures and Python version thanks to Jean Kossaifi.
Loader for libsvm/svmlight format by Mathieu Blondel and Lars Buitinck
Documentation improvements: thumbnails in example gallery by Fabian Pedregosa.
Important bugfixes in Support Vector Machines module (segfaults, bad performance) by Fabian Pedregosa.
Added Multinomial Naive Bayes and Bernoulli Naive Bayes by Lars Buitinck
Text feature extraction optimizations by Lars Buitinck
Chi-Square feature selection (
feature_selection.chi2) by Lars Buitinck.Generated datasets module refactoring by Gilles Louppe
Ball tree rewrite by Jake Vanderplas
Implementation of DBSCAN algorithm by Robert Layton
Kmeans predict and transform by Robert Layton
Preprocessing module refactoring by Olivier Grisel
Faster mean shift by Conrad Lee
New
Bootstrap, Random permutations cross-validation a.k.a. Shuffle & Split and various other improvements in cross validation schemes by Olivier Grisel and Gael VaroquauxAdjusted Rand index and V-Measure clustering evaluation metrics by Olivier Grisel
Added
Orthogonal Matching Pursuitby Vlad NiculaeAdded 2D-patch extractor utilities in the Feature extraction module by Vlad Niculae
Implementation of
LassoLarsCV(cross-validated Lasso solver using the Lars algorithm) andLassoLarsIC(BIC/AIC model selection in Lars) by Gael Varoquaux and Alexandre GramfortScalability improvements to
metrics.roc_curveby Olivier HervieuDistance helper functions
metrics.pairwise_distancesandmetrics.pairwise.pairwise_kernelsby Robert LaytonMini-Batch K-Meansby Nelle Varoquaux and Peter Prettenhofer.mldata utilities by Pietro Berkes.
API changes summary#
Here are the code migration instructions when upgrading from scikit-learn version 0.8:
The
scikits.learnpackage was renamedsklearn. There is still ascikits.learnpackage alias for backward compatibility.Third-party projects with a dependency on scikit-learn 0.9+ should upgrade their codebase. For instance, under Linux / MacOSX just run (make a backup first!):
find -name "*.py" | xargs sed -i 's/\bscikits.learn\b/sklearn/g'
Estimators no longer accept model parameters as
fitarguments: instead all parameters must be only be passed as constructor arguments or using the now publicset_paramsmethod inherited fromBaseEstimator.Some estimators can still accept keyword arguments on the
fitbut this is restricted to data-dependent values (e.g. a Gram matrix or an affinity matrix that are precomputed from theXdata matrix.The
cross_valpackage has been renamed tocross_validationalthough there is also across_valpackage alias in place for backward compatibility.Third-party projects with a dependency on scikit-learn 0.9+ should upgrade their codebase. For instance, under Linux / MacOSX just run (make a backup first!):
find -name "*.py" | xargs sed -i 's/\bcross_val\b/cross_validation/g'
The
score_funcargument of thesklearn.cross_validation.cross_val_scorefunction is now expected to accepty_testandy_predictedas only arguments for classification and regression tasks orX_testfor unsupervised estimators.gammaparameter for support vector machine algorithms is set to1 / n_featuresby default, instead of1 / n_samples.The
sklearn.hmmhas been marked as orphaned: it will be removed from scikit-learn in version 0.11 unless someone steps up to contribute documentation, examples and fix lurking numerical stability issues.sklearn.neighborshas been made into a submodule. The two previously available estimators,NeighborsClassifierandNeighborsRegressorhave been marked as deprecated. Their functionality has been divided among five new classes:NearestNeighborsfor unsupervised neighbors searches,KNeighborsClassifier&RadiusNeighborsClassifierfor supervised classification problems, andKNeighborsRegressor&RadiusNeighborsRegressorfor supervised regression problems.sklearn.ball_tree.BallTreehas been moved tosklearn.neighbors.BallTree. Using the former will generate a warning.sklearn.linear_model.LARS()and related classes (LassoLARS, LassoLARSCV, etc.) have been renamed tosklearn.linear_model.Lars().All distance metrics and kernels in
sklearn.metrics.pairwisenow have a Y parameter, which by default is None. If not given, the result is the distance (or kernel similarity) between each sample in Y. If given, the result is the pairwise distance (or kernel similarity) between samples in X to Y.sklearn.metrics.pairwise.l1_distanceis now calledmanhattan_distance, and by default returns the pairwise distance. For the component wise distance, set the parametersum_over_featurestoFalse.
Backward compatibility package aliases and other deprecated classes and functions will be removed in version 0.11.
People#
38 people contributed to this release.
387 Vlad Niculae
320 Olivier Grisel
192 Lars Buitinck
179 Gael Varoquaux
168 Fabian Pedregosa (INRIA, Parietal Team)
127 Jake Vanderplas
120 Mathieu Blondel
42 Robert Layton
38 Nelle Varoquaux
30 Conrad Lee
22 Pietro Berkes
18 andy
17 David Warde-Farley
12 Brian Holt
11 Robert
8 Amit Aides
6 Salvatore Masecchia
5 Paolo Losi
4 Vincent Schut
3 Alexis Metaireau
3 Bryan Silverthorn
2 Minwoo Jake Lee
1 Emmanuelle Gouillart
1 Keith Goodman
1 Lucas Wiman
1 Thouis (Ray) Jones
1 Tim Sheerman-Chase
Version 0.8#
May 11, 2011
scikit-learn 0.8 was released on May 2011, one month after the first “international” scikit-learn coding sprint and is marked by the inclusion of important modules: Hierarchical clustering, Cross decomposition, Non-negative matrix factorization (NMF or NNMF), initial support for Python 3 and by important enhancements and bug fixes.
Changelog#
Several new modules were introduced during this release:
New Hierarchical clustering module by Vincent Michel, Bertrand Thirion, Alexandre Gramfort and Gael Varoquaux.
Kernel Principal Component Analysis (kPCA) implementation by Mathieu Blondel
The Labeled Faces in the Wild face recognition dataset by Olivier Grisel.
New Cross decomposition module by Edouard Duchesnay.
Non-negative matrix factorization (NMF or NNMF) module Vlad Niculae
Implementation of the Oracle Approximating Shrinkage algorithm by Virgile Fritsch in the Covariance estimation module.
Some other modules benefited from significant improvements or cleanups.
Initial support for Python 3: builds and imports cleanly, some modules are usable while others have failing tests by Fabian Pedregosa.
PCAis now usable from the Pipeline object by Olivier Grisel.Guide How to optimize for speed by Olivier Grisel.
Fixes for memory leaks in libsvm bindings, 64-bit safer BallTree by Lars Buitinck.
bug and style fixing in K-means algorithm by Jan Schlüter.
Add attribute converged to Gaussian Mixture Models by Vincent Schut.
Implemented
transform,predict_log_probainLinearDiscriminantAnalysisBy Mathieu Blondel.Refactoring in the Support Vector Machines module and bug fixes by Fabian Pedregosa, Gael Varoquaux and Amit Aides.
Refactored SGD module (removed code duplication, better variable naming), added interface for sample weight by Peter Prettenhofer.
Wrapped BallTree with Cython by Thouis (Ray) Jones.
Added function
svm.l1_min_cby Paolo Losi.Typos, doc style, etc. by Yaroslav Halchenko, Gael Varoquaux, Olivier Grisel, Yann Malet, Nicolas Pinto, Lars Buitinck and Fabian Pedregosa.
People#
People that made this release possible preceded by number of commits:
159 Olivier Grisel
96 Vlad Niculae
32 Paolo Losi
7 Lars Buitinck
6 Vincent Michel
4 Thouis (Ray) Jones
4 Vincent Schut
3 Jan Schlüter
2 Julien Miotte
2 Yann Malet
1 Amit Aides
1 Feth Arezki
1 Meng Xinfan
Version 0.7#
March 2, 2011
scikit-learn 0.7 was released in March 2011, roughly three months after the 0.6 release. This release is marked by the speed improvements in existing algorithms like k-Nearest Neighbors and K-Means algorithm and by the inclusion of an efficient algorithm for computing the Ridge Generalized Cross Validation solution. Unlike the preceding release, no new modules were added to this release.
Changelog#
Performance improvements for Gaussian Mixture Model sampling [Jan Schlüter].
Implementation of efficient leave-one-out cross-validated Ridge in
RidgeCV[Mathieu Blondel]Better handling of collinearity and early stopping in
linear_model.lars_path[Alexandre Gramfort and Fabian Pedregosa].Fixes for liblinear ordering of labels and sign of coefficients [Dan Yamins, Paolo Losi, Mathieu Blondel and Fabian Pedregosa].
Performance improvements for Nearest Neighbors algorithm in high-dimensional spaces [Fabian Pedregosa].
Performance improvements for
KMeans[Gael Varoquaux and James Bergstra].Sanity checks for SVM-based classes [Mathieu Blondel].
Refactoring of
neighbors.NeighborsClassifierandneighbors.kneighbors_graph: added different algorithms for the k-Nearest Neighbor Search and implemented a more stable algorithm for finding barycenter weights. Also added some developer documentation for this module, see notes_neighbors for more information [Fabian Pedregosa].Documentation improvements: Added
pca.RandomizedPCAandLogisticRegressionto the class reference. Also added references of matrices used for clustering and other fixes [Gael Varoquaux, Fabian Pedregosa, Mathieu Blondel, Olivier Grisel, Virgile Fritsch , Emmanuelle Gouillart]Binded decision_function in classes that make use of liblinear, dense and sparse variants, like
LinearSVCorLogisticRegression[Fabian Pedregosa].Performance and API improvements to
metrics.pairwise.euclidean_distancesand topca.RandomizedPCA[James Bergstra].Fix compilation issues under NetBSD [Kamel Ibn Hassen Derouiche]
Allow input sequences of different lengths in
hmm.GaussianHMM[Ron Weiss].Fix bug in affinity propagation caused by incorrect indexing [Xinfan Meng]
People#
People that made this release possible preceded by number of commits:
14 Dan Yamins
2 Satrajit Ghosh
2 Vincent Dubourg
1 Emmanuelle Gouillart
1 Kamel Ibn Hassen Derouiche
1 Paolo Losi
1 VirgileFritsch
1 Xinfan Meng
Version 0.6#
December 21, 2010
scikit-learn 0.6 was released on December 2010. It is marked by the inclusion of several new modules and a general renaming of old ones. It is also marked by the inclusion of new example, including applications to real-world datasets.
Changelog#
New stochastic gradient descent module by Peter Prettenhofer. The module comes with complete documentation and examples.
Improved svm module: memory consumption has been reduced by 50%, heuristic to automatically set class weights, possibility to assign weights to samples (see SVM: Weighted samples for an example).
New Gaussian Processes module by Vincent Dubourg. This module also has great documentation and some very neat examples. See example_gaussian_process_plot_gp_regression.py or example_gaussian_process_plot_gp_probabilistic_classification_after_regression.py for a taste of what can be done.
It is now possible to use liblinear’s Multi-class SVC (option multi_class in
LinearSVC)New features and performance improvements of text feature extraction.
Improved sparse matrix support, both in main classes (
GridSearchCV) as in modules sklearn.svm.sparse and sklearn.linear_model.sparse.Lots of cool new examples and a new section that uses real-world datasets was created. These include: Faces recognition example using eigenfaces and SVMs, Species distribution modeling, Wikipedia principal eigenvector and others.
Faster Least Angle Regression algorithm. It is now 2x faster than the R version on worst case and up to 10x times faster on some cases.
Faster coordinate descent algorithm. In particular, the full path version of lasso (
linear_model.lasso_path) is more than 200x times faster than before.It is now possible to get probability estimates from a
LogisticRegressionmodel.module renaming: the glm module has been renamed to linear_model, the gmm module has been included into the more general mixture model and the sgd module has been included in linear_model.
Lots of bug fixes and documentation improvements.
People#
People that made this release possible preceded by number of commits:
207 Olivier Grisel
167 Fabian Pedregosa
33 Vincent Dubourg
21 Ron Weiss
9 Bertrand Thirion
3 Anne-Laure Fouque
2 Ronan Amicel
Version 0.5#
October 11, 2010
Changelog#
New classes#
Support for sparse matrices in some classifiers of modules
svmandlinear_model(seesvm.sparse.SVC,svm.sparse.SVR,svm.sparse.LinearSVC,linear_model.sparse.Lasso,linear_model.sparse.ElasticNet)New
Pipelineobject to compose different estimators.Recursive Feature Elimination routines in module Feature selection.
Addition of various classes capable of cross validation in the linear_model module (
LassoCV,ElasticNetCV, etc.).New, more efficient LARS algorithm implementation. The Lasso variant of the algorithm is also implemented. See
lars_path,LarsandLassoLars.New Hidden Markov Models module (see classes
hmm.GaussianHMM,hmm.MultinomialHMM,hmm.GMMHMM)New module feature_extraction (see class reference)
New FastICA algorithm in module sklearn.fastica
Documentation#
Improved documentation for many modules, now separating narrative documentation from the class reference. As an example, see documentation for the SVM module and the complete class reference.
Fixes#
API changes: adhere variable names to PEP-8, give more meaningful names.
Fixes for svm module to run on a shared memory context (multiprocessing).
It is again possible to generate latex (and thus PDF) from the sphinx docs.
Examples#
new examples using some of the mlcomp datasets:
sphx_glr_auto_examples_mlcomp_sparse_document_classification.py(since removed) and Classification of text documents using sparse featuresMany more examples. See here the full list of examples.
External dependencies#
Joblib is now a dependency of this package, although it is shipped with (sklearn.externals.joblib).
Removed modules#
Module ann (Artificial Neural Networks) has been removed from the distribution. Users wanting this sort of algorithms should take a look into pybrain.
Misc#
New sphinx theme for the web page.
Version 0.4#
August 26, 2010
Changelog#
Major changes in this release include:
Coordinate Descent algorithm (Lasso, ElasticNet) refactoring & speed improvements (roughly 100x times faster).
Coordinate Descent Refactoring (and bug fixing) for consistency with R’s package GLMNET.
New metrics module.
New GMM module contributed by Ron Weiss.
Implementation of the LARS algorithm (without Lasso variant for now).
feature_selection module redesign.
Migration to GIT as version control system.
Removal of obsolete attrselect module.
Rename of private compiled extensions (added underscore).
Removal of legacy unmaintained code.
Documentation improvements (both docstring and rst).
Improvement of the build system to (optionally) link with MKL. Also, provide a lite BLAS implementation in case no system-wide BLAS is found.
Lots of new examples.
Many, many bug fixes …
Authors#
The committer list for this release is the following (preceded by number of commits):
143 Fabian Pedregosa
35 Alexandre Gramfort
34 Olivier Grisel
11 Gael Varoquaux
5 Yaroslav Halchenko
2 Vincent Michel
1 Chris Filo Gorgolewski
Earlier versions#
Earlier versions included contributions by Fred Mailhot, David Cooke, David Huard, Dave Morrill, Ed Schofield, Travis Oliphant, Pearu Peterson.