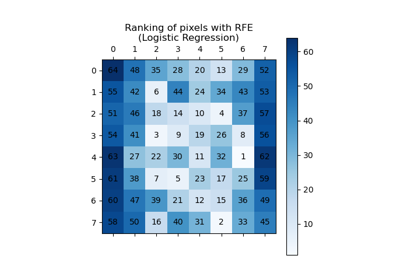
RFE#
- class sklearn.feature_selection.RFE(estimator, *, n_features_to_select=None, step=1, verbose=0, importance_getter='auto')[source]#
Feature ranking with recursive feature elimination.
Given an external estimator that assigns weights to features (e.g., the coefficients of a linear model), the goal of recursive feature elimination (RFE) is to select features by recursively considering smaller and smaller sets of features. First, the estimator is trained on the initial set of features and the importance of each feature is obtained either through any specific attribute or callable. Then, the least important features are pruned from current set of features. That procedure is recursively repeated on the pruned set until the desired number of features to select is eventually reached.
Read more in the User Guide.
- Parameters:
- estimator
Estimatorinstance A supervised learning estimator with a
fitmethod that provides information about feature importance (e.g.coef_,feature_importances_).- n_features_to_selectint or float, default=None
The number of features to select. If
None, half of the features are selected. If integer, the parameter is the absolute number of features to select. If float between 0 and 1, it is the fraction of features to select.Changed in version 0.24: Added float values for fractions.
- stepint or float, default=1
If greater than or equal to 1, then
stepcorresponds to the (integer) number of features to remove at each iteration. If within (0.0, 1.0), thenstepcorresponds to the percentage (rounded down) of features to remove at each iteration.- verboseint, default=0
Controls verbosity of output.
- importance_getterstr or callable, default=’auto’
If ‘auto’, uses the feature importance either through a
coef_orfeature_importances_attributes of estimator.Also accepts a string that specifies an attribute name/path for extracting feature importance (implemented with
attrgetter). For example, giveregressor_.coef_in case ofTransformedTargetRegressorornamed_steps.clf.feature_importances_in case of class:~sklearn.pipeline.Pipelinewith its last step namedclf.If
callable, overrides the default feature importance getter. The callable is passed with the fitted estimator and it should return importance for each feature.Added in version 0.24.
- estimator
- Attributes:
classes_ndarray of shape (n_classes,)Classes labels available when
estimatoris a classifier.- estimator_
Estimatorinstance The fitted estimator used to select features.
- n_features_int
The number of selected features.
- n_features_in_int
Number of features seen during fit. Only defined if the underlying estimator exposes such an attribute when fit.
Added in version 0.24.
- feature_names_in_ndarray of shape (
n_features_in_,) Names of features seen during fit. Defined only when
Xhas feature names that are all strings.Added in version 1.0.
- ranking_ndarray of shape (n_features,)
The feature ranking, such that
ranking_[i]corresponds to the ranking position of the i-th feature. Selected (i.e., estimated best) features are assigned rank 1.- support_ndarray of shape (n_features,)
The mask of selected features.
See also
RFECVRecursive feature elimination with built-in cross-validated selection of the best number of features.
SelectFromModelFeature selection based on thresholds of importance weights.
SequentialFeatureSelectorSequential cross-validation based feature selection. Does not rely on importance weights.
Notes
Allows NaN/Inf in the input if the underlying estimator does as well.
References
[1]Guyon, I., Weston, J., Barnhill, S., & Vapnik, V., “Gene selection for cancer classification using support vector machines”, Mach. Learn., 46(1-3), 389–422, 2002.
Examples
The following example shows how to retrieve the 5 most informative features in the Friedman #1 dataset.
>>> from sklearn.datasets import make_friedman1 >>> from sklearn.feature_selection import RFE >>> from sklearn.svm import SVR >>> X, y = make_friedman1(n_samples=50, n_features=10, random_state=0) >>> estimator = SVR(kernel="linear") >>> selector = RFE(estimator, n_features_to_select=5, step=1) >>> selector = selector.fit(X, y) >>> selector.support_ array([ True, True, True, True, True, False, False, False, False, False]) >>> selector.ranking_ array([1, 1, 1, 1, 1, 6, 4, 3, 2, 5])
- property classes_#
Classes labels available when
estimatoris a classifier.- Returns:
- ndarray of shape (n_classes,)
- decision_function(X)[source]#
Compute the decision function of
X.- Parameters:
- X{array-like or sparse matrix} of shape (n_samples, n_features)
The input samples. Internally, it will be converted to
dtype=np.float32and if a sparse matrix is provided to a sparsecsr_matrix.
- Returns:
- scorearray, shape = [n_samples, n_classes] or [n_samples]
The decision function of the input samples. The order of the classes corresponds to that in the attribute classes_. Regression and binary classification produce an array of shape [n_samples].
- fit(X, y, **fit_params)[source]#
Fit the RFE model and then the underlying estimator on the selected features.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples.
- yarray-like of shape (n_samples,)
The target values.
- **fit_paramsdict
If
enable_metadata_routing=False(default): Parameters directly passed to thefitmethod of the underlying estimator.If
enable_metadata_routing=True: Parameters safely routed to thefitmethod of the underlying estimator.
Changed in version 1.6: See Metadata Routing User Guide for more details.
- Returns:
- selfobject
Fitted estimator.
- fit_transform(X, y=None, **fit_params)[source]#
Fit to data, then transform it.
Fits transformer to
Xandywith optional parametersfit_paramsand returns a transformed version ofX.- Parameters:
- Xarray-like of shape (n_samples, n_features)
Input samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs), default=None
Target values (None for unsupervised transformations).
- **fit_paramsdict
Additional fit parameters.
- Returns:
- X_newndarray array of shape (n_samples, n_features_new)
Transformed array.
- get_feature_names_out(input_features=None)[source]#
Mask feature names according to selected features.
- Parameters:
- input_featuresarray-like of str or None, default=None
Input features.
If
input_featuresisNone, thenfeature_names_in_is used as feature names in. Iffeature_names_in_is not defined, then the following input feature names are generated:["x0", "x1", ..., "x(n_features_in_ - 1)"].If
input_featuresis an array-like, theninput_featuresmust matchfeature_names_in_iffeature_names_in_is defined.
- Returns:
- feature_names_outndarray of str objects
Transformed feature names.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
Added in version 1.6.
- Returns:
- routingMetadataRouter
A
MetadataRouterencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- get_support(indices=False)[source]#
Get a mask, or integer index, of the features selected.
- Parameters:
- indicesbool, default=False
If True, the return value will be an array of integers, rather than a boolean mask.
- Returns:
- supportarray
An index that selects the retained features from a feature vector. If
indicesis False, this is a boolean array of shape [# input features], in which an element is True iff its corresponding feature is selected for retention. Ifindicesis True, this is an integer array of shape [# output features] whose values are indices into the input feature vector.
- inverse_transform(X)[source]#
Reverse the transformation operation.
- Parameters:
- Xarray of shape [n_samples, n_selected_features]
The input samples.
- Returns:
- X_rarray of shape [n_samples, n_original_features]
Xwith columns of zeros inserted where features would have been removed bytransform.
- predict(X, **predict_params)[source]#
Reduce X to the selected features and predict using the estimator.
- Parameters:
- Xarray of shape [n_samples, n_features]
The input samples.
- **predict_paramsdict
Parameters to route to the
predictmethod of the underlying estimator.Added in version 1.6: Only available if
enable_metadata_routing=True, which can be set by usingsklearn.set_config(enable_metadata_routing=True). See Metadata Routing User Guide for more details.
- Returns:
- yarray of shape [n_samples]
The predicted target values.
- predict_log_proba(X)[source]#
Predict class log-probabilities for X.
- Parameters:
- Xarray of shape [n_samples, n_features]
The input samples.
- Returns:
- parray of shape (n_samples, n_classes)
The class log-probabilities of the input samples. The order of the classes corresponds to that in the attribute classes_.
- predict_proba(X)[source]#
Predict class probabilities for X.
- Parameters:
- X{array-like or sparse matrix} of shape (n_samples, n_features)
The input samples. Internally, it will be converted to
dtype=np.float32and if a sparse matrix is provided to a sparsecsr_matrix.
- Returns:
- parray of shape (n_samples, n_classes)
The class probabilities of the input samples. The order of the classes corresponds to that in the attribute classes_.
- score(X, y, **score_params)[source]#
Reduce X to the selected features and return the score of the estimator.
- Parameters:
- Xarray of shape [n_samples, n_features]
The input samples.
- yarray of shape [n_samples]
The target values.
- **score_paramsdict
If
enable_metadata_routing=False(default): Parameters directly passed to thescoremethod of the underlying estimator.If
enable_metadata_routing=True: Parameters safely routed to thescoremethod of the underlying estimator.
Added in version 1.0.
Changed in version 1.6: See Metadata Routing User Guide for more details.
- Returns:
- scorefloat
Score of the underlying base estimator computed with the selected features returned by
rfe.transform(X)andy.
- set_output(*, transform=None)[source]#
Set output container.
See Introducing the set_output API for an example on how to use the API.
- Parameters:
- transform{“default”, “pandas”, “polars”}, default=None
Configure output of
transformandfit_transform."default": Default output format of a transformer"pandas": DataFrame output"polars": Polars outputNone: Transform configuration is unchanged
Added in version 1.4:
"polars"option was added.
- Returns:
- selfestimator instance
Estimator instance.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.