RepeatedStratifiedKFold#
- class sklearn.model_selection.RepeatedStratifiedKFold(*, n_splits=5, n_repeats=10, random_state=None)[source]#
Repeated Stratified K-Fold cross validator.
Repeats Stratified K-Fold n times with different randomization in each repetition.
Read more in the User Guide.
- Parameters:
- n_splitsint, default=5
Number of folds. Must be at least 2.
- n_repeatsint, default=10
Number of times cross-validator needs to be repeated.
- random_stateint, RandomState instance or None, default=None
Controls the generation of the random states for each repetition. Pass an int for reproducible output across multiple function calls. See Glossary.
See also
RepeatedKFoldRepeats K-Fold n times.
Notes
Randomized CV splitters may return different results for each call of split. You can make the results identical by setting
random_stateto an integer.Examples
>>> import numpy as np >>> from sklearn.model_selection import RepeatedStratifiedKFold >>> X = np.array([[1, 2], [3, 4], [1, 2], [3, 4]]) >>> y = np.array([0, 0, 1, 1]) >>> rskf = RepeatedStratifiedKFold(n_splits=2, n_repeats=2, ... random_state=36851234) >>> rskf.get_n_splits(X, y) 4 >>> print(rskf) RepeatedStratifiedKFold(n_repeats=2, n_splits=2, random_state=36851234) >>> for i, (train_index, test_index) in enumerate(rskf.split(X, y)): ... print(f"Fold {i}:") ... print(f" Train: index={train_index}") ... print(f" Test: index={test_index}") ... Fold 0: Train: index=[1 2] Test: index=[0 3] Fold 1: Train: index=[0 3] Test: index=[1 2] Fold 2: Train: index=[1 3] Test: index=[0 2] Fold 3: Train: index=[0 2] Test: index=[1 3]
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_n_splits(X=None, y=None, groups=None)[source]#
Returns the number of splitting iterations in the cross-validator.
- Parameters:
- Xobject
Always ignored, exists for compatibility.
np.zeros(n_samples)may be used as a placeholder.- yobject
Always ignored, exists for compatibility.
np.zeros(n_samples)may be used as a placeholder.- groupsarray-like of shape (n_samples,), default=None
Group labels for the samples used while splitting the dataset into train/test set.
- Returns:
- n_splitsint
Returns the number of splitting iterations in the cross-validator.
- split(X, y, groups=None)[source]#
Generate indices to split data into training and test set.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Training data, where
n_samplesis the number of samples andn_featuresis the number of features.Note that providing
yis sufficient to generate the splits and hencenp.zeros(n_samples)may be used as a placeholder forXinstead of actual training data.- yarray-like of shape (n_samples,)
The target variable for supervised learning problems. Stratification is done based on the y labels.
- groupsobject
Always ignored, exists for compatibility.
- Yields:
- trainndarray
The training set indices for that split.
- testndarray
The testing set indices for that split.
Notes
Randomized CV splitters may return different results for each call of split. You can make the results identical by setting
random_stateto an integer.
Gallery examples#
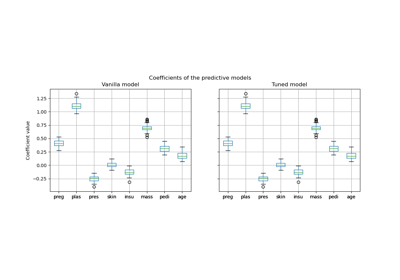
Post-hoc tuning the cut-off point of decision function
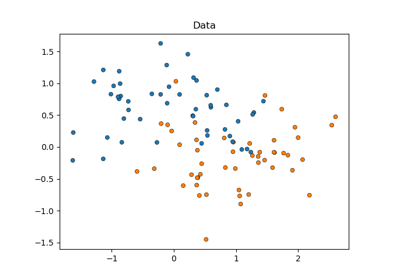
Statistical comparison of models using grid search