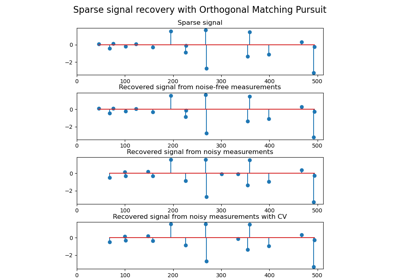
3.2.4.1.8. sklearn.linear_model.OrthogonalMatchingPursuitCV¶
-
class
sklearn.linear_model.OrthogonalMatchingPursuitCV(*, copy=True, fit_intercept=True, normalize=True, max_iter=None, cv=None, n_jobs=None, verbose=False)[source]¶ Cross-validated Orthogonal Matching Pursuit model (OMP).
See glossary entry for cross-validation estimator.
Read more in the User Guide.
- Parameters
- copybool, optional
Whether the design matrix X must be copied by the algorithm. A false value is only helpful if X is already Fortran-ordered, otherwise a copy is made anyway.
- fit_interceptboolean, optional
whether to calculate the intercept for this model. If set to false, no intercept will be used in calculations (i.e. data is expected to be centered).
- normalizeboolean, optional, default True
This parameter is ignored when
fit_interceptis set to False. If True, the regressors X will be normalized before regression by subtracting the mean and dividing by the l2-norm. If you wish to standardize, please usesklearn.preprocessing.StandardScalerbefore callingfiton an estimator withnormalize=False.- max_iterinteger, optional
Maximum numbers of iterations to perform, therefore maximum features to include. 10% of
n_featuresbut at least 5 if available.- cvint, cross-validation generator or an iterable, optional
Determines the cross-validation splitting strategy. Possible inputs for cv are:
None, to use the default 5-fold cross-validation,
integer, to specify the number of folds.
An iterable yielding (train, test) splits as arrays of indices.
For integer/None inputs,
KFoldis used.Refer User Guide for the various cross-validation strategies that can be used here.
Changed in version 0.22:
cvdefault value if None changed from 3-fold to 5-fold.- n_jobsint or None, optional (default=None)
Number of CPUs to use during the cross validation.
Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.- verboseboolean or integer, optional
Sets the verbosity amount
- Attributes
- intercept_float or array, shape (n_targets,)
Independent term in decision function.
- coef_array, shape (n_features,) or (n_targets, n_features)
Parameter vector (w in the problem formulation).
- n_nonzero_coefs_int
Estimated number of non-zero coefficients giving the best mean squared error over the cross-validation folds.
- n_iter_int or array-like
Number of active features across every target for the model refit with the best hyperparameters got by cross-validating across all folds.
See also
orthogonal_mporthogonal_mp_gramlars_pathLarsLassoLarsOrthogonalMatchingPursuitLarsCVLassoLarsCVdecomposition.sparse_encode
Examples
>>> from sklearn.linear_model import OrthogonalMatchingPursuitCV >>> from sklearn.datasets import make_regression >>> X, y = make_regression(n_features=100, n_informative=10, ... noise=4, random_state=0) >>> reg = OrthogonalMatchingPursuitCV(cv=5).fit(X, y) >>> reg.score(X, y) 0.9991... >>> reg.n_nonzero_coefs_ 10 >>> reg.predict(X[:1,]) array([-78.3854...])
Methods
fit(X, y)Fit the model using X, y as training data.
get_params([deep])Get parameters for this estimator.
predict(X)Predict using the linear model.
score(X, y[, sample_weight])Return the coefficient of determination R^2 of the prediction.
set_params(**params)Set the parameters of this estimator.
-
__init__(*, copy=True, fit_intercept=True, normalize=True, max_iter=None, cv=None, n_jobs=None, verbose=False)[source]¶ Initialize self. See help(type(self)) for accurate signature.
-
fit(X, y)[source]¶ Fit the model using X, y as training data.
- Parameters
- Xarray-like, shape [n_samples, n_features]
Training data.
- yarray-like, shape [n_samples]
Target values. Will be cast to X’s dtype if necessary
- Returns
- selfobject
returns an instance of self.
-
get_params(deep=True)[source]¶ Get parameters for this estimator.
- Parameters
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns
- paramsmapping of string to any
Parameter names mapped to their values.
-
predict(X)[source]¶ Predict using the linear model.
- Parameters
- Xarray_like or sparse matrix, shape (n_samples, n_features)
Samples.
- Returns
- Carray, shape (n_samples,)
Returns predicted values.
-
score(X, y, sample_weight=None)[source]¶ Return the coefficient of determination R^2 of the prediction.
The coefficient R^2 is defined as (1 - u/v), where u is the residual sum of squares ((y_true - y_pred) ** 2).sum() and v is the total sum of squares ((y_true - y_true.mean()) ** 2).sum(). The best possible score is 1.0 and it can be negative (because the model can be arbitrarily worse). A constant model that always predicts the expected value of y, disregarding the input features, would get a R^2 score of 0.0.
- Parameters
- Xarray-like of shape (n_samples, n_features)
Test samples. For some estimators this may be a precomputed kernel matrix or a list of generic objects instead, shape = (n_samples, n_samples_fitted), where n_samples_fitted is the number of samples used in the fitting for the estimator.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True values for X.
- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns
- scorefloat
R^2 of self.predict(X) wrt. y.
Notes
The R2 score used when calling
scoreon a regressor usesmultioutput='uniform_average'from version 0.23 to keep consistent with default value ofr2_score. This influences thescoremethod of all the multioutput regressors (except forMultiOutputRegressor).
-
set_params(**params)[source]¶ Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as pipelines). The latter have parameters of the form
<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters
- **paramsdict
Estimator parameters.
- Returns
- selfobject
Estimator instance.