Note
Go to the end to download the full example code or to run this example in your browser via JupyterLite or Binder
Feature agglomeration vs. univariate selection¶
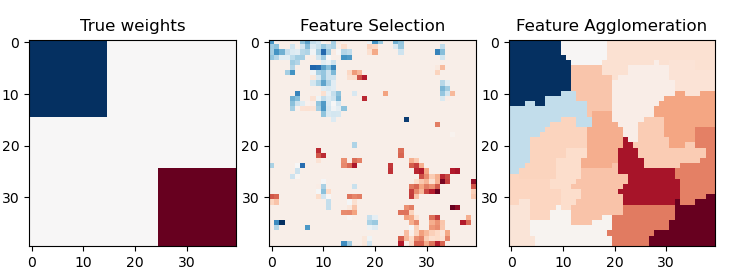
This example compares 2 dimensionality reduction strategies:
univariate feature selection with Anova
feature agglomeration with Ward hierarchical clustering
Both methods are compared in a regression problem using a BayesianRidge as supervised estimator.
# Author: Alexandre Gramfort <alexandre.gramfort@inria.fr>
# License: BSD 3 clause
import shutil
import tempfile
import matplotlib.pyplot as plt
import numpy as np
from joblib import Memory
from scipy import linalg, ndimage
from sklearn import feature_selection
from sklearn.cluster import FeatureAgglomeration
from sklearn.feature_extraction.image import grid_to_graph
from sklearn.linear_model import BayesianRidge
from sklearn.model_selection import GridSearchCV, KFold
from sklearn.pipeline import Pipeline
Set parameters
n_samples = 200
size = 40 # image size
roi_size = 15
snr = 5.0
np.random.seed(0)
Generate data
coef = np.zeros((size, size))
coef[0:roi_size, 0:roi_size] = -1.0
coef[-roi_size:, -roi_size:] = 1.0
X = np.random.randn(n_samples, size**2)
for x in X: # smooth data
x[:] = ndimage.gaussian_filter(x.reshape(size, size), sigma=1.0).ravel()
X -= X.mean(axis=0)
X /= X.std(axis=0)
y = np.dot(X, coef.ravel())
add noise
noise = np.random.randn(y.shape[0])
noise_coef = (linalg.norm(y, 2) / np.exp(snr / 20.0)) / linalg.norm(noise, 2)
y += noise_coef * noise
Compute the coefs of a Bayesian Ridge with GridSearch
cv = KFold(2) # cross-validation generator for model selection
ridge = BayesianRidge()
cachedir = tempfile.mkdtemp()
mem = Memory(location=cachedir, verbose=1)
Ward agglomeration followed by BayesianRidge
connectivity = grid_to_graph(n_x=size, n_y=size)
ward = FeatureAgglomeration(n_clusters=10, connectivity=connectivity, memory=mem)
clf = Pipeline([("ward", ward), ("ridge", ridge)])
# Select the optimal number of parcels with grid search
clf = GridSearchCV(clf, {"ward__n_clusters": [10, 20, 30]}, n_jobs=1, cv=cv)
clf.fit(X, y) # set the best parameters
coef_ = clf.best_estimator_.steps[-1][1].coef_
coef_ = clf.best_estimator_.steps[0][1].inverse_transform(coef_)
coef_agglomeration_ = coef_.reshape(size, size)
________________________________________________________________________________
[Memory] Calling sklearn.cluster._agglomerative.ward_tree...
ward_tree(array([[-0.451933, ..., -0.675318],
...,
[ 0.275706, ..., -1.085711]]), connectivity=<1600x1600 sparse matrix of type '<class 'numpy.int64'>'
with 7840 stored elements in COOrdinate format>, n_clusters=None, return_distance=False)
________________________________________________________ward_tree - 0.0s, 0.0min
________________________________________________________________________________
[Memory] Calling sklearn.cluster._agglomerative.ward_tree...
ward_tree(array([[ 0.905206, ..., 0.161245],
...,
[-0.849835, ..., -1.091621]]), connectivity=<1600x1600 sparse matrix of type '<class 'numpy.int64'>'
with 7840 stored elements in COOrdinate format>, n_clusters=None, return_distance=False)
________________________________________________________ward_tree - 0.0s, 0.0min
________________________________________________________________________________
[Memory] Calling sklearn.cluster._agglomerative.ward_tree...
ward_tree(array([[ 0.905206, ..., -0.675318],
...,
[-0.849835, ..., -1.085711]]), connectivity=<1600x1600 sparse matrix of type '<class 'numpy.int64'>'
with 7840 stored elements in COOrdinate format>, n_clusters=None, return_distance=False)
________________________________________________________ward_tree - 0.0s, 0.0min
Anova univariate feature selection followed by BayesianRidge
f_regression = mem.cache(feature_selection.f_regression) # caching function
anova = feature_selection.SelectPercentile(f_regression)
clf = Pipeline([("anova", anova), ("ridge", ridge)])
# Select the optimal percentage of features with grid search
clf = GridSearchCV(clf, {"anova__percentile": [5, 10, 20]}, cv=cv)
clf.fit(X, y) # set the best parameters
coef_ = clf.best_estimator_.steps[-1][1].coef_
coef_ = clf.best_estimator_.steps[0][1].inverse_transform(coef_.reshape(1, -1))
coef_selection_ = coef_.reshape(size, size)
________________________________________________________________________________
[Memory] Calling sklearn.feature_selection._univariate_selection.f_regression...
f_regression(array([[-0.451933, ..., 0.275706],
...,
[-0.675318, ..., -1.085711]]),
array([ 25.267703, ..., -25.026711]))
_____________________________________________________f_regression - 0.0s, 0.0min
________________________________________________________________________________
[Memory] Calling sklearn.feature_selection._univariate_selection.f_regression...
f_regression(array([[ 0.905206, ..., -0.849835],
...,
[ 0.161245, ..., -1.091621]]),
array([ -27.447268, ..., -112.638768]))
_____________________________________________________f_regression - 0.0s, 0.0min
________________________________________________________________________________
[Memory] Calling sklearn.feature_selection._univariate_selection.f_regression...
f_regression(array([[ 0.905206, ..., -0.849835],
...,
[-0.675318, ..., -1.085711]]),
array([-27.447268, ..., -25.026711]))
_____________________________________________________f_regression - 0.0s, 0.0min
Inverse the transformation to plot the results on an image
plt.close("all")
plt.figure(figsize=(7.3, 2.7))
plt.subplot(1, 3, 1)
plt.imshow(coef, interpolation="nearest", cmap=plt.cm.RdBu_r)
plt.title("True weights")
plt.subplot(1, 3, 2)
plt.imshow(coef_selection_, interpolation="nearest", cmap=plt.cm.RdBu_r)
plt.title("Feature Selection")
plt.subplot(1, 3, 3)
plt.imshow(coef_agglomeration_, interpolation="nearest", cmap=plt.cm.RdBu_r)
plt.title("Feature Agglomeration")
plt.subplots_adjust(0.04, 0.0, 0.98, 0.94, 0.16, 0.26)
plt.show()
Attempt to remove the temporary cachedir, but don’t worry if it fails
shutil.rmtree(cachedir, ignore_errors=True)
Total running time of the script: (0 minutes 0.499 seconds)
Related examples

A demo of structured Ward hierarchical clustering on an image of coins
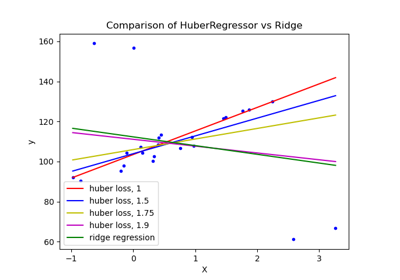
HuberRegressor vs Ridge on dataset with strong outliers