sklearn.preprocessing.PowerTransformer¶
-
class
sklearn.preprocessing.PowerTransformer(method='yeo-johnson', standardize=True, copy=True)[source]¶ Apply a power transform featurewise to make data more Gaussian-like.
Power transforms are a family of parametric, monotonic transformations that are applied to make data more Gaussian-like. This is useful for modeling issues related to heteroscedasticity (non-constant variance), or other situations where normality is desired.
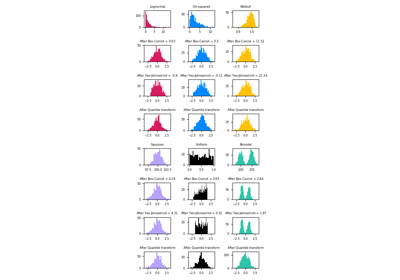
Currently, PowerTransformer supports the Box-Cox transform and the Yeo-Johnson transform. The optimal parameter for stabilizing variance and minimizing skewness is estimated through maximum likelihood.
Box-Cox requires input data to be strictly positive, while Yeo-Johnson supports both positive or negative data.
By default, zero-mean, unit-variance normalization is applied to the transformed data.
Read more in the User Guide.
New in version 0.20.
- Parameters
- methodstr, (default=’yeo-johnson’)
The power transform method. Available methods are:
‘yeo-johnson’ [Rf3e1504535de-1], works with positive and negative values
‘box-cox’ [Rf3e1504535de-2], only works with strictly positive values
- standardizeboolean, default=True
Set to True to apply zero-mean, unit-variance normalization to the transformed output.
- copyboolean, optional, default=True
Set to False to perform inplace computation during transformation.
- Attributes
- lambdas_array of float, shape (n_features,)
The parameters of the power transformation for the selected features.
See also
power_transformEquivalent function without the estimator API.
QuantileTransformerMaps data to a standard normal distribution with the parameter
output_distribution='normal'.
Notes
NaNs are treated as missing values: disregarded in
fit, and maintained intransform.For a comparison of the different scalers, transformers, and normalizers, see examples/preprocessing/plot_all_scaling.py.
References
- Rf3e1504535de-1
I.K. Yeo and R.A. Johnson, “A new family of power transformations to improve normality or symmetry.” Biometrika, 87(4), pp.954-959, (2000).
- Rf3e1504535de-2
G.E.P. Box and D.R. Cox, “An Analysis of Transformations”, Journal of the Royal Statistical Society B, 26, 211-252 (1964).
Examples
>>> import numpy as np >>> from sklearn.preprocessing import PowerTransformer >>> pt = PowerTransformer() >>> data = [[1, 2], [3, 2], [4, 5]] >>> print(pt.fit(data)) PowerTransformer() >>> print(pt.lambdas_) [ 1.386... -3.100...] >>> print(pt.transform(data)) [[-1.316... -0.707...] [ 0.209... -0.707...] [ 1.106... 1.414...]]
Methods
fit(self, X[, y])Estimate the optimal parameter lambda for each feature.
fit_transform(self, X[, y])Fit to data, then transform it.
get_params(self[, deep])Get parameters for this estimator.
inverse_transform(self, X)Apply the inverse power transformation using the fitted lambdas.
set_params(self, \*\*params)Set the parameters of this estimator.
transform(self, X)Apply the power transform to each feature using the fitted lambdas.
-
__init__(self, method='yeo-johnson', standardize=True, copy=True)[source]¶ Initialize self. See help(type(self)) for accurate signature.
-
fit(self, X, y=None)[source]¶ Estimate the optimal parameter lambda for each feature.
The optimal lambda parameter for minimizing skewness is estimated on each feature independently using maximum likelihood.
- Parameters
- Xarray-like, shape (n_samples, n_features)
The data used to estimate the optimal transformation parameters.
- yIgnored
- Returns
- selfobject
-
fit_transform(self, X, y=None)[source]¶ Fit to data, then transform it.
Fits transformer to X and y with optional parameters fit_params and returns a transformed version of X.
- Parameters
- Xnumpy array of shape [n_samples, n_features]
Training set.
- ynumpy array of shape [n_samples]
Target values.
- **fit_paramsdict
Additional fit parameters.
- Returns
- X_newnumpy array of shape [n_samples, n_features_new]
Transformed array.
-
get_params(self, deep=True)[source]¶ Get parameters for this estimator.
- Parameters
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns
- paramsmapping of string to any
Parameter names mapped to their values.
-
inverse_transform(self, X)[source]¶ Apply the inverse power transformation using the fitted lambdas.
The inverse of the Box-Cox transformation is given by:
if lambda_ == 0: X = exp(X_trans) else: X = (X_trans * lambda_ + 1) ** (1 / lambda_)
The inverse of the Yeo-Johnson transformation is given by:
if X >= 0 and lambda_ == 0: X = exp(X_trans) - 1 elif X >= 0 and lambda_ != 0: X = (X_trans * lambda_ + 1) ** (1 / lambda_) - 1 elif X < 0 and lambda_ != 2: X = 1 - (-(2 - lambda_) * X_trans + 1) ** (1 / (2 - lambda_)) elif X < 0 and lambda_ == 2: X = 1 - exp(-X_trans)
- Parameters
- Xarray-like, shape (n_samples, n_features)
The transformed data.
- Returns
- Xarray-like, shape (n_samples, n_features)
The original data
-
set_params(self, **params)[source]¶ Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as pipelines). The latter have parameters of the form
<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters
- **paramsdict
Estimator parameters.
- Returns
- selfobject
Estimator instance.