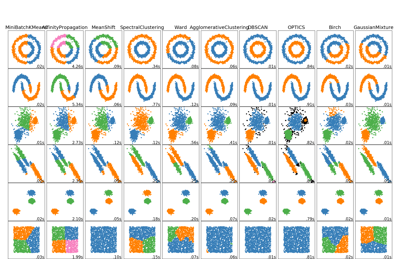
sklearn.cluster.SpectralClustering¶
-
class
sklearn.cluster.SpectralClustering(n_clusters=8, eigen_solver=None, n_components=None, random_state=None, n_init=10, gamma=1.0, affinity='rbf', n_neighbors=10, eigen_tol=0.0, assign_labels='kmeans', degree=3, coef0=1, kernel_params=None, n_jobs=None)[source]¶ Apply clustering to a projection of the normalized Laplacian.
In practice Spectral Clustering is very useful when the structure of the individual clusters is highly non-convex or more generally when a measure of the center and spread of the cluster is not a suitable description of the complete cluster. For instance when clusters are nested circles on the 2D plane.
If affinity is the adjacency matrix of a graph, this method can be used to find normalized graph cuts.
When calling
fit, an affinity matrix is constructed using either kernel function such the Gaussian (aka RBF) kernel of the euclidean distancedd(X, X):np.exp(-gamma * d(X,X) ** 2)
or a k-nearest neighbors connectivity matrix.
Alternatively, using
precomputed, a user-provided affinity matrix can be used.Read more in the User Guide.
- Parameters
- n_clustersinteger, optional
The dimension of the projection subspace.
- eigen_solver{None, ‘arpack’, ‘lobpcg’, or ‘amg’}
The eigenvalue decomposition strategy to use. AMG requires pyamg to be installed. It can be faster on very large, sparse problems, but may also lead to instabilities.
- n_componentsinteger, optional, default=n_clusters
Number of eigen vectors to use for the spectral embedding
- random_stateint, RandomState instance or None (default)
A pseudo random number generator used for the initialization of the lobpcg eigen vectors decomposition when
eigen_solver='amg'and by the K-Means initialization. Use an int to make the randomness deterministic. See Glossary.- n_initint, optional, default: 10
Number of time the k-means algorithm will be run with different centroid seeds. The final results will be the best output of n_init consecutive runs in terms of inertia.
- gammafloat, default=1.0
Kernel coefficient for rbf, poly, sigmoid, laplacian and chi2 kernels. Ignored for
affinity='nearest_neighbors'.- affinitystring or callable, default ‘rbf’
- How to construct the affinity matrix.
‘nearest_neighbors’ : construct the affinity matrix by computing a graph of nearest neighbors.
‘rbf’ : construct the affinity matrix using a radial basis function (RBF) kernel.
‘precomputed’ : interpret
Xas a precomputed affinity matrix.‘precomputed_nearest_neighbors’ : interpret
Xas a sparse graph of precomputed nearest neighbors, and constructs the affinity matrix by selecting then_neighborsnearest neighbors.one of the kernels supported by
pairwise_kernels.
Only kernels that produce similarity scores (non-negative values that increase with similarity) should be used. This property is not checked by the clustering algorithm.
- n_neighborsinteger
Number of neighbors to use when constructing the affinity matrix using the nearest neighbors method. Ignored for
affinity='rbf'.- eigen_tolfloat, optional, default: 0.0
Stopping criterion for eigendecomposition of the Laplacian matrix when
eigen_solver='arpack'.- assign_labels{‘kmeans’, ‘discretize’}, default: ‘kmeans’
The strategy to use to assign labels in the embedding space. There are two ways to assign labels after the laplacian embedding. k-means can be applied and is a popular choice. But it can also be sensitive to initialization. Discretization is another approach which is less sensitive to random initialization.
- degreefloat, default=3
Degree of the polynomial kernel. Ignored by other kernels.
- coef0float, default=1
Zero coefficient for polynomial and sigmoid kernels. Ignored by other kernels.
- kernel_paramsdictionary of string to any, optional
Parameters (keyword arguments) and values for kernel passed as callable object. Ignored by other kernels.
- n_jobsint or None, optional (default=None)
The number of parallel jobs to run.
Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.
- Attributes
- affinity_matrix_array-like, shape (n_samples, n_samples)
Affinity matrix used for clustering. Available only if after calling
fit.- labels_array, shape (n_samples,)
Labels of each point
Notes
If you have an affinity matrix, such as a distance matrix, for which 0 means identical elements, and high values means very dissimilar elements, it can be transformed in a similarity matrix that is well suited for the algorithm by applying the Gaussian (RBF, heat) kernel:
np.exp(- dist_matrix ** 2 / (2. * delta ** 2))
Where
deltais a free parameter representing the width of the Gaussian kernel.Another alternative is to take a symmetric version of the k nearest neighbors connectivity matrix of the points.
If the pyamg package is installed, it is used: this greatly speeds up computation.
References
Normalized cuts and image segmentation, 2000 Jianbo Shi, Jitendra Malik http://citeseer.ist.psu.edu/viewdoc/summary?doi=10.1.1.160.2324
A Tutorial on Spectral Clustering, 2007 Ulrike von Luxburg http://citeseerx.ist.psu.edu/viewdoc/summary?doi=10.1.1.165.9323
Multiclass spectral clustering, 2003 Stella X. Yu, Jianbo Shi https://www1.icsi.berkeley.edu/~stellayu/publication/doc/2003kwayICCV.pdf
Examples
>>> from sklearn.cluster import SpectralClustering >>> import numpy as np >>> X = np.array([[1, 1], [2, 1], [1, 0], ... [4, 7], [3, 5], [3, 6]]) >>> clustering = SpectralClustering(n_clusters=2, ... assign_labels="discretize", ... random_state=0).fit(X) >>> clustering.labels_ array([1, 1, 1, 0, 0, 0]) >>> clustering SpectralClustering(assign_labels='discretize', n_clusters=2, random_state=0)
Methods
fit(self, X[, y])Perform spectral clustering from features, or affinity matrix.
fit_predict(self, X[, y])Perform spectral clustering from features, or affinity matrix, and return cluster labels.
get_params(self[, deep])Get parameters for this estimator.
set_params(self, \*\*params)Set the parameters of this estimator.
-
__init__(self, n_clusters=8, eigen_solver=None, n_components=None, random_state=None, n_init=10, gamma=1.0, affinity='rbf', n_neighbors=10, eigen_tol=0.0, assign_labels='kmeans', degree=3, coef0=1, kernel_params=None, n_jobs=None)[source]¶ Initialize self. See help(type(self)) for accurate signature.
-
fit(self, X, y=None)[source]¶ Perform spectral clustering from features, or affinity matrix.
- Parameters
- Xarray-like or sparse matrix, shape (n_samples, n_features), or array-like, shape (n_samples, n_samples)
Training instances to cluster, or similarities / affinities between instances if
affinity='precomputed'. If a sparse matrix is provided in a format other thancsr_matrix,csc_matrix, orcoo_matrix, it will be converted into a sparsecsr_matrix.- yIgnored
Not used, present here for API consistency by convention.
- Returns
- self
-
fit_predict(self, X, y=None)[source]¶ Perform spectral clustering from features, or affinity matrix, and return cluster labels.
- Parameters
- Xarray-like or sparse matrix, shape (n_samples, n_features), or array-like, shape (n_samples, n_samples)
Training instances to cluster, or similarities / affinities between instances if
affinity='precomputed'. If a sparse matrix is provided in a format other thancsr_matrix,csc_matrix, orcoo_matrix, it will be converted into a sparsecsr_matrix.- yIgnored
Not used, present here for API consistency by convention.
- Returns
- labelsndarray, shape (n_samples,)
Cluster labels.
-
get_params(self, deep=True)[source]¶ Get parameters for this estimator.
- Parameters
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns
- paramsmapping of string to any
Parameter names mapped to their values.
-
set_params(self, **params)[source]¶ Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as pipelines). The latter have parameters of the form
<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters
- **paramsdict
Estimator parameters.
- Returns
- selfobject
Estimator instance.