sklearn.feature_extraction.FeatureHasher¶
- class sklearn.feature_extraction.FeatureHasher(n_features=1048576, *, input_type='dict', dtype=<class 'numpy.float64'>, alternate_sign=True)[source]¶
Implements feature hashing, aka the hashing trick.
This class turns sequences of symbolic feature names (strings) into scipy.sparse matrices, using a hash function to compute the matrix column corresponding to a name. The hash function employed is the signed 32-bit version of Murmurhash3.
Feature names of type byte string are used as-is. Unicode strings are converted to UTF-8 first, but no Unicode normalization is done. Feature values must be (finite) numbers.
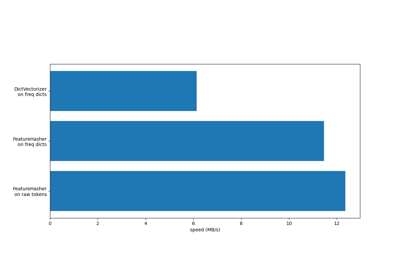
This class is a low-memory alternative to DictVectorizer and CountVectorizer, intended for large-scale (online) learning and situations where memory is tight, e.g. when running prediction code on embedded devices.
Read more in the User Guide.
New in version 0.13.
- Parameters:
- n_featuresint, default=2**20
The number of features (columns) in the output matrices. Small numbers of features are likely to cause hash collisions, but large numbers will cause larger coefficient dimensions in linear learners.
- input_typestr, default=’dict’
Choose a string from {‘dict’, ‘pair’, ‘string’}. Either “dict” (the default) to accept dictionaries over (feature_name, value); “pair” to accept pairs of (feature_name, value); or “string” to accept single strings. feature_name should be a string, while value should be a number. In the case of “string”, a value of 1 is implied. The feature_name is hashed to find the appropriate column for the feature. The value’s sign might be flipped in the output (but see non_negative, below).
- dtypenumpy dtype, default=np.float64
The type of feature values. Passed to scipy.sparse matrix constructors as the dtype argument. Do not set this to bool, np.boolean or any unsigned integer type.
- alternate_signbool, default=True
When True, an alternating sign is added to the features as to approximately conserve the inner product in the hashed space even for small n_features. This approach is similar to sparse random projection.
Changed in version 0.19:
alternate_signreplaces the now deprecatednon_negativeparameter.
See also
DictVectorizerVectorizes string-valued features using a hash table.
sklearn.preprocessing.OneHotEncoderHandles nominal/categorical features.
Notes
This estimator is stateless and does not need to be fitted. However, we recommend to call
fit_transforminstead oftransform, as parameter validation is only performed infit.Examples
>>> from sklearn.feature_extraction import FeatureHasher >>> h = FeatureHasher(n_features=10) >>> D = [{'dog': 1, 'cat':2, 'elephant':4},{'dog': 2, 'run': 5}] >>> f = h.transform(D) >>> f.toarray() array([[ 0., 0., -4., -1., 0., 0., 0., 0., 0., 2.], [ 0., 0., 0., -2., -5., 0., 0., 0., 0., 0.]])
With
input_type="string", the input must be an iterable over iterables of strings:>>> h = FeatureHasher(n_features=8, input_type="string") >>> raw_X = [["dog", "cat", "snake"], ["snake", "dog"], ["cat", "bird"]] >>> f = h.transform(raw_X) >>> f.toarray() array([[ 0., 0., 0., -1., 0., -1., 0., 1.], [ 0., 0., 0., -1., 0., -1., 0., 0.], [ 0., -1., 0., 0., 0., 0., 0., 1.]])
Methods
fit([X, y])Only validates estimator's parameters.
fit_transform(X[, y])Fit to data, then transform it.
get_params([deep])Get parameters for this estimator.
set_output(*[, transform])Set output container.
set_params(**params)Set the parameters of this estimator.
transform(raw_X)Transform a sequence of instances to a scipy.sparse matrix.
- fit(X=None, y=None)[source]¶
Only validates estimator’s parameters.
This method allows to: (i) validate the estimator’s parameters and (ii) be consistent with the scikit-learn transformer API.
- Parameters:
- XIgnored
Not used, present here for API consistency by convention.
- yIgnored
Not used, present here for API consistency by convention.
- Returns:
- selfobject
FeatureHasher class instance.
- fit_transform(X, y=None, **fit_params)[source]¶
Fit to data, then transform it.
Fits transformer to
Xandywith optional parametersfit_paramsand returns a transformed version ofX.- Parameters:
- Xarray-like of shape (n_samples, n_features)
Input samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs), default=None
Target values (None for unsupervised transformations).
- **fit_paramsdict
Additional fit parameters.
- Returns:
- X_newndarray array of shape (n_samples, n_features_new)
Transformed array.
- get_params(deep=True)[source]¶
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- set_output(*, transform=None)[source]¶
Set output container.
See Introducing the set_output API for an example on how to use the API.
- Parameters:
- transform{“default”, “pandas”}, default=None
Configure output of
transformandfit_transform."default": Default output format of a transformer"pandas": DataFrame outputNone: Transform configuration is unchanged
- Returns:
- selfestimator instance
Estimator instance.
- set_params(**params)[source]¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
- transform(raw_X)[source]¶
Transform a sequence of instances to a scipy.sparse matrix.
- Parameters:
- raw_Xiterable over iterable over raw features, length = n_samples
Samples. Each sample must be iterable an (e.g., a list or tuple) containing/generating feature names (and optionally values, see the input_type constructor argument) which will be hashed. raw_X need not support the len function, so it can be the result of a generator; n_samples is determined on the fly.
- Returns:
- Xsparse matrix of shape (n_samples, n_features)
Feature matrix, for use with estimators or further transformers.