sklearn.manifold.LocallyLinearEmbedding¶
- class sklearn.manifold.LocallyLinearEmbedding(n_neighbors=5, n_components=2, reg=0.001, eigen_solver='auto', tol=1e-06, max_iter=100, method='standard', hessian_tol=0.0001, modified_tol=1e-12, neighbors_algorithm='auto', random_state=None)[source]¶
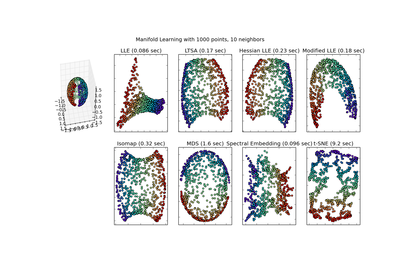
Locally Linear Embedding
Parameters: n_neighbors : integer
number of neighbors to consider for each point.
n_components : integer
number of coordinates for the manifold
reg : float
regularization constant, multiplies the trace of the local covariance matrix of the distances.
eigen_solver : string, {‘auto’, ‘arpack’, ‘dense’}
auto : algorithm will attempt to choose the best method for input data
- arpack : use arnoldi iteration in shift-invert mode.
For this method, M may be a dense matrix, sparse matrix, or general linear operator. Warning: ARPACK can be unstable for some problems. It is best to try several random seeds in order to check results.
- dense : use standard dense matrix operations for the eigenvalue
decomposition. For this method, M must be an array or matrix type. This method should be avoided for large problems.
tol : float, optional
Tolerance for ‘arpack’ method Not used if eigen_solver==’dense’.
max_iter : integer
maximum number of iterations for the arpack solver. Not used if eigen_solver==’dense’.
method : string (‘standard’, ‘hessian’, ‘modified’ or ‘ltsa’)
- standard : use the standard locally linear embedding algorithm. see
reference [1]
- hessian : use the Hessian eigenmap method. This method requires
n_neighbors > n_components * (1 + (n_components + 1) / 2 see reference [2]
- modified : use the modified locally linear embedding algorithm.
see reference [3]
- ltsa : use local tangent space alignment algorithm
see reference [4]
hessian_tol : float, optional
Tolerance for Hessian eigenmapping method. Only used if method == 'hessian'
modified_tol : float, optional
Tolerance for modified LLE method. Only used if method == 'modified'
neighbors_algorithm : string [‘auto’|’brute’|’kd_tree’|’ball_tree’]
algorithm to use for nearest neighbors search, passed to neighbors.NearestNeighbors instance
random_state: numpy.RandomState or int, optional :
The generator or seed used to determine the starting vector for arpack iterations. Defaults to numpy.random.
Attributes: embedding_vectors_ : array-like, shape [n_components, n_samples]
Stores the embedding vectors
reconstruction_error_ : float
Reconstruction error associated with embedding_vectors_
nbrs_ : NearestNeighbors object
Stores nearest neighbors instance, including BallTree or KDtree if applicable.
References
[R147] Roweis, S. & Saul, L. Nonlinear dimensionality reduction by locally linear embedding. Science 290:2323 (2000). [R148] Donoho, D. & Grimes, C. Hessian eigenmaps: Locally linear embedding techniques for high-dimensional data. Proc Natl Acad Sci U S A. 100:5591 (2003). [R149] Zhang, Z. & Wang, J. MLLE: Modified Locally Linear Embedding Using Multiple Weights. http://citeseerx.ist.psu.edu/viewdoc/summary?doi=10.1.1.70.382 [R150] Zhang, Z. & Zha, H. Principal manifolds and nonlinear dimensionality reduction via tangent space alignment. Journal of Shanghai Univ. 8:406 (2004) Methods
fit(X[, y]) Compute the embedding vectors for data X fit_transform(X[, y]) Compute the embedding vectors for data X and transform X. get_params([deep]) Get parameters for this estimator. set_params(**params) Set the parameters of this estimator. transform(X) Transform new points into embedding space. - __init__(n_neighbors=5, n_components=2, reg=0.001, eigen_solver='auto', tol=1e-06, max_iter=100, method='standard', hessian_tol=0.0001, modified_tol=1e-12, neighbors_algorithm='auto', random_state=None)[source]¶
- fit(X, y=None)[source]¶
Compute the embedding vectors for data X
Parameters: X : array-like of shape [n_samples, n_features]
training set.
Returns: self : returns an instance of self.
- fit_transform(X, y=None)[source]¶
Compute the embedding vectors for data X and transform X.
Parameters: X : array-like of shape [n_samples, n_features]
training set.
Returns: X_new: array-like, shape (n_samples, n_components) :
- get_params(deep=True)[source]¶
Get parameters for this estimator.
Parameters: deep: boolean, optional :
If True, will return the parameters for this estimator and contained subobjects that are estimators.
Returns: params : mapping of string to any
Parameter names mapped to their values.
- set_params(**params)[source]¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as pipelines). The former have parameters of the form <component>__<parameter> so that it’s possible to update each component of a nested object.
Returns: self :
- transform(X)[source]¶
Transform new points into embedding space.
Parameters: X : array-like, shape = [n_samples, n_features] Returns: X_new : array, shape = [n_samples, n_components] Notes
Because of scaling performed by this method, it is discouraged to use it together with methods that are not scale-invariant (like SVMs)