sklearn.mixture.DPGMM¶
- class sklearn.mixture.DPGMM(n_components=1, covariance_type='diag', alpha=1.0, random_state=None, thresh=0.01, verbose=False, min_covar=None, n_iter=10, params='wmc', init_params='wmc')¶
Variational Inference for the Infinite Gaussian Mixture Model.
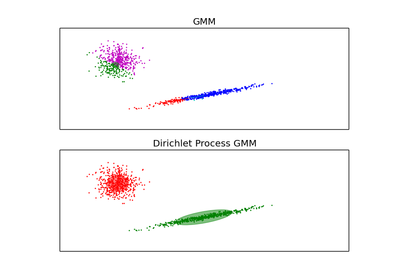
DPGMM stands for Dirichlet Process Gaussian Mixture Model, and it is an infinite mixture model with the Dirichlet Process as a prior distribution on the number of clusters. In practice the approximate inference algorithm uses a truncated distribution with a fixed maximum number of components, but almost always the number of components actually used depends on the data.
Stick-breaking Representation of a Gaussian mixture model probability distribution. This class allows for easy and efficient inference of an approximate posterior distribution over the parameters of a Gaussian mixture model with a variable number of components (smaller than the truncation parameter n_components).
Initialization is with normally-distributed means and identity covariance, for proper convergence.
Parameters: n_components: int, optional :
Number of mixture components. Defaults to 1.
covariance_type: string, optional :
String describing the type of covariance parameters to use. Must be one of ‘spherical’, ‘tied’, ‘diag’, ‘full’. Defaults to ‘diag’.
alpha: float, optional :
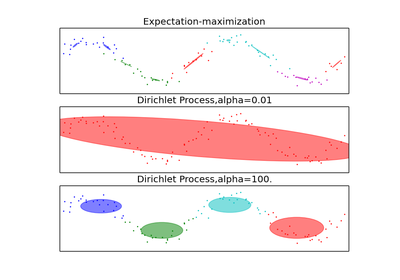
Real number representing the concentration parameter of the dirichlet process. Intuitively, the Dirichlet Process is as likely to start a new cluster for a point as it is to add that point to a cluster with alpha elements. A higher alpha means more clusters, as the expected number of clusters is alpha*log(N). Defaults to 1.
thresh : float, optional
Convergence threshold.
n_iter : int, optional
Maximum number of iterations to perform before convergence.
params : string, optional
Controls which parameters are updated in the training process. Can contain any combination of ‘w’ for weights, ‘m’ for means, and ‘c’ for covars. Defaults to ‘wmc’.
init_params : string, optional
Controls which parameters are updated in the initialization process. Can contain any combination of ‘w’ for weights, ‘m’ for means, and ‘c’ for covars. Defaults to ‘wmc’.
Attributes: covariance_type : string
String describing the type of covariance parameters used by the DP-GMM. Must be one of ‘spherical’, ‘tied’, ‘diag’, ‘full’.
n_components : int
Number of mixture components.
`weights_` : array, shape (n_components,)
Mixing weights for each mixture component.
`means_` : array, shape (n_components, n_features)
Mean parameters for each mixture component.
`precs_` : array
Precision (inverse covariance) parameters for each mixture component. The shape depends on covariance_type:
(`n_components`, 'n_features') if 'spherical', (`n_features`, `n_features`) if 'tied', (`n_components`, `n_features`) if 'diag', (`n_components`, `n_features`, `n_features`) if 'full'
`converged_` : bool
True when convergence was reached in fit(), False otherwise.
See also
Methods
aic(X) Akaike information criterion for the current model fit bic(X) Bayesian information criterion for the current model fit eval(*args, **kwargs) DEPRECATED: DPGMM.eval was renamed to DPGMM.score_samples in 0.14 and will be removed in 0.16. fit(X) Estimate model parameters with the variational algorithm. get_params([deep]) Get parameters for this estimator. lower_bound(X, z) returns a lower bound on model evidence based on X and membership predict(X) Predict label for data. predict_proba(X) Predict posterior probability of data under each Gaussian in the model. sample([n_samples, random_state]) Generate random samples from the model. score(X) Compute the log probability under the model. score_samples(X) Return the likelihood of the data under the model. set_params(**params) Set the parameters of this estimator. - __init__(n_components=1, covariance_type='diag', alpha=1.0, random_state=None, thresh=0.01, verbose=False, min_covar=None, n_iter=10, params='wmc', init_params='wmc')¶
- aic(X)¶
Akaike information criterion for the current model fit and the proposed data
Parameters: X : array of shape(n_samples, n_dimensions) Returns: aic: float (the lower the better) :
- bic(X)¶
Bayesian information criterion for the current model fit and the proposed data
Parameters: X : array of shape(n_samples, n_dimensions) Returns: bic: float (the lower the better) :
- eval(*args, **kwargs)¶
DEPRECATED: DPGMM.eval was renamed to DPGMM.score_samples in 0.14 and will be removed in 0.16.
- fit(X)¶
Estimate model parameters with the variational algorithm.
For a full derivation and description of the algorithm see doc/modules/dp-derivation.rst or http://scikit-learn.org/stable/modules/dp-derivation.html#
A initialization step is performed before entering the em algorithm. If you want to avoid this step, set the keyword argument init_params to the empty string ‘’ when when creating the object. Likewise, if you would like just to do an initialization, set n_iter=0.
Parameters: X : array_like, shape (n, n_features)
List of n_features-dimensional data points. Each row corresponds to a single data point.
- get_params(deep=True)¶
Get parameters for this estimator.
Parameters: deep: boolean, optional :
If True, will return the parameters for this estimator and contained subobjects that are estimators.
Returns: params : mapping of string to any
Parameter names mapped to their values.
- lower_bound(X, z)¶
returns a lower bound on model evidence based on X and membership
- predict(X)¶
Predict label for data.
Parameters: X : array-like, shape = [n_samples, n_features] Returns: C : array, shape = (n_samples,)
- predict_proba(X)¶
Predict posterior probability of data under each Gaussian in the model.
Parameters: X : array-like, shape = [n_samples, n_features]
Returns: responsibilities : array-like, shape = (n_samples, n_components)
Returns the probability of the sample for each Gaussian (state) in the model.
- sample(n_samples=1, random_state=None)¶
Generate random samples from the model.
Parameters: n_samples : int, optional
Number of samples to generate. Defaults to 1.
Returns: X : array_like, shape (n_samples, n_features)
List of samples
- score(X)¶
Compute the log probability under the model.
Parameters: X : array_like, shape (n_samples, n_features)
List of n_features-dimensional data points. Each row corresponds to a single data point.
Returns: logprob : array_like, shape (n_samples,)
Log probabilities of each data point in X
- score_samples(X)¶
Return the likelihood of the data under the model.
Compute the bound on log probability of X under the model and return the posterior distribution (responsibilities) of each mixture component for each element of X.
This is done by computing the parameters for the mean-field of z for each observation.
Parameters: X : array_like, shape (n_samples, n_features)
List of n_features-dimensional data points. Each row corresponds to a single data point.
Returns: logprob : array_like, shape (n_samples,)
Log probabilities of each data point in X
responsibilities: array_like, shape (n_samples, n_components) :
Posterior probabilities of each mixture component for each observation
- set_params(**params)¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as pipelines). The former have parameters of the form <component>__<parameter> so that it’s possible to update each component of a nested object.
Returns: self :