Note
Click here to download the full example code or to run this example in your browser via Binder
Cross-validation on diabetes Dataset Exercise¶
A tutorial exercise which uses cross-validation with linear models.
This exercise is used in the Cross-validated estimators part of the Model selection: choosing estimators and their parameters section of the A tutorial on statistical-learning for scientific data processing.
Load dataset and apply GridSearchCV¶
import matplotlib.pyplot as plt
import numpy as np
from sklearn import datasets
from sklearn.linear_model import Lasso
from sklearn.model_selection import GridSearchCV
X, y = datasets.load_diabetes(return_X_y=True)
X = X[:150]
y = y[:150]
lasso = Lasso(random_state=0, max_iter=10000)
alphas = np.logspace(-4, -0.5, 30)
tuned_parameters = [{"alpha": alphas}]
n_folds = 5
clf = GridSearchCV(lasso, tuned_parameters, cv=n_folds, refit=False)
clf.fit(X, y)
scores = clf.cv_results_["mean_test_score"]
scores_std = clf.cv_results_["std_test_score"]
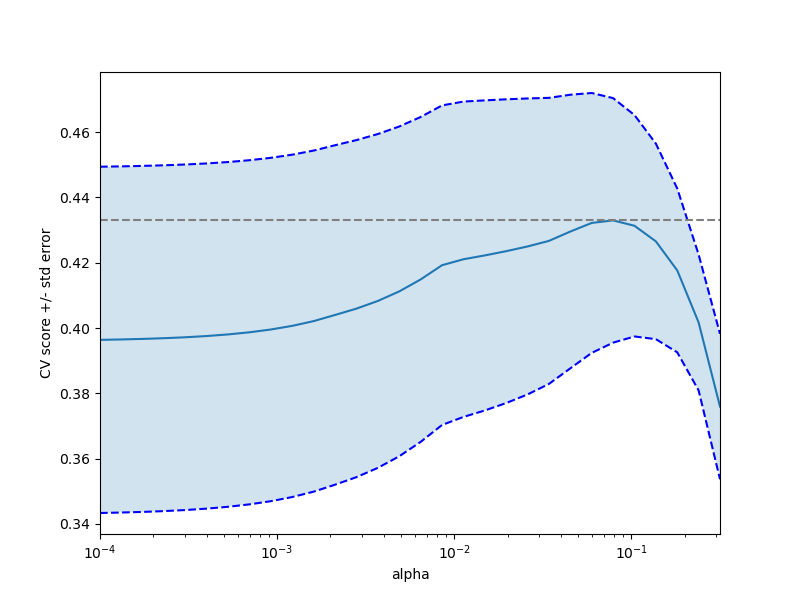
Plot error lines showing +/- std. errors of the scores¶
plt.figure().set_size_inches(8, 6)
plt.semilogx(alphas, scores)
std_error = scores_std / np.sqrt(n_folds)
plt.semilogx(alphas, scores + std_error, "b--")
plt.semilogx(alphas, scores - std_error, "b--")
# alpha=0.2 controls the translucency of the fill color
plt.fill_between(alphas, scores + std_error, scores - std_error, alpha=0.2)
plt.ylabel("CV score +/- std error")
plt.xlabel("alpha")
plt.axhline(np.max(scores), linestyle="--", color=".5")
plt.xlim([alphas[0], alphas[-1]])
(9.999999999999999e-05, 0.31622776601683794)
Bonus: how much can you trust the selection of alpha?¶
# To answer this question we use the LassoCV object that sets its alpha
# parameter automatically from the data by internal cross-validation (i.e. it
# performs cross-validation on the training data it receives).
# We use external cross-validation to see how much the automatically obtained
# alphas differ across different cross-validation folds.
from sklearn.linear_model import LassoCV
from sklearn.model_selection import KFold
lasso_cv = LassoCV(alphas=alphas, random_state=0, max_iter=10000)
k_fold = KFold(3)
print("Answer to the bonus question:", "how much can you trust the selection of alpha?")
print()
print("Alpha parameters maximising the generalization score on different")
print("subsets of the data:")
for k, (train, test) in enumerate(k_fold.split(X, y)):
lasso_cv.fit(X[train], y[train])
print(
"[fold {0}] alpha: {1:.5f}, score: {2:.5f}".format(
k, lasso_cv.alpha_, lasso_cv.score(X[test], y[test])
)
)
print()
print("Answer: Not very much since we obtained different alphas for different")
print("subsets of the data and moreover, the scores for these alphas differ")
print("quite substantially.")
plt.show()
Answer to the bonus question: how much can you trust the selection of alpha?
Alpha parameters maximising the generalization score on different
subsets of the data:
[fold 0] alpha: 0.05968, score: 0.54209
[fold 1] alpha: 0.04520, score: 0.15521
[fold 2] alpha: 0.07880, score: 0.45192
Answer: Not very much since we obtained different alphas for different
subsets of the data and moreover, the scores for these alphas differ
quite substantially.
Total running time of the script: ( 0 minutes 0.477 seconds)