sklearn.model_selection.StratifiedKFold¶
-
class
sklearn.model_selection.StratifiedKFold(n_splits=5, *, shuffle=False, random_state=None)[source]¶ Stratified K-Folds cross-validator
Provides train/test indices to split data in train/test sets.
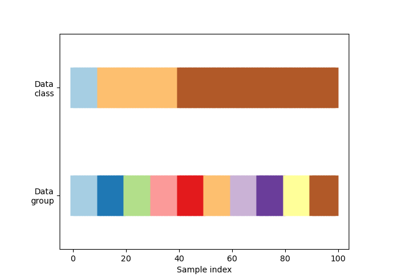
This cross-validation object is a variation of KFold that returns stratified folds. The folds are made by preserving the percentage of samples for each class.
Read more in the User Guide.
- Parameters
- n_splitsint, default=5
Number of folds. Must be at least 2.
Changed in version 0.22:
n_splitsdefault value changed from 3 to 5.- shufflebool, default=False
Whether to shuffle each class’s samples before splitting into batches. Note that the samples within each split will not be shuffled.
- random_stateint or RandomState instance, default=None
When
shuffleis True,random_stateaffects the ordering of the indices, which controls the randomness of each fold for each class. Otherwise, leaverandom_stateasNone. Pass an int for reproducible output across multiple function calls. See Glossary.
See also
RepeatedStratifiedKFoldRepeats Stratified K-Fold n times.
Notes
The implementation is designed to:
Generate test sets such that all contain the same distribution of classes, or as close as possible.
Be invariant to class label: relabelling
y = ["Happy", "Sad"]toy = [1, 0]should not change the indices generated.Preserve order dependencies in the dataset ordering, when
shuffle=False: all samples from class k in some test set were contiguous in y, or separated in y by samples from classes other than k.Generate test sets where the smallest and largest differ by at most one sample.
Changed in version 0.22: The previous implementation did not follow the last constraint.
Examples
>>> import numpy as np >>> from sklearn.model_selection import StratifiedKFold >>> X = np.array([[1, 2], [3, 4], [1, 2], [3, 4]]) >>> y = np.array([0, 0, 1, 1]) >>> skf = StratifiedKFold(n_splits=2) >>> skf.get_n_splits(X, y) 2 >>> print(skf) StratifiedKFold(n_splits=2, random_state=None, shuffle=False) >>> for train_index, test_index in skf.split(X, y): ... print("TRAIN:", train_index, "TEST:", test_index) ... X_train, X_test = X[train_index], X[test_index] ... y_train, y_test = y[train_index], y[test_index] TRAIN: [1 3] TEST: [0 2] TRAIN: [0 2] TEST: [1 3]
Methods
get_n_splits([X, y, groups])Returns the number of splitting iterations in the cross-validator
split(X, y[, groups])Generate indices to split data into training and test set.
-
__init__(n_splits=5, *, shuffle=False, random_state=None)[source]¶ Initialize self. See help(type(self)) for accurate signature.
-
get_n_splits(X=None, y=None, groups=None)[source]¶ Returns the number of splitting iterations in the cross-validator
- Parameters
- Xobject
Always ignored, exists for compatibility.
- yobject
Always ignored, exists for compatibility.
- groupsobject
Always ignored, exists for compatibility.
- Returns
- n_splitsint
Returns the number of splitting iterations in the cross-validator.
-
split(X, y, groups=None)[source]¶ Generate indices to split data into training and test set.
- Parameters
- Xarray-like of shape (n_samples, n_features)
Training data, where n_samples is the number of samples and n_features is the number of features.
Note that providing
yis sufficient to generate the splits and hencenp.zeros(n_samples)may be used as a placeholder forXinstead of actual training data.- yarray-like of shape (n_samples,)
The target variable for supervised learning problems. Stratification is done based on the y labels.
- groupsobject
Always ignored, exists for compatibility.
- Yields
- trainndarray
The training set indices for that split.
- testndarray
The testing set indices for that split.
Notes
Randomized CV splitters may return different results for each call of split. You can make the results identical by setting
random_stateto an integer.