Robust linear estimator fitting¶
Here a sine function is fit with a polynomial of order 3, for values close to zero.
Robust fitting is demoed in different situations:
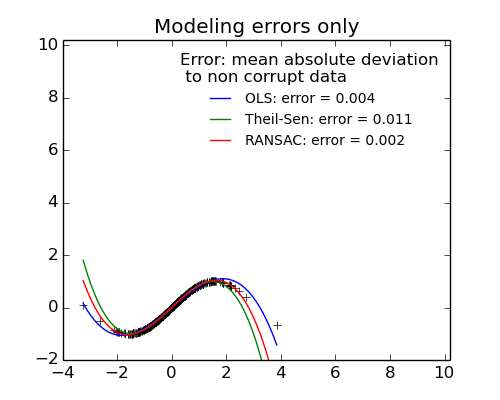
- No measurement errors, only modelling errors (fitting a sine with a polynomial)
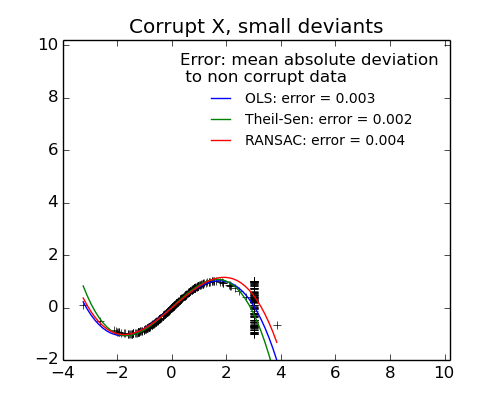
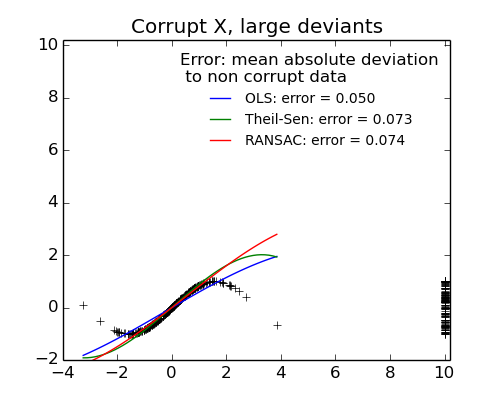
- Measurement errors in X
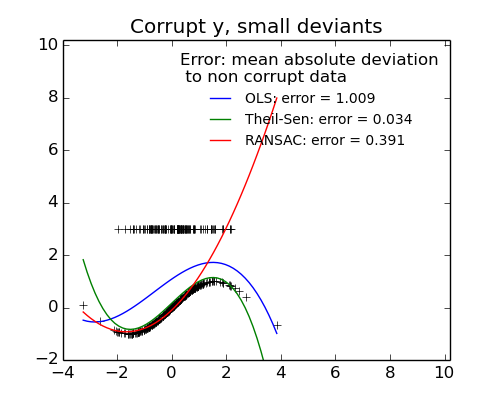
- Measurement errors in y
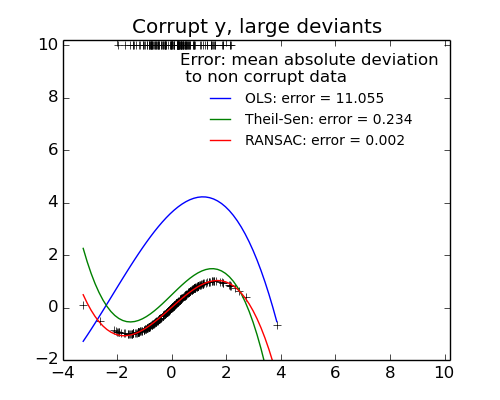
The median absolute deviation to non corrupt new data is used to judge the quality of the prediction.
What we can see that:
- RANSAC is good for strong outliers in the y direction
- TheilSen is good for small outliers, both in direction X and y, but has a break point above which it performs worst than OLS.
Python source code: plot_robust_fit.py
from matplotlib import pyplot as plt
import numpy as np
from sklearn import linear_model, metrics
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
np.random.seed(42)
X = np.random.normal(size=400)
y = np.sin(X)
# Make sure that it X is 2D
X = X[:, np.newaxis]
X_test = np.random.normal(size=200)
y_test = np.sin(X_test)
X_test = X_test[:, np.newaxis]
y_errors = y.copy()
y_errors[::3] = 3
X_errors = X.copy()
X_errors[::3] = 3
y_errors_large = y.copy()
y_errors_large[::3] = 10
X_errors_large = X.copy()
X_errors_large[::3] = 10
estimators = [('OLS', linear_model.LinearRegression()),
('Theil-Sen', linear_model.TheilSenRegressor(random_state=42)),
('RANSAC', linear_model.RANSACRegressor(random_state=42)), ]
x_plot = np.linspace(X.min(), X.max())
for title, this_X, this_y in [
('Modeling errors only', X, y),
('Corrupt X, small deviants', X_errors, y),
('Corrupt y, small deviants', X, y_errors),
('Corrupt X, large deviants', X_errors_large, y),
('Corrupt y, large deviants', X, y_errors_large)]:
plt.figure(figsize=(5, 4))
plt.plot(this_X[:, 0], this_y, 'k+')
for name, estimator in estimators:
model = make_pipeline(PolynomialFeatures(3), estimator)
model.fit(this_X, this_y)
mse = metrics.mean_squared_error(model.predict(X_test), y_test)
y_plot = model.predict(x_plot[:, np.newaxis])
plt.plot(x_plot, y_plot,
label='%s: error = %.3f' % (name, mse))
plt.legend(loc='best', frameon=False,
title='Error: mean absolute deviation\n to non corrupt data')
plt.xlim(-4, 10.2)
plt.ylim(-2, 10.2)
plt.title(title)
plt.show()
Total running time of the example: 6.00 seconds ( 0 minutes 6.00 seconds)