sklearn.calibration.CalibratedClassifierCV¶
- class sklearn.calibration.CalibratedClassifierCV(base_estimator=None, method='sigmoid', cv=3)[source]¶
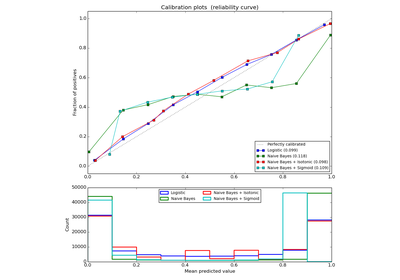
Probability calibration with isotonic regression or sigmoid.
With this class, the base_estimator is fit on the train set of the cross-validation generator and the test set is used for calibration. The probabilities for each of the folds are then averaged for prediction. In case that cv=”prefit” is passed to __init__, it is it is assumed that base_estimator has been fitted already and all data is used for calibration. Note that data for fitting the classifier and for calibrating it must be disjpint.
Parameters: base_estimator : instance BaseEstimator
The classifier whose output decision function needs to be calibrated to offer more accurate predict_proba outputs. If cv=prefit, the classifier must have been fit already on data.
method : ‘sigmoid’ | ‘isotonic’
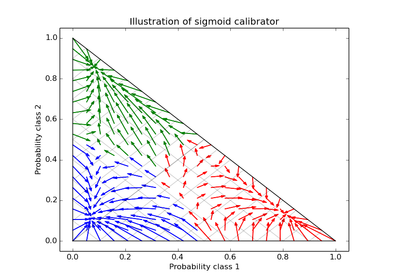
The method to use for calibration. Can be ‘sigmoid’ which corresponds to Platt’s method or ‘isotonic’ which is a non-parameteric approach. It is not advised to use isotonic calibration with too few calibration samples (<<1000) since it tends to overfit. Use sigmoids (Platt’s calibration) in this case.
cv : integer or cross-validation generator or “prefit”, optional
If an integer is passed, it is the number of folds (default 3). Specific cross-validation objects can be passed, see sklearn.cross_validation module for the list of possible objects. If “prefit” is passed, it is assumed that base_estimator has been fitted already and all data is used for calibration.
Attributes: classes_ : array, shape (n_classes)
The class labels.
calibrated_classifiers_: list (len() equal to cv or 1 if cv == “prefit”) :
The list of calibrated classifiers, one for each crossvalidation fold, which has been fitted on all but the validation fold and calibrated on the validation fold.
References
[R103] Obtaining calibrated probability estimates from decision trees and naive Bayesian classifiers, B. Zadrozny & C. Elkan, ICML 2001 [R104] Transforming Classifier Scores into Accurate Multiclass Probability Estimates, B. Zadrozny & C. Elkan, (KDD 2002) [R105] Probabilistic Outputs for Support Vector Machines and Comparisons to Regularized Likelihood Methods, J. Platt, (1999) [R106] Predicting Good Probabilities with Supervised Learning, A. Niculescu-Mizil & R. Caruana, ICML 2005 Methods
fit(X, y[, sample_weight]) Fit the calibrated model get_params([deep]) Get parameters for this estimator. predict(X) Predict the target of new samples. predict_proba(X) Posterior probabilities of classification score(X, y[, sample_weight]) Returns the mean accuracy on the given test data and labels. set_params(**params) Set the parameters of this estimator. - fit(X, y, sample_weight=None)[source]¶
Fit the calibrated model
Parameters: X : array-like, shape (n_samples, n_features)
Training data.
y : array-like, shape (n_samples,)
Target values.
sample_weight : array-like, shape = [n_samples] or None
Sample weights. If None, then samples are equally weighted.
Returns: self : object
Returns an instance of self.
- get_params(deep=True)[source]¶
Get parameters for this estimator.
Parameters: deep: boolean, optional :
If True, will return the parameters for this estimator and contained subobjects that are estimators.
Returns: params : mapping of string to any
Parameter names mapped to their values.
- predict(X)[source]¶
Predict the target of new samples. Can be different from the prediction of the uncalibrated classifier.
Parameters: X : array-like, shape (n_samples, n_features)
The samples.
Returns: C : array, shape (n_samples,)
The predicted class.
- predict_proba(X)[source]¶
Posterior probabilities of classification
This function returns posterior probabilities of classification according to each class on an array of test vectors X.
Parameters: X : array-like, shape (n_samples, n_features)
The samples.
Returns: C : array, shape (n_samples, n_classes)
The predicted probas.
- score(X, y, sample_weight=None)[source]¶
Returns the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
Parameters: X : array-like, shape = (n_samples, n_features)
Test samples.
y : array-like, shape = (n_samples) or (n_samples, n_outputs)
True labels for X.
sample_weight : array-like, shape = [n_samples], optional
Sample weights.
Returns: score : float
Mean accuracy of self.predict(X) wrt. y.
- set_params(**params)[source]¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as pipelines). The former have parameters of the form <component>__<parameter> so that it’s possible to update each component of a nested object.
Returns: self :